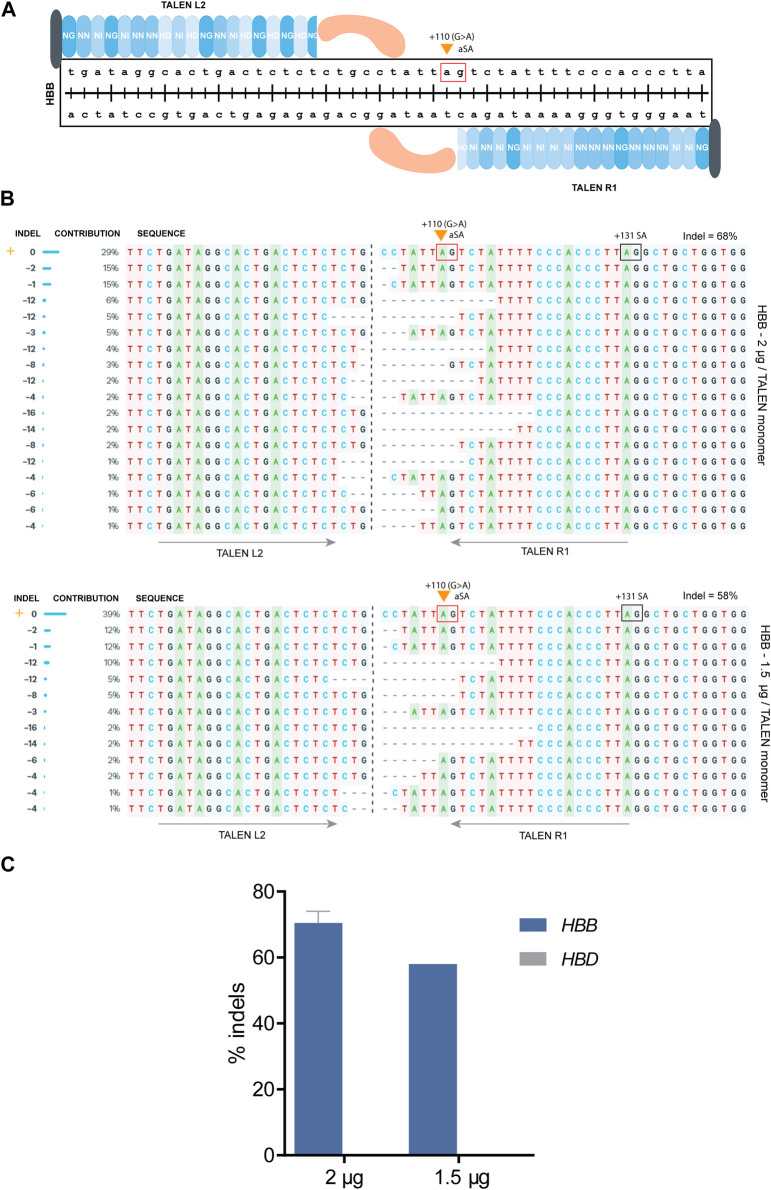
FIGURE 4.
TALEN-based targeting of the HBB IVSI−110(G>A) mutation in patient-derived CD34+ cells. (A) A schematic diagram showing the design of TALENs targeting the HBB IVSI−110(G>A) mutation (orange arrow). The red box shows the aSA (AG dinucleotide) that is created by the mutation. The TALEN monomers L2 and R1 were used to introduce DSBs and create indels upstream of the splice acceptor site. The blue shaded repeated shapes represent the specific RVDs used for the design, the black shape shows the N-terminus, while the orange shape represents the Fok1 endonuclease monomer. (B) Alignment of the various editing events resulting from the disruption of the upstream splice acceptor site using the TALENs R1/L2 pair. For the top alignment 2 μg, for the bottom 1.5 μg were used per TALEN monomer. The INDEL column shows the exact number of deleted (<0) nucleotides, the CONTRIBUTION column the percentage of these indels in the bulk population, and the SEQUENCE column the edited sequences containing the various indels aligned with the wild type (orange cross). The orange arrow indicates the HBB IVSI−110(G>A) mutation as the first base of the aSA (red box); the normal +131 splice acceptor site (SA) is indicated by a black box. The grey arrows show the binding site of each TALEN monomer. The percentage of the total indels is also displayed. (C) Chart showing the percentage of indels detected for HBB and HBD after delivery of either 2 or 1.5 μg per TALEN monomer. For the 2 μg/TALEN monomer, the data are plotted as mean ± s.d. (n = 2.)