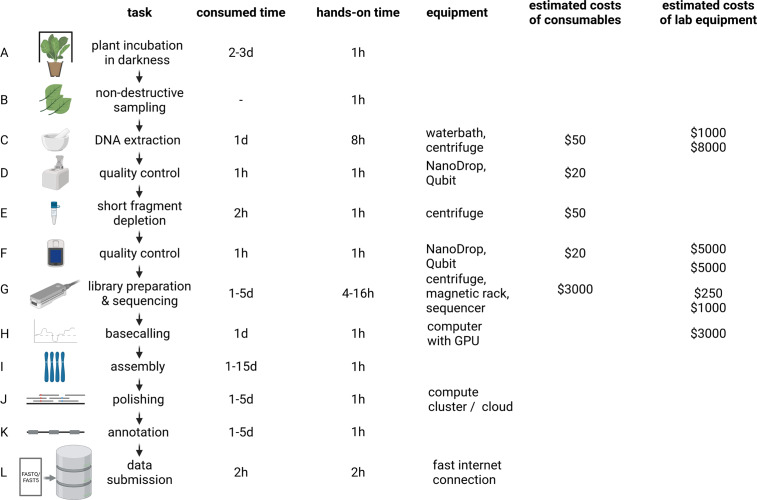
Fig. 2.
Plant genome project workflow from DNA extraction over Oxford Nanopore Technologies (ONT) sequencing to data submission. The indicated durations depend on the size and complexity of the investigated plant genome, with larger genomes generally taking longer to analyse. To reduce sugar content, plants are incubated in the dark for a few days prior to DNA extraction (a). Non-destructive sampling is important to allow additional genomic sequencing and also RNA-Seq if required in later stages of a project (b). Mechanical disruption of cell walls is required for the DNA extraction (c). Photometric analysis of the DNA solution (including quantification) is often the first step of quality control (d and f). Removal of short DNA fragments is highly recommended to improve the sequencing output and quality (e). ONT library preparation and sequencing can be repeated several times to increase the output (g). Graphic cards are an efficient resource to convert electric signal into sequence information in real time (h). Multiple tools are available to generate a chromosome-arm level assembly based on long reads (i). Additional polishing in multiple rounds can be necessary due to the noisy character of long reads (j). The value of a genome sequence can be enriched through the identification of relevant genetic elements like genes and transposable elements (k). All data should be shared with the community via submission to a public repository which ensures long-term storage (l). d, day(s); hr, hour(s). The given time estimates for assembly, polishing and annotation are the minimal run time required for the analyses. Manual curation and iterative improvements can take substantially longer. The estimated costs of consumables are based on a haploid 1-Gbp genome and a targeted coverage of 30× which would require six libraries to be sequenced on three MinION/GridION flow cells when assuming an average output of 10 GB per flow cell with two libraries sequenced per flow cell. Investment costs for non-standard lab equipment are independent of the specific sequencing project and only required for high-output experiments in the lab. There is an option to perform rapid sequencing without these instruments in the field, but the lower output does not make that option attractive for large plant genomes.