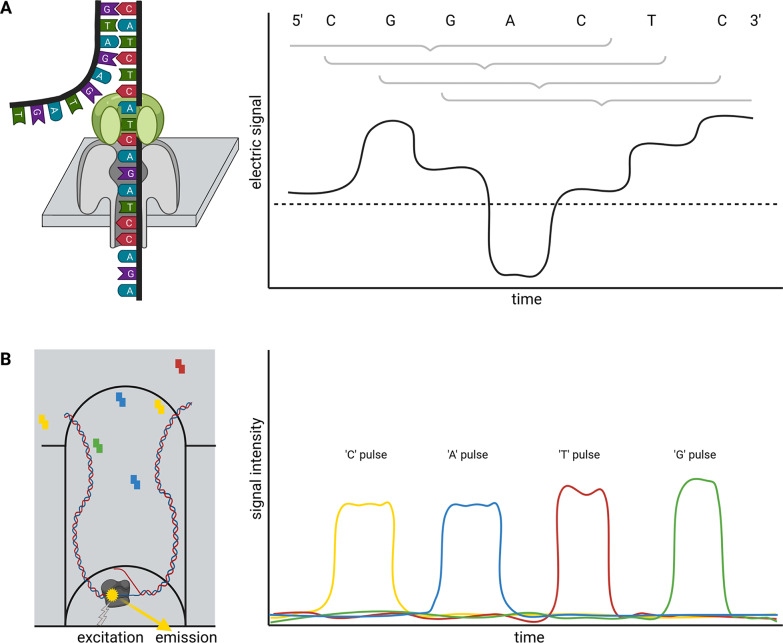
Fig. 1.
Schematic illustration of nanopore sequencing (a) and Single-Molecule Real Time (SMRT) sequencing (b). Nanopore sequencing is based on the translocation of a DNA or RNA strand through a nanopore located in an artificial membrane. Multiple nucleotides located in the nanopore determine the flow of ions through this nanopore in a specific way by physically blocking the space. This change in ion flux is recorded as an electric signal and further converted into sequence information. The illustration shows the contribution of six bases to the signal, but the number of bases depends on the pore type. SMRT sequencing detects fluorescent light emitted from nucleotides upon incorporation into a DNA strand. The DNA polymerase is located at the bottom of a well and synthesises a new DNA strand. The integration into the new DNA strand keeps the nucleotide for a sufficiently long time in the well to allow detection.