Abstract
Chromosomal translocations resulting in chimeric fusion genes are prototypic for pediatric leukemia patients. The most known fusions are ETV6-RUNX1 or BCR-ABL1 in B-cell progenitor (BCP)-ALL, and rearrangements of MLL in pediatric ALL and AML. Genome-wide sequencing projects have revealed additional, recurrent gene mutations in B cell malignancies. One of these mutations comprises the IKZF1 gene, encoding the IKAROS transcription factor which is one of the essential transcription factors driving lymphoid development. IKZF1 deletions were first identified by SNP arrays in ALL patients, and later identified with a high prevalence in BCR-ABL1+ patients. IKZF1 deletions turned out to be an independent prognostic marker associated with a poor outcome. Here, we characterized IKZF1 deletions in pediatric BCP-ALL patients by combining MLPA mapping experiments with long distance inverse PCR. The aim of our study was also to compare existing methods with our approach. Our attempt confirmed many of the existing data but revealed a more complex pattern of recombination sites, including a total of 4 recombination hotspots. This extended knowledge was translated into a novel, multiplex PCR assay that allows to perform IKZF1 deletion analyses by using a 2-tube PCR approach.
Keywords: Childhood leukemia, cancer genetics, gene deletion, IKAROS, IKZF1, leukemia markers
Introduction
Genetic prerequisites for developing acute leukemia have been characterized during the past decades. Besides well known fusion genes novel genome-wide technologies (e.g. high-resolution SNP arrays or whole genome sequencing) also allow to identify complementing gene mutations. This allows to draw important conclusions about genetic prerequisites of hematological malignancies [1]. Mutant genes have been identified that affect general pathways important for DNA repair, cell cycle, epigenetics, transcription, differentiation and signaling. All these changes are helping to render pre-cancerous cells independent of their environment and to convert them into a tumor-initiating cells.
One of the identified gene mutations concerns the IKZF1 gene located at 7p12, encoding the IKAROS transcription factor. IKAROS has been identified to be involved in B lineage development [2]. The biology of IKZF1 is quite complex because this gene, consisting of 8 exons, encodes 11 different splice variants (see Figure 1A) [3-5]. Five of those translate into proteins that are transcriptional activators (1, 2, 2a, 3, 3a) while six result in dominant-negative IKAROS versions (4, 4a, 5, 6, 7, 8). Importantly, exons 4 to 6 of the IKZF1 gene encode four independent zinc finger domains of which three are necessary for DNA binding. The large exon 8 encodes 2 additional zinc fingers which, however, are required for homodimerization of the different IKAROS variants. Binding of any dominant-negative IKAROS version to an active one always result in transcriptional repression. Moreover, IKAROS heterodimers are frequently associated with pericentromeric heterochromatin, which is in line with the finding that IKAROS transcription factors are mainly involved in pathways that control the shut-off of genes during development [5,6]. On the other hand, IKAROS is also involved in the regulation of the hypothalamic-pituitary-adrenal (HPA) axis, known to control body stress reactions and to coordinate the inflammatory/immune response (see Figure 1B) [7].
Figure 1.
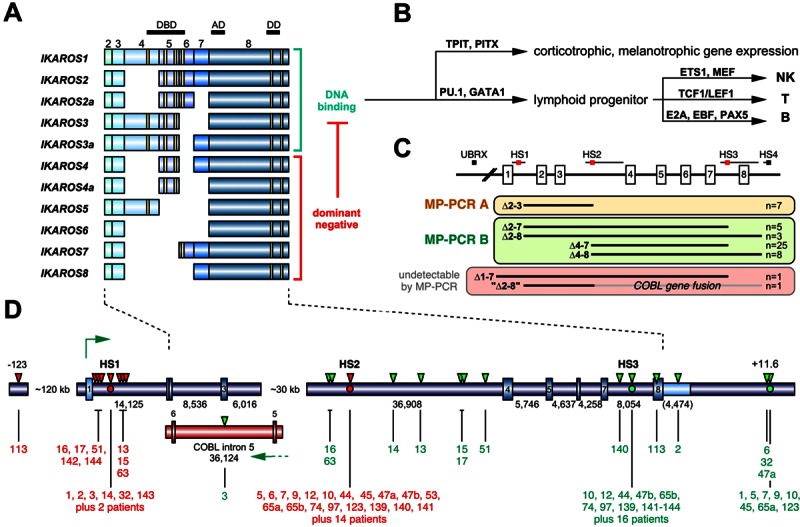
The human IKZF1 gene and its role in the biology. A. Summary of IKZF1 transcripts. IKZF1 is composed of 8 exons, of which the first one is noncoding. Several splice forms of IKZF1 transcripts have been described in the literature (n=11). Important functional domains are encoded by exon 4-6 (4 zinc fingers) and exon 8 (2 zinc fingers). At least 3 N-terminal zinc fingers are necessary for DNA binding. Therefore, 6 out of 11 IKZF1 splice variants are described to encode dominant-negative versions of IKAROS, because they are still able to dimerize with bona fide protein variants via the C-terminal zinc finger domains. B. The complex biology behind IKAROS protein variants is necessary to (a) regulate hypothalamic-pituitary-adrenal axis. IKAROS steers with TPIT and PITX transcription factors the corticotrophic and melanotrophic gene expression; (b) control hematopoietic development. IKAROS determines the devlopment of NK, T and B cells in concert with PU.1, GATA factors and the other depicted transcription factors. C. Overview of the results of the MLPA analysis of the 48 identified IKZF1-deleted BCP-ALL patients (that exhibited 50 breakpoints). All deletions within the IKZF1 gene can be clustered into 4 different groups (indicated by the 3 colored panels and a fourth group of leukemia patients that lost their complete IKZF1 allele). Four different recombination hotspots (HS1-4) and a single upstream breakpoint (UBRX) were identified. Two of these 4 patient classes can be readily analyzed by the established multiplex PCR reactions A and B, respectively. The third group can only be investigated by LDI-PCR analyses. The fourth group can only be detected by MLPA analyses. D. The gene structure of human IKZF1 is shown. Exons are depicted as rectangles. Below: Sizes of the individual introns and the 3-NTR (in brackets). There is a gap of about 30 kb for which no sequence data are available from public databases. Thus intron 3 must be divided in intron 3a (6,016 bp) and intron 3b (36,908 bp), separated by a gap (hampering our LDI-PCR analyses because no oligonucleotides can be designed). Published fusion sites between HS1 and HS3 or HS2 and HS3 are indicated by red or green dots. All fusion sites identified in this study are indicated by red (5’-breakpoint) and green (3’-breakpoint) triangles above the gene structure and by black lines below the gene structure. Each breakpoint could be assigned to a patient by its UPN. In addition, a small portion of the COBL gene is shown. The tail-to-tail gene fusion between IKZF1 intron 1 and COBL intron 5 is due to an interstitial deletion of about 800,000 kb between 7p12 and 7p12.1. The novel HS4 breakpoint is located about 11.6 kb downstream of the IKZF1 gene. Patients identified by the published direct genomic PCR approach are indicated as non-numbered “patients” (n=16).
IKAROS is part of a complex regulatory network involving the hematopoietic transcription factors E2A, EBF(1-3) and PAX5 to steer lymphocyte development. A homozygous knock-out of the murine Ikaros gene was associated with a complete loss of B, NK and T cells in the transgenic model [8]. In addition, IKZF1 belongs to of a gene family (IKZF1=IKAROS, IKZF2=HELIOS and IKZF3=AIOLOS), whose members share similar gene structures, exhibit a number of splice variants and are all involved in lymphocytic lineage decisions (see Figure 1B) [5]. Noteworthy, mutated members of this gene family are frequently identified in different hematological cancers (IKZF1*: BCP-ALL, BCR-ABL1, BCR-ABL1-like; IKZF2*: T-ALL; IKZF3*: BCP-ALL).
The first study demonstrating the importance of IKZF1 deletions in leukemia was published in 2007. Genome wide SNP array analyses of 242 patients revealed that several genes involved in B-lineage development can be mutated in BCP-ALL patients [9]. These efforts identified - besides PAX5, E2A, EBF1, LEF1 - the first IKZF1 and IKZF3 deletions. These initial findings have been validated by a second study that investigated 40 leukemia patients [10]. The study identified again recurrent submicroscopic deletions in several genes linked to B lineage development (PAX5, EBF1, TCF4, IKZF1 and others). Blocking key proteins required for proper B cell development will result in more immature cells which consequently will increase the risk for poorer outcome. Exactly this has been noted in clinical studies where either a BCR-ABL1-like subgroup with poor clinical course was defined by microarray transcription studies [11], or, where it was demonstrated that leukemia patients with IKZF1 deletions have a poorer clinical outcome [12]. Noteworthy, about 80% of all investigated BCR-ABL1+ ALL patients displayed an IKZF1 deletions [13], and leukemia patients bearing an IKZF1 deletion are all classified as high-risk ALL patients. The clinical effects of IKZF1 deletions and the results from basic research have been thoroughly reviewed [14].
Iacobucci and coworkers investigated a series of 106 patients with BCR-ABL1+ ALL [15]. Of those, 80 patients were positive for an IKZF1 deletion as detected by high-resolution SNP arrays. By using different genomic PCR strategies, 63 out of these 80 patients could be successfully analyzed at the genomic level. Interestingly, 44 patients (of which 32 could be characterized at the nucleotide level) displayed a Δ4-7 deletion that results in the expression of a dominant-negative IKAROS variant. The other 19 patients (of which 18 could be characterized at the nucleotide level) displayed a Δ2-7 IKZF1 deletion. This kind of deletion will result in an IKAROS protein that is composed of 13 amino acids of the first exon and 237 amino acids encoded by exon 8. The function of this protein variant is unknown, as it is not represented in the naturally occuring 11 splice variations.
The aim of our study was to extend the current knowledge about genetic events affecting the IKZF1 gene and to further improve the routine analysis of IKZF1 deletion. We applied long-distance inverse PCR (LDI-PCR) and long distance PCR (LD-PCR), that have been successfully used to clone genomic fusion sites in many acute leukemia patients [16,17]. We investigated a cohort of 72 pediatric BCP-ALL patients that was positively screened for a genetic rearrangement of IKZF1. Since the applied LDI-PCR method is completely unbiased, we were also able to identify Δ2-3, Δ2-8, Δ4-7 and Δ4-8 deletione of IKZF1 and a first tail to tail fusion of IKZF1 with COBL due to an interstitial deletion at 7p. Based on our data, we refined the current knowledge on submicroscopic deletions of the IKZF1 gene which in turn will enable other investigators to more precisely map IKZF1 deletions in the future.
Material and methods
Patient material
Seventy-two pediatric BCP-ALL patient samples with a deletion in IKZF1 as assessed by MLPA analysis were included in this study. All patients with copy number alterations of greater than 2 IKZF1 exons were classified as IKZF1-deleted patients. Fifty-six patients younger than 18 years with newly diagnosed ALL were registered in the German CoALL study, and sixteen Brazilian children with BCP-ALL were enrolled in the Immunophenotyping-Genotyping Study of Childhood Acute Leukemia [18]. Informed consent was obtained from all patients or patients’ parents/legal guardians and control individuals.
MLPA analyses to detect IKZF1 deletions
Deletions within the IKZF1 gene were analyzed by using multiplex ligation-dependent probe amplification (SALSA MLPA P202 probemix and/or SALSA MLPA P335-A3/4, MRC Holland, Amsterdam, The Netherlands). All analyses were performed according to the manufacturer’s recommendations.
Establishing an LDI-PCR method for the detection of genomic IKZF1 deletions
LDI-PCR experiments were in principle performed as described recently [19], with the following modification: genomic patient DNA samples were digested either with the restriction endonuclease BamHI/BglII (BB), AvrII/NheI/SpeI (ANS) or Acc65I/BsrGI (AB) before DNA religation reactions were carried out. After PCR analyses, all non-germline PCR fragments were cut out from the gel and subjected to DNA sequence analysis. Obtained sequence data were blasted against the human genome (NCBI database) and aligned to germline gene sequences. All primer used to perform LDI-PCR or multiplex PCR experiments are listed in Supplementary Table S1A-C.
Results
MLPA analyses revealed different groups of IKZF1 deletion patients
MLPA analyses were performed in a cohort of about 356 pediatric BCP-ALL patients. Several of the patients were selected because they had a relapse, and thus, our cohort was presumably biased towards a poorer outcome. In 72 patients (20%), copy number aberrations of IKZF1 exons were identified. The outline of our study is schematically depicted in Figure 2. According to the obtained MLPA data, several patient groups can be defined. A first group exhibited only IKZF1 Δ2-7 or Δ4-7 deletions. This group (n=16) was analyzed by LD-PCR with the already established primer sets according to Iacobucci at al 2009 [15]. A second group comprised 40 MLPA+ leukemia patients that were classified as IKZF1 Δ1-3 (n=1), Δ1-7 (n=1), Δ1-8 (n=16), Δ2-3 (n=6), Δ2-7 (n=3), Δ2-8 (n=4), Δ3-7 (n=1), Δ4-7 (n=4) and Δ4-8 (n=4). This group was investigated by LDI-PCR. A third group came from Brazil (n=16) and contained leukemia patients with IKZF1 Δ1-8 (n=6), Δ4-7 (n=3), Δ4-8 (n=2), Δ2,3,6 (n=1), Δ6 (n=3) and Δ8 (n=1). This group was also analyzed by LDI-PCR to investigate the unbiased incidence in a test cohort.
Figure 2.

Design of the study. A total of 72 patients were identified as “IKZF1-deleted” (more than 2 exons of IKZF1 were lost). Some patients have been classified by MLPA analyses as “Δ6” (n=3) and “Δ8” (n=1); these patients have been included in our study. The patients with IKZF1 Δ2-7 and Δ4-7 were directly analyzed by available PCR tests (n=16). All others (n=40) and the 16 patients from Brazil were directly subjected to LDI-PCR analyses. The obtained information was translated into the establishment of a multiplex PCR method which allows for identification of IKZF1-deleted patients by two independent PCR reaction (MP-PCR A and B). The novel method was validated in a cohort of 88 unscreened pediatric ALL patients. These analyses revealed 8 positive patients.
Fine mapping of internally deleted IKZF1 genes
All 16 patients of MLPA+-group 1 exhibited the classical IKZF1 Δ2-7 and Δ4-7 deletions which were successfully analyzed by applying the published LD-PCR strategy [15]. All other patients of MLPA+-group 2 and 3 were characterized as follows: two patients bear an IKZF1 allele that resulted from recombination event between IKZF1 intron1 (see Figure 1D, Hot Spot 1; HS1) and intron 7 (Hot Spot 3; HS3). Fourteen patients bear an IKZF1 allele that resulted from a recombination event between IKZF1 intron 3b (Hot Spot 2; HS2) and intron 7 (HS3). All breakpoints are indicated as red (5’-break) and green numbers (3’-break) below the IKZF1 gene structure (Figure 1D), while the primary DNA sequence of all genomic breakpoints are listed in Supplementary Table S2.
LDI-PCR experiments provide the possibility to identify any kind of genetic rearrangement (small deletions, inversions or duplications; chromosomal translocations, larger interstitial deletions, etc). Thus, we conducted LDI-PCR experiments as outlined in material and methods to clone the recombination sites in different IKZF1-deleted patients. However, LDI-PCR analyses of IKZF1-deleted patients turned out to be rather complicated. As summarized in Figure 1C and in Figure 2, only 20 out of 40 MLPA+ patients of group 2 and only 4 out of 16 MLPA+ patients of group 3 were successfully analyzed. One reason for the low success rate was the high number of patients that carried a deletion of all IKZF1 exons (Δ1-8 patients; n=22). In these cases neither LDI-PCR nor any other type of PCR technique can be applied because the chromosomal breakpoints are located far outside of the IKZF1 gene.
During our studies, we identified a fourth recombination hotspot (HS4) that is located about 11 kb downstream of the IKZF1 polyA site. This novel HS4 represents a 3’-break site which was identified in 11 patients (1, 5, 6, 7, 9, 10, 32, 45, 47a, 65a and 123). This novel finding gave us the opportunity to perform additional LDI-PCR experiments from this downstream site, but all 22 patients which were negative in the first round of screening, remained also negative in the second round of analyses. Thus, the larger chromosomal deletions present in these patients hampered any kind of reasonable analysis.
The mapped deletion sites of 24 patients belonging to group 2 and 3 (displayed in Figure 1D) revealed that not all patients have their breakpoints in vicinity of the already published hotspots (HS1-3). 5’-breaks within IKZF1 intron 1 (HS1) are not only limited to recombinations at HS1 but are rather clustering over a larger genomic region (n=16) or were even located far upstream (~120 kb; n=1). Similar findings were obtained for 5’-breaks within IKZF1 intron 3b (HS2), where most patients exhibited their gene-internal recombination events (n=33). Nearly all 3’-breaks clustered at HS3 (n=28) located within IKZF1 intron 7, but we also identified a breakpoint upstream of HS3 (n=1), one in IKZF1 exon 8 (n=1), and another one in the 3’-UTR (n=1). As already mentioned above, a large series of patients exhibited their 3’-breaks in a tiny region located about 11.6 kb downstream of the poly(A) site (n=11). This recombination hotspot is responsible for all characterized IKZF1 Δ4-8 deletions, resulting in a protein variant that is per se not capable of forming heterodimers with other bona fide IKAROS protein variants. Thus, these genetic events can be regarded as functional knock-out of one IKZF1 allele. Interestingly, about 120 and 240 bp downstream of the novel 11 kb downstream recombination site, two putative 3’-splice acceptor sites were identified (+123: TCCCTCCTGCCAG; +243: TCATCCTCCTCTCAG), together with a putative bona fide poly-A site (+350: AATAAA). Therefore, transcripts encoded by these mutant IKZF1 genes contain exon 1 and exon 2 sequences fused to the 11 kb downstream region, where the putative signal sequences may allow proper post-transcriptional processing, which is a prerequisite for nuclear export of such mRNAs and their translation in the cytosol.
In addition, we identified a yet undescribed interstitial deletion that involved the IKZF1 and the COBL genes located at 7p12 and 7p12.1, respectively. This larger deletion has a size of ~800 kb and fused IKZF1 intron 1 with COBL intron 5 in a tail-to-tail fashion. The resulting gene fusion exhibit two promoter regions which may allow transcription of the first non-coding exon of IKZF1 and the first 5 COBL exons (the first 5 exons encode 261 out of 1,261 amino acids), but none of these transcripts will result in a functional protein. The interstitial deletion also led to a deletion of 3 additional genes (FIGNL1, DCC, GRB10). The FIGNL1 protein is assumed to play a role in osteoblast proliferation, while the DCC protein represents a tumor suppressor protein that is frequently deleted in colon cancer cells. GRB10 is a SH2-domain protein that is involved in the insuline/IGF1 signaling pathway, and is linking the IGF1 with the RAS/RAF signaling pathway.
Analysis of RAG1/2 recognition signal sequences in recombined IKZF1 alleles
We also investigated the hypothesis that most of these illegitimate recombination events were presumably caused by RAG1/RAG2 mediated recombination. Therefore we aimed to identify potential RAG1/RAG2 recognition signal sequences (RSS) that flank the recombination sites. Per definition, these sites should be located within the deleted region. As shown in the Supplementary Table S2, sequence homologous to the known RAG1/RAG2 RSS (CACAGTG-12/23-ACAAAACC or GGTTTTGT-23/12-CACTGTG) has been identified for all characterized fusion alleles of the investigated patients near to the sequenced recombination sites. Based on these analyses, we validated the hypothesis that mutant forms of the IKZF1 gene, identified in leukemia patients, lost one of their IKZF1 allele due to an illegitimate, RAG1/RAG2-mediated recombination process. Moreover, the four hotspots are presumably exhibiting the best consensus sites for these RAG1/RAG2-mediated illegitimate recombination events.
Establishing an Multiplex-PCR method for the detection of genomic IKZF1 deletions
Based on the data obtained in this study, a novel diagnostic PCR assay was established to simplify the diagnosis of genomic IKZF1 deletions. By using 2 different multiplex PCR reactions (A and B) we were able to demonstrate the presence of different IKZF1 deletions (Δ2-3, Δ2-8, Δ4-7 and Δ4-8). Unfortunately, larger deletions affecting the whole IKZF1 gene or deletions that comprise exon 1 of IKZF1 (IKZF1 Δ1-7, Δ1-8) cannot be detected by this method. As shown in Figure 3, the A-reaction mix of our established multiplex PCR was able to identify the genomic fusion in 4 out of 4 patients (Pts #13, #14, #15 and #17) that we characterized to carry an IKZF1 Δ2-3 deletion. The B-reaction mix of our established multiplex PCR was able to identify the genomic fusion in 11 out of 12 patients. Three out of 4 patients with IKZF1 Δ2-8 deletions were positively identified. Patient 3 must be negative in this analysis because this patient exhibits the above described tail-to-tail fusion with the COBL gene (see above). All three patients selected for the IKZF1 Δ4-7 fusion and 5 out of 5 patients that carried the IKZF1 Δ4-8 deletion were also positively identified. Similarly, patient #4 in the panel Δ4-8 was used as a negative control, because this patient carries a complete IKZF1 gene deletion.
Figure 3.

PCR results of investigated IKZF1 deleted patients. Results of the multiplex PCR in different patient subgroups are displayed. Left: multiplex PCR reaction A is able to readily identify IKZF1 Δ2-3 leukemia patients. Middle: multiplex PCR reaction B was used to characterize IKZF1 Δ2-8, IKZF1 Δ4-1 and IKZF1 Δ4-8 leukemia patients. Right: two examples of our unbiased screening of 88 BCP-ALL patients revealed two patients with both IKZF1 alleles partially deleted: both patients (Pts #47 and #65) displayed an IKZF1 Δ4-7 and an IKZF1 Δ4-8 deletion. Lane 1 in panel “Δ4-8” was a negative control (Pt #4). Patient #4 was defined by MLPA analysis to carry an IKZF1 Δ1-8 deletion, and thus, exhibits a larger deletion of chromosome 7. Such patients can neither be positively analyzed by MP-PCR nor by LDI-PCR. Lane 3 in panel “Δ2-8” represents patient #3 that was identified to carry the IKZF1-COBL fusion. Therefore, this patient fails to show any amplification by MP-PCR.
After having established this novel diagnostic method, we validated the MP-PCR method on a cohort of 88 unselected pediatric ALL patients (see Figure 2). Within this validation group, we were able to identify 8 additional patients to carry an IKZF1 deletion. Surprisingly, 2 of these 8 patients had both alleles of IKZF1 deleted (see Figure 3, right panel: Pts 47 and 65). Both displayed an HS2-HS3 and HS2-HS4 recombination event. The other 6 patients displayed IKZF1 Δ 2-3 (n=2; HS1-HS2), Δ 4-7 (n=3; HS2-HS3) and Δ4-8 (n=1; HS2-HS4) deletions. These experiments successfully validated our novel multiplex PCR method.
Discussion
Using a combined experimental approach (MLPA screening in combination with LDI- or LD-PCR) we attempted to assess IKZF1 deletions in pediatric BCP-ALL patients. By applying these technologies, we extended the current knowledge of recombination hotspot 1 (HS1) and 2 (HS2), because we obtained a more scattered pattern of recombination events within IKZF1 intron 1 and intron 3b (see Figure 2D). We also confirmed the existence of HS3 and HS4, of which the latter one was recently identified to be located about 11.6 kb downstream of the IKZF1 gene [20]. Recombination events involving HS4 were mainly found in patients displaying IKZF1 Δ2-8 and Δ4-8 deletions. These genetic hotspots are presumably caused by the action of RAG1/RAG2 heterodimers that are normally used in lymphoid precursor cells to recombine Immunoglobuline and T-cell receptor genes. This assumption is strongly supported by DNA sequence motifs that are flanking the deletion site and which are highly similar to the known RAG1/RAG2 recognitions signal sequences (see Supplementary Table S2).
Based on these extended findings we developed a 2-tube multiplex PCR assay (tube A and B), able to readily identify intragenic IKZF1 deletions. This method was validated in the cohort of 88 pediatric BCP-ALL patients. This unselected cohort revealed about 9% positive cases. According to our data, about half of IKZF1 deleted leukemia patients can be diagnosed by the 2-tube multiplex PCR method, while the remaining 50% of patients carry larger deletions at chromosome 7p that could neither be analyzed by the established MLPA (all negative) nor by any other type of PCR analysis. These patients can only be identified by more costly CNA analyses.
IKZF1 deletions are frequently observed in BCR-ABL1 positive ALL patients (up to 80%), in BCR-ABL-like leukemia patients and in BCP-ALL patients [9,13,14]. Since IKZF1-deletions have a prognostic impact on the outcome of patients, the yet established MLPA test can be combined with any available PCR system to generate a rapid diagnostic read-out to identify patients. Sequencing the obtained PCR amplimers allows to establish a patient-specific biomarker which might be useful for subsequent MRD monitoring. According to our experience with genomic fusion sites as MRD markers, the sensitivity of these assays allow to trace residual tumor cells in a concentration between 10-4 and 10-5, depending on the available primary DNA sequences. This could be important for those patients for which alternative MRD markers are not available or lost during therapy. Two recent studies have already demonstrated that it is a feasible strategy when applied to leukemia patients that bear IKZF1 deletions [20,21]. Thus, the analysis of IKZF1 deletions may not only be of scientific interest but may also be of benefit for the molecular surveillance of therapeutic treatment response.
Acknowledgements
This work is supported by grant DKS 2011.09 from the German Pediatric Cancer Foundation to R.M. MSPO and ME were supported by CAPES-PROBRAL (#345/10) and CNPq (#309091/2007-1).
Supporting Information
References
- 1.Moorman AV. The clinical relevance of chromosomal and genomic abnormalities in B-cell precursor acute lymphoblastic leukemia. Blood Rev. 2012;26:123–135. doi: 10.1016/j.blre.2012.01.001. [DOI] [PubMed] [Google Scholar]
- 2.Georgopoulos K, Moore DD, Derfler B. Ikaros, an early lymphoid-specific transcription factor and a putative mediator for T cell commitment. Science. 1992;258:808–812. doi: 10.1126/science.1439790. [DOI] [PubMed] [Google Scholar]
- 3.Molnár A, Wu P, Largespada DA, Vortkamp A, Scherer S, Copeland NG, Jenkins NA, Bruns G, Georgopoulos K. The Ikaros gene encodes a family of lymphocyte-restricted zinc finger DNA binding proteins, highly conserved in human and mouse. J Immunol. 1996;156:585–592. [PubMed] [Google Scholar]
- 4.Klug CA, Morrison SJ, Masek M, Hahm K, Smale ST, Weissman IL. Hematopoietic stem cells and lymphoid progenitors express different Ikaros isoforms, and Ikaros is localized to heterochromatin in immature lymphocytes. Proc Natl Acad Sci U S A. 1998;95:657–662. doi: 10.1073/pnas.95.2.657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rebollo A, Schmitt C. Ikaros, Aiolos and Helios: transcription regulators and lymphoid malignancies. Immunol Cell Biol. 2003;81:171–175. doi: 10.1046/j.1440-1711.2003.01159.x. [DOI] [PubMed] [Google Scholar]
- 6.Liberg D, Smale ST, Merkenschlager M. Upstream of Ikaros. Trends Immunol. 2003;24:567–570. doi: 10.1016/j.it.2003.09.008. [DOI] [PubMed] [Google Scholar]
- 7.Chrousos GP, Kino T. Ikaros transcription factors: flying between stress and inflammation. J Clin Invest. 2005;115:844–848. doi: 10.1172/JCI24886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Georgopoulos K, Bigby M, Wang JH, Molnar A, Wu P, Winandy S, Sharpe A. The Ikaros gene is required for the development of all lymphoid lineages. Cell. 1994;79:143–156. doi: 10.1016/0092-8674(94)90407-3. [DOI] [PubMed] [Google Scholar]
- 9.Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, Girtman K, Mathew S, Ma J, Pounds SB, Su X, Pui CH, Relling MV, Evans WE, Shurtleff SA, Downing JR. Genome-wide analysis of genetic alterations in acute lymphoblastic leukemia. Nature. 2007;446:758–764. doi: 10.1038/nature05690. [DOI] [PubMed] [Google Scholar]
- 10.Kuiper RP, Schoenmakers EF, van Reijmersdal SV, Hehir-Kwa JY, van Kessel AG, van Leeuwen FN, Hoogerbrugge PM. High-resolution genomic profiling of childhood ALL reveals novel recurrent genetic lesions affecting pathways involved in lymphocyte differentiation and cell cycle progression. Leukemia. 2007;21:1258–1266. doi: 10.1038/sj.leu.2404691. [DOI] [PubMed] [Google Scholar]
- 11.Den Boer ML, van Slegtenhorst M, De Menezes RX, Cheok MH, Buijs-Gladdines JG, Peters ST, Van Zutven LJ, Beverloo HB, Van der Spek PJ, Escherich G, Horstmann MA, Janka-Schaub GE, Kamps WA, Evans WE, Pieters R. A subtype of childhood acute lymphoblastic leukemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol. 2009;10:125–134. doi: 10.1016/S1470-2045(08)70339-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Martinelli G, Iacobucci I, Storlazzi CT, Vignetti M, Paoloni F, Cilloni D, Soverini S, Vitale A, Chiaretti S, Cimino G, Papayannidis C, Paolini S, Elia L, Fazi P, Meloni G, Amadori S, Saglio G, Pane F, Baccarani M, Foà R. IKZF1 (Ikaros) deletions in BCR-ABL1-positive acute lymphoblastic leukemia are associated with short disease-free survival and high rate of cumulative incidence of relapse: a GIMEMA AL WP report. J. Clin. Oncol. 2009;27:5202–5207. doi: 10.1200/JCO.2008.21.6408. [DOI] [PubMed] [Google Scholar]
- 13.Mullighan CG, Miller CB, Radtke I, Phillips LA, Dalton J, Ma J, White D, Hughes TP, Le Beau MM, Pui CH, Relling MV, Shurtleff SA, Downing JR. BCR-ABL1 lymphoblastic leukemia is characterized by the deletion of Ikaros. Nature. 2008;453:110–114. doi: 10.1038/nature06866. [DOI] [PubMed] [Google Scholar]
- 14.Roberts KG, Mullighan CG. How new advances in genetic analysis are influencing the understanding and treatment of childhood acute leukemia. Curr Opin Pediatr. 2011;23:34–40. doi: 10.1097/MOP.0b013e3283426260. [DOI] [PubMed] [Google Scholar]
- 15.Iacobucci I, Storlazzi CT, Cilloni D, Lonetti A, Ottaviani E, Soverini S, Astolfi A, Chiaretti S, Vitale A, Messa F, Impera L, Baldazzi C, D’Addabbo P, Papayannidis C, Lonoce A, Colarossi S, Vignetti M, Piccaluga PP, Paolini S, Russo D, Pane F, Saglio G, Baccarani M, Foà R, Martinelli G. Identification and molecular characterization of recurrent genomic deletions on 7p12 in the IKZF1 gene in a large cohort of BCR-ABL1-positive acute lymphoblastic leukemia patients: on behalf of Gruppo Italiano Malattie Ematologiche dell’Adulto Acute Leukemia Working Party (GIMEMA AL WP) Blood. 2009;114:2159–2167. doi: 10.1182/blood-2008-08-173963. [DOI] [PubMed] [Google Scholar]
- 16.Meyer C, Schneider B, Jakob S, Strehl S, Attarbaschi A, Schnittger S, Schoch C, Jansen MW, van Dongen JJ, den Boer ML, Pieters R, Ennas MG, Angelucci E, Koehl U, Greil J, Griesinger F, Zur Stadt U, Eckert C, Szczepanski T, Niggli FK, Schäfer BW, Kempski H, Brady HJ, Zuna J, Trka J, Nigro LL, Biondi A, Delabesse E, Macintyre E, Stanulla M, Schrappe M, Haas OA, Burmeister T, Dingermann T, Klingebiel T, Marschalek R. The MLL recombinome of acute leukemias. Leukemia. 2006;20:777–784. doi: 10.1038/sj.leu.2404150. [DOI] [PubMed] [Google Scholar]
- 17.Meyer C, Kowarz E, Hofmann J, Renneville A, Zuna J, Trka J, Ben Abdelali R, Macintyre E, De Braekeleer E, De Braekeleer M, Delabesse E, de Oliveira MP, Cavé H, Clappier E, van Dongen JJ, Balgobind BV, van den Heuvel-Eibrink MM, Beverloo HB, Panzer-Grümayer R, Teigler-Schlegel A, Harbott J, Kjeldsen E, Schnittger S, Koehl U, Gruhn B, Heidenreich O, Chan LC, Yip SF, Krzywinski M, Eckert C, Möricke A, Schrappe M, Alonso CN, Schäfer BW, Krauter J, Lee DA, Zur Stadt U, Te Kronnie G, Sutton R, Izraeli S, Trakhtenbrot L, Lo Nigro L, Tsaur G, Fechina L, Szczepanski T, Strehl S, Ilencikova D, Molkentin M, Burmeister T, Dingermann T, Klingebiel T, Marschalek R. New insights to the MLL recombinome of acute leukemias. Leukemia. 2009;23:1490–1499. doi: 10.1038/leu.2009.33. [DOI] [PubMed] [Google Scholar]
- 18.Pombo de Oliveira MS, Koifman S, Vasconcelos GM, Emerenciano M, de Oliveira Novaes C Brazilian Collaborative Study Group of Infant Acute Leukemia. Development and perspective of current Brazilian studies on the epidemiology of childhood leukemia. Blood Cells Mol Dis. 2009;42:121–125. doi: 10.1016/j.bcmd.2008.10.012. [DOI] [PubMed] [Google Scholar]
- 19.Meyer C, Schneider B, Reichel M, Angermueller S, Strehl S, Schnittger S, Schoch C, Jansen MW, van Dongen JJ, Pieters R, Haas OA, Dingermann T, Klingebiel T, Marschalek R. A new diagnostic tool for the identification of MLL rearrangements including unknown partner genes. Proc Natl Acad Sci U S A. 2005;102:449–454. doi: 10.1073/pnas.0406994102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Caye A, Beldjord K, Mass-Malo K, Drunat S, Soulier J, Gandemer V, Baruchel A, Bertrand Y, Cavé H, Clappier E. Breakpoint-specific multiplex PCR allows the detection of IKZF1 intragenic deletions and minimal residual disease monitoring in B-cell precursor acute lymphoblastic leukemia. Haematologica. 2013;98:597–601. doi: 10.3324/haematol.2012.073965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Venn NC, van der Velden VH, de Bie M, Waanders E, Giles JE, Law T, Kuiper RP, de Haas V, Mullighan CG, Haber M, Marshall GM, Md N, van Dongen JJ, Sutton R. Highly sensitive MRD tests for ALL based on the IKZF1 Δ3-6 microdeletion. Leukemia. 2012;26:1414–1416. doi: 10.1038/leu.2011.348. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


