Abstract
Cell divisions are accurately positioned to generate cells of the correct size and shape. In plant cells, the new cell wall is built in the middle of the cell by vesicles trafficked along an antiparallel microtubule and a microfilament array called the phragmoplast. The phragmoplast expands toward a specific location at the cell cortex called the division site, but how it accurately reaches the division site is unclear. We observed microtubule arrays that accumulate at the cell cortex during the telophase transition in maize (Zea mays) leaf epidermal cells. Before the phragmoplast reaches the cell cortex, these cortical-telophase microtubules transiently interact with the division site. Increased microtubule plus end capture and pausing occur when microtubules contact the division site-localized protein TANGLED1 or other closely associated proteins. Microtubule capture and pausing align the cortical microtubules perpendicular to the division site during telophase. Once the phragmoplast reaches the cell cortex, cortical-telophase microtubules are incorporated into the phragmoplast primarily by parallel bundling. The addition of microtubules into the phragmoplast promotes fine-tuning of the positioning at the division site. Our hypothesis is that division site-localized proteins such as TANGLED1 organize cortical microtubules during telophase to mediate phragmoplast positioning at the final division plane.
Microtubules accumulate at the cell cortex during telophase and interact with the plant division machinery to direct its movement towards the division site.
IN A NUTSHELL.
Background: Both cell division and proper orientation of the division are important for plant development and growth. Cell division is initiated in the middle of the cell by a structure called the phragmoplast. The phragmoplast is composed of filaments including microtubules and it expands outward to form the new cell wall. Phragmoplast positioning is mediated by proteins that localize in a ring at the future division location or division site. It is not known how these proteins promote division positioning.
Question: How do proteins at the division site contribute to the phragmoplast reaching the correct location?
Findings: We propose a potential mechanism linking phragmoplast positioning with division site-localized proteins using maize epidermal cells expressing a live-cell microtubule marker. We discovered that an extensive network of microtubule filaments accumulates at the cell periphery and is captured at the division site by a microtubule-binding protein called TANGLED1, thereby leading to microtubules that are oriented perpendicular to the division site. Pre-oriented microtubules are added into the phragmoplast as it reaches the cell periphery to accurately direct the movement of the phragmoplast.
Next steps: Whether microtubules participate in division positioning and how TANGLED1 might modulate their dynamics in other plant cells are unknown.
Introduction
Cell division in plants occurs via the transport of vesicles along an antiparallel microtubule array called the phragmoplast to build a new cell wall (Smertenko et al. 2017). The phragmoplast grows toward the cell cortex via microtubule nucleation on preexisting phragmoplast microtubules. Microtubule-dependent microtubule nucleation on the phragmoplast is mediated by gamma-tubulin and augmin complex proteins (Hotta et al. 2012; Nakaoka et al. 2012; Murata et al. 2013; Lee et al. 2017; Lee and Liu 2019). Microtubule bundling in the phragmoplast may be mediated by MICROTUBULE ORGANIZATION 1 (MOR1)/TMBP200/GEM1, which localizes to the phragmoplast and also crosslinks microtubules in vitro (Yasuhara et al. 2002; Hamada et al. 2004). This activity is consistent with its role in rapid phragmoplast expansion, as demonstrated by quantitative live-cell imaging (Kawamura et al. 2006). In addition to nucleation and bundling, microtubule disassembly at the phragmoplast lagging edge also promotes phragmoplast expansion. Mitogen-activated protein kinase (MAPK) phosphorylates MAP65-1, which, in turn, reduces the microtubule bundling efficiency of MAP65-1. MAP65-1 phosphorylation allows lagging edge microtubules to disassemble, thereby promoting phragmoplast expansion (Sasabe et al. 2006; Sasabe and Machida 2012). MAP65-1 is also phosphorylated by alpha Aurora Kinase, which similarly promotes timely phragmoplast expansion (Boruc et al. 2016). PHRAGMOPLAST ORIENTING KINESIN2 (POK2) promotes timely phragmoplast expansion, possibly through interaction with the phragmoplast midzone crosslinker MAP65-3 (Herrmann et al. 2018).
Although there has been progress in identifying factors that mediate phragmoplast expansion, how the phragmoplast is directed toward a specific cortical location, called the division site, is still unknown (Rasmussen and Bellinger 2018; Livanos and Müller 2019). In land plants, the location of the future division site can be accurately predicted by a microtubule and microfilament structure that assembles during the G2 phase of the cell cycle at the cell cortex called the preprophase band (PPB). Several proteins co-localize with the PPB and remain at the division site until the division is completed. These division site-localized proteins promote phragmoplast guidance to the division site: phragmoplasts in mutant cells often do not return to the division site (Cleary and Smith 1998; Müller et al. 2006; Martinez et al. 2017). Several division site-localized proteins are microtubule- or microfilament-bundling or motor proteins (Müller et al. 2006; Wu and Bezanilla 2014; Hermann et al. 2018; Martinez et al. 2020), suggesting that division site positioning may be mediated by local alterations in cytoskeletal dynamics.
Current models of division plane positioning propose that division site- and phragmoplast-localized proteins pull or push cytoskeletal filaments within the phragmoplast to guide it to the division site. More specifically, these models propose that microtubules attached to and nucleated from the phragmoplast (peripheral microtubules) interact with division site-localized proteins such as the PHRAGMOPLAST ORIENTING KINESIN2 (POK2) or MYOSINVIII (Wu and Bezanilla 2014; Chugh et al. 2018). A proposed function of POK2, a microtubule plus end-directed kinesin, is to bind peripheral microtubules and, through plus end-directed movement, push them away from the division site, thereby positioning the phragmoplast (Chugh et al. 2018). MYOSINVIII, a division site-localized protein that also localizes to the plus ends of peripheral phragmoplast microtubules, mediates an interaction between microtubules and actin filaments to guide the phragmoplast to the division site (Wu and Bezanilla 2014). The microtubule-binding protein TANGLED1 (TAN1) localizes to the division site in maize (Zea mays) and Arabidopsis (Arabidopsis thaliana) (Walker et al. 2007; Martinez et al. 2017). In vitro, TAN1 bundles and promotes transient microtubule capture and crosslinking. Furthermore, it co-localizes with a small population of phragmoplast microtubules at the division site in vivo and promotes timely phragmoplast expansion (Martinez et al. 2017, 2020). Maize tan1 mutants have defects in phragmoplast guidance to the division site, indicating that TAN1 contributes to positioning the phragmoplast, but the underlying mechanisms are not yet clear (Cleary and Smith 1998; Martinez et al. 2017).
The plant cell at telophase has often been considered devoid of microtubules outside of the phragmoplast (Smertenko et al. 2017). However, many reports indicate that both microtubules and microtubule nucleators such as gamma-tubulin are present at the cell cortex during telophase in monocots, dicots, ferns, and moss (Gunning et al. 1978; Wick 1985; Liu et al. 1995; Panteris et al. 1995; Kong et al. 2010; Wu and Bezanilla 2014). Additionally, in dicots, microtubules nucleated from the nuclear envelope in late telophase contact the cell cortex but their function was not described (Chan et al. 2005; Van Damme and Geelen 2008). More recently, cortical microtubules were shown to accumulate independently of the nuclear envelope in Arabidopsis during the late telophase (Lucas 2021). A previously proposed function of cortical-telophase microtubules is to prepopulate the cortex for microtubule reorganization during G1 (Flanders et al. 1990).
Here, we demonstrate that cortical microtubules are organized by a transient interaction with the division site, specifically near TAN1 puncta, in maize during telophase. This interaction may be directly mediated by the division site-localized protein TAN1. The cortical-telophase microtubules are prearranged via interactions with TAN1 or other nearby proteins at the division site. Cortical-telophase microtubules are then added by parallel bundling into the phragmoplast at the cortex. Therefore, cortical-telophase microtubules direct the movement of the phragmoplast toward the division site in maize cells.
Results
Analysis of cortical microtubule accumulation and orientation during telophase
Live-cell imaging of symmetrically dividing maize leaf epidermal cells revealed an unexpected population of cell-cortex-localized microtubules that typically accumulated during telophase (n = 45/49 cells) (Fig. 1, A and B; Supplemental Fig. S1, A and B). Cortical microtubules were sparse or nonexistent during metaphase (10% n = 2/20 cells from 8 plants, Supplemental Fig. S1, G and H) and anaphase (0% n = 0/8 cells from 8 plants). These cortical-telophase microtubules were spatially distinct from the phragmoplast and accumulated before the phragmoplast reached the cortex, as shown by time-lapse imaging (Fig. 1, A and B). We measured the density of cortical-telophase microtubules using the BoneJ plug-in in ImageJ (Doube et al. 2010), as described in the “Materials and methods” section. Using confocal z stacks with 0.5-µm resolution, we observed that the cortical-telophase array started to accumulate when the phragmoplast was as far as 3 to 4.5 microns away from the cell cortex (5% density, n = 2 cells). When the phragmoplast was between 0.5 and 1.5 microns from the cell cortex, the cortical-telophase array covered 33% of the cortex [n = 6 cells, standard error (SE) ± 7%]. The density of the cortical-telophase microtubules increased as the leading edge of the phragmoplast neared the cell cortex. The density of the cortical-telophase array when the phragmoplast touched the cortex was 39% (n = 24 cells, SE ± 4%, from 12 plants). Together, these results indicate that cortical-telophase microtubules accumulate during telophase but before the phragmoplast reaches the cortex.
Figure 1.
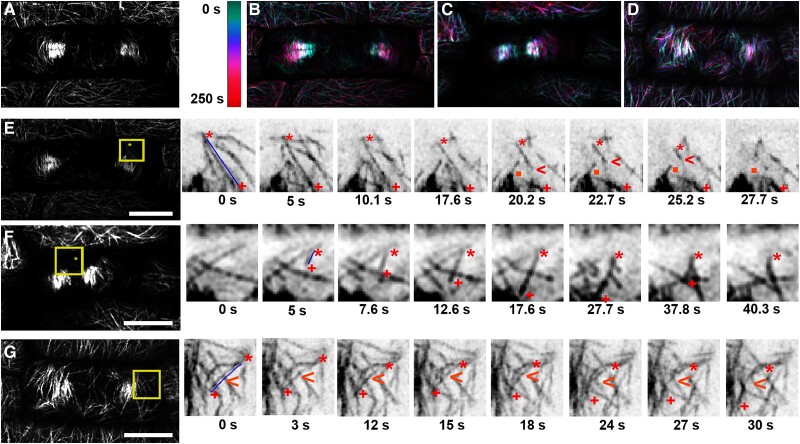
Cortical-telophase microtubules accumulate at the cortex before the phragmoplast contacts the cortex in wild-type maize epidermal cells. Times are indicated in hours:minutes:seconds at the bottom left corner. A) Time-lapse imaging of a cell expressing YFP-TUBULIN from metaphase to telophase. Microtubules at the cortex (top panel), microtubules at the midplane (middle panel), and merged images (cortex: green and midplane: magenta; bottom panel) are shown. Cortical-telophase microtubules faintly accumulate at 14:11, with more accumulating by 29:16; additional intermediary timepoints are shown in Supplemental Fig. S1. B) Time-lapse imaging of microtubules from anaphase to telophase. Merged images (cortex: green, and cell plate and plasma membrane dyed with FM-4-64: magenta). X–Z projections show that the cortical-telophase arrays accumulate before the phragmoplast reaches the cell cortex. Bar is 10 µm for X–Y images and ∼10 µm for the X–Z projection (estimated due to sample drift). Images were acquired using a Zeiss LSM 880 equipped with Airyscan at 100× (NA = 1.46).
Cortical-telophase microtubules were present in over 90% of wild-type cells during telophase (n = 173/190 cells from 26 plants, Fig. 2A). Cortical-telophase microtubule arrays covered 33 ± 2% (mean ± standard error (SE) of the cell cortex (Fig. 2D), with an average anisotropy of 0.12 ± 0.01 arbitrary units (Fig. 2B). These anisotropy values (reflecting the relative orientation of cortical-telophase arrays) were similar to those of microtubule arrays in Arabidopsis shoot meristems during interphase (Boudaoud et al. 2014). The cortical-telophase microtubules were on average typically arranged into anti-parallel arrays perpendicular to the division site (∼50% within 10 degrees of perpendicular relative to the phragmoplast midline, n = 38 microtubule arrays from 19 cells, 7 plants, Fig. 2D), with most plus ends facing the division site. We also observed cortical-telophase microtubule arrays in Arabidopsis root cells (Supplemental Fig. S1C). The results were similar to previous reports showing microtubule-nucleating protein accumulation at the cell cortex (e.g. Kong et al. 2010; Vavrdová et al. 2019; Lucas 2021) or cortical microtubules in moss (Physcomitrium patens) (e.g. Wu and Bezanilla 2014). Therefore, cortical-telophase microtubule arrays were abundant and on average oriented perpendicular to the division site during telophase in maize epidermal cells. Although they are difficult to see, cortical-telophase microtubules may be a conserved feature of plant cells.
Figure 2.
Cortical-telophase microtubules are typically abundant and arranged toward the division site in wild-type cells but are more variable in abundance and organization in tan1 mutant cells. A) A wild-type maize epidermal cell with abundant cortical-telophase microtubules (far left), tan1 mutant cells with abundant (left), asymmetric (middle), or sparse cortical-telophase microtubules (right). Merged images show a midplane view (magenta) and cortex view (green). X–Z shows the X–Z projection, with orange arrowheads indicating cortical microtubules at the top of the cell. B) Cortical-telophase microtubule array anisotropy, n = 38 wild-type arrays (19 cells from 5 plants) and 50 tan1 arrays (25 cells from 9 plants) with median and quartiles indicated by black bars (2-tailed Mann–Whitney U test P-value = 0.005). Schematic diagrams of cells with high and low anisotropy (right). C) Histogram of the average microtubule orientation of the cortical-telophase microtubule array (n = 38 arrays for wild-type and 50 for tan1 mutant cells, 2-tailed Mann–Whitney U test comparing angle values, P value <0.001). Schematic diagram showing angle measurements compared to the division site, D) Relative cortical-telophase coverage, represented as a fraction, was significantly higher in wild-type (38 arrays) than tan1 (54 arrays) cells, 2-tailed Mann–Whitney U test, median and quartiles are indicated with black bars (P value <0.0001). Schematic diagrams with examples of high and low microtubule coverage. Micrographs were taken under a Zeiss LSM 880 (Airyscan,100×, NA = 1.46). Bars are 10 µm.
Analysis of microtubule dynamics
To assess whether cortical-telophase microtubules had different properties compared to phragmoplast microtubules, we treated maize epidermal cells with the microtubule destabilizer propyzamide (5 µM). Propyzamide inhibits microtubule assembly by binding to beta-tubulin (Young and Lewandowski 2000). Cortical-telophase microtubules were more sensitive to 5-µM propyzamide treatment than phragmoplast microtubules. The majority of cortical-telophase microtubules were depolymerized within 30 min of treatment (Supplemental Fig. S2, n = 50/56 cells from 3 plants). After an additional 30 min, all remaining cortical-telophase microtubules were disassembled. In contrast, interphase cortical microtubules were intact after 30 min of propyzamide treatment, depolymerizing after 45 min to 1 h of treatment (n = 117 cells from 2 plants). In contrast, phragmoplast microtubules were resistant to propyzamide treatment and remained intact after 1 h (n = 68/68 phragmoplasts). When incubated with DMSO (negative control), cortical-telophase microtubules were still intact 30 min after treatment (n = 34/69 cortical-telophase microtubule arrays). Phragmoplast (n = 62) and interphase microtubules (n = 82 cells) were also intact. Together, these results indicate that cortical-telophase microtubules are more sensitive to treatment with 5 µM propyzamide than either interphase cortical microtubules or phragmoplast microtubules, suggesting that they are more likely to be single, highly dynamic, and unbundled microtubules that are distinct from the phragmoplast, which is consistent with our imaging data.
To understand how cortical-telophase microtubules formed arrays with their plus ends facing the division site, we examined individual microtubules interacting with the division site in addition to other locations within the cell (Supplemental Table S1 and Supplemental Data Set S1). In wild-type cells, microtubule plus ends were transiently stabilized by pausing or capturing at the division site (Fig. 3, A and C; see Movie 1). When cortical-telophase microtubules contacted the division site, 59.4% of microtubules paused (n = 60/101 microtubules, 4 cells, 3 plants, Table 1), 4.9% underwent immediate catastrophe after touching the division site, and 35.6% passed through the division site without altering of their trajectories. When cortical-telophase microtubules interacted with the division site, 59% (n = 53/90) buckled, indicating that the microtubule was still growing as it was transiently captured at the division site. Median pausing or capture time was 15 s at the division site but 10.0 s in other locations (Fig. 3, A and C; Table 2, P value = 0.03). Transient stabilization of microtubule plus ends at the division site may promote overall perpendicular orientation.
Figure 3.
Cortical-telophase microtubules pause at the division site near TAN1 puncta. A) and B) Time-lapse images of cortical-telophase microtubules (YFP-TUBULIN, green) pausing at the division site (top panels) or passing (bottom panels) through the division site ahead of the phragmoplast in wild-type (WT) A) and tangled1 (tan1) B) cells. Red arrowheads indicate the microtubule plus end. Dotted lines in time-lapse insets mark the division site, as predicted through the extension of FM4-64 cell plate staining (magenta). The scale bar is 10 µm and 5 µm in insets. C) Dot plot of microtubule pause times (s) at division site and other cortex locations in wild type and tan1. Bars represent median with interquartile range. D) Time-lapse images of a wild-type cell cortex with cortical-telophase microtubules and cortical TAN1 localization. Cortical-telophase microtubules ahead of the phragmoplast pause at the division site with no TAN1 puncta (i, top panel) and at the division site with TAN1-puncta (ii, bottom panel). Microtubules are labeled with CFP-TUBULIN (green) and TAN1 by TAN1-YFP (magenta). Arrowheads indicate the respective microtubule plus end. The scale bar is 10 microns and 1 micron in insets. E) Dot plot comparing microtubule pause times(s) at division site locations with or without TAN1 puncta in wild-type cells expressing TAN1-YFP. Each dot represents one microtubule pause time. Error bars are median with interquartile range. P-values ns not significant, * < 0.05, ** < 0.01, *** < 0.001 by Kruskal–Wallis and Dunn's test. Images were acquired using a Zeiss LSM 880 equipped with Airyscan with a 100× (NA = 1.46) lens.
Movie 1.

Time-lapse imaging of cortical-telophase microtubules interacting with the future division site.
Table 1.
Quantification of individual interaction and bundling events between cortical-telophase microtubules and the phragmoplast. (A) Summary of cortical-telophase microtubule bundling times and angles with the phragmoplast. (B) Summary of cortical-telophase microtubule interaction types with the phragmoplast. (B) Fisher's exact test was used, and significant differences are indicated by (**) P < 0.01, (****) P < 0.0001. Phragmoplast-interacting MTs: WT (n = 252, 5 cells, 3 individuals), tan1 (n = 163, 5 cells, 3 individuals)
| A | ||||||
|---|---|---|---|---|---|---|
| Phragmoplast trailing edge | Phragmoplast leading edge | |||||
| Sample | Time bundled (seconds, mean ± SEM) | Angle bundled (degrees, mean ± SEM) | Proportion of microtubules (%, n) | Time bundled (seconds, mean ± SEM) | Angle bundled (degrees, mean ± SEM) | Proportion of microtubules (%, n) |
| Wild type | 45.75 ± 6.781 | 26.57 ± 1.804 | 34 (86) | 108.6 ± 6.922 | 21.48 ± 1.113 | 66 (166) |
| tangled1 | ****63.07 ± 6.305 | 28.44 ± 1.901 | **47 (77) | 97.96 ± 7.745 | 25.14 ± 2.032 | **53 (86) |
| P value | 0.0001 | ns | 0.0074 | ns | ns | 0.0074 |
| B | ||||||
|---|---|---|---|---|---|---|
| Phragmoplast interaction types | ||||||
| Sample | Depolymerized (%, n) | Stayed (%, n) | Severed (%, n) | Stabilized (%, n) | ||
| Wild type | 22 (54) | 78 (197) | 37 (94) | 41 (103) | ||
| tangled1 | **10 (16) | **90 (147) | **56 (92) | 34 (55) | ||
| P value | 0.0019 | 0.0019 | 0.002 | ns | ||
Table 2.
Percentage of cortical-telophase microtubule pausing at or passing through the division site in wild-type and tangled1 plants. Comparison of microtubule interactions with the division site between wild type (n = 3 plants, 4 cells) and tangled1 (n = 4 plants, 4 cells). P-values ns not significant, *** ≤ 0.001 by Fisher's exact test
| Wild Type | tangled1 | |
|---|---|---|
| % Pause | 59.4% (60/101) | 27% (26/96)*** |
| % Pass | 35.6% (36/101) | 65% (62/96)*** |
| % Depolymerize | 4.9% (5/101) | 8% (8/96)ns |
To determine if microtubule pausing at the division site was due to contact with opposing microtubules, we examined how often microtubules contacted antiparallel microtubules at the division site. Microtubule pausing at the cortical division site ahead of the phragmoplast did not typically occur through antiparallel interactions with microtubules located on the other side of the division site. For wild-type cortical-telophase microtubules, antiparallel contact at the division site occurred 13% of the time (n = 8/61 microtubules from 12 cells from 6 plants). This number of antiparallel contacts was similar to the number of cortical-telophase microtubule contacts that did not occur at the division site, 2% (n = 1/48 microtubules from 12 cells from 6 plants, no significant difference, 2-tailed Fisher's exact test). Similarly, 25% of phragmoplast leading-edge microtubules had antiparallel contacts at the division site (n = 5/20 microtubules from 8 cells, 5 plants, no significant difference compared to cortical-telophase microtubules at the division site, 2-tailed Fisher's exact test). Together, these results suggest that cortical-telophase microtubules did not pause at the division site solely due to contact with antiparallel microtubules. Instead, microtubules paused at the division site regardless of whether a microtubule from the other side was present, suggesting that a protein (or proteins) located at the division site mediated microtubule pausing.
TAN1 functions in microtubule plus end capture
To analyze the role of the microtubule-binding protein TAN1 in microtubule plus end capture, we measured microtubule plus end pausing using time-lapse imaging of wild-type cells expressing both TAN1-YFP and CFP-TUBULIN. TAN1-YFP forms discrete puncta at the division site during telophase (Fig. 3D, see Movie 2) (Walker et al. 2007; Rasmussen et al. 2011; Martinez et al. 2017). In time-lapse images, cortical-telophase microtubules appear to interact with stable TAN1 puncta at the division site (see Movie 2). We found that cortical-telophase microtubules remained near TAN1-YFP puncta longer than other regions within the division site in wild-type maize cells (Fig. 3, D and E). Microtubule plus ends paused near TAN-YFP1 puncta for ∼20 s (20 ± 1.6 s, median ± SEM, n = 39 microtubules, 4 plants). By contrast, microtubule plus ends that contacted regions of the division site distinct from TAN1-YFP puncta paused for ∼10 s (10 ± 1.1, median ± SEM n = 48 microtubules, 4 plants, Fig. 3E). Together, these findings suggest that TAN1 or other division site-localized proteins in close proximity (within Airy Scan resolution limits of ∼150 nm) promote cortical microtubule plus end pausing or capture during telophase. This microtubule interaction is consistent with the results of in vitro dynamic assays where TAN1 transiently captured microtubules at high contact angles (Martinez et al. 2020). Together, these results suggest that TAN1 or other nearby division site-localized proteins increase microtubule pausing or capture at the division site, which over time leads to cortical-telophase arrays that are on average perpendicular to the division site, as observed in Fig. 2C.
Movie 2.
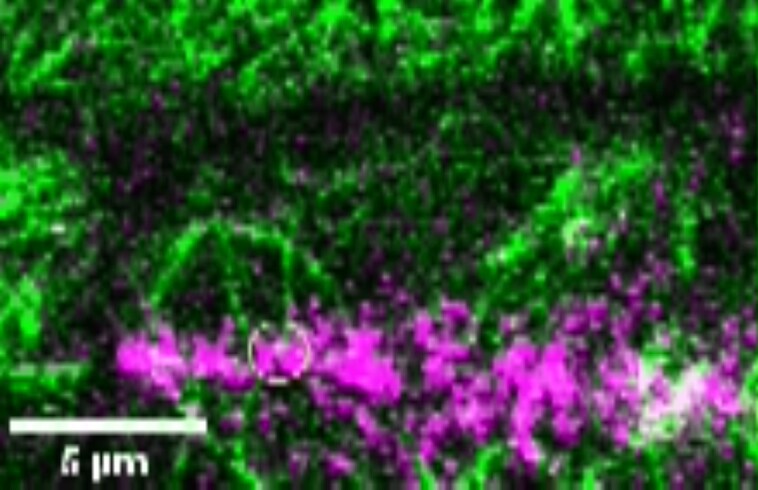
Time-lapse imaging of a cortical-telophase microtubule interacting with TAN1-YFP puncta.
We hypothesized that loss of TAN1 from the division site would lead to defects in cortical-telophase microtubule organization, so we examined cortical-telophase microtubules in the maize tan1 mutant. Cortical-telophase microtubule arrays were sparse or missing in nearly 30% of tan1 cells (n = 24/122 cells from 24 plants, e.g. Supplemental Fig. S1, E and F). When cortical-telophase arrays were present in tan1 cells, they were often unevenly distributed (asymmetric) (Fig. 2A). Furthermore, cortical-telophase microtubule arrays in the tan1 mutant were less anisotropic (Fig. 2B) than wild-type cortical-telophase arrays. In addition, unlike wild-type cortical-telophase arrays, tan1 mutant arrays were not typically oriented toward the division site (Fig. 2C, median orientation 49.5 ± 3 degrees relative to the phragmoplast midline, P < 0.0001 Mann–Whitney U test). These data suggest that TAN1 promotes proper cortical-telophase microtubule array organization.
In contrast to wild-type cells, cortical-telophase microtubules were not transiently stabilized at the division site in tan1 mutant cells, showing no significant difference in microtubule pausing at the division site versus other cortical locations (Fig. 3, B and C; Supplemental Fig. S3; Supplemental Data Set S1; Kruskal–Wallis test and Dunn's test, P = 0.11; adjusted P-value = 1). Unlike wild-type cells, where ∼60% of cortical-telophase microtubules paused at the division site, significantly fewer cortical-telophase microtubules paused in the tan1 mutant (27%, n = 26/96, Fisher's exact test, P value < 0.00001, Table 2). Instead, the majority of cortical-telophase microtubules in the tan1 mutant grew past the division site without any alteration in their trajectories (65% n = 62/96, example in Fig. 3B, 6 cells from 5 plants, Table 2) or shrunk immediately (8%, n = 8/96). Microtubules that interacted with the division site displayed similar buckling frequency compared to wild-type cortical microtubules (58%, n = 22/38 compared to 59%, 53/90, P value = 1, Fisher's exact test). TAN1 also plays a role in mediating other microtubule dynamics during telophase, as measured using the Dynamic Kymograph Plugin in FIJI (Zhou et al. 2020) Supplemental Fig. S4). Microtubule growth, shrinkage, and pause during telophase tended to be slower in tan1 mutants compared to wild type (Supplemental Fig. S3). These data, together with the finding that microtubule pausing or capture increased at TAN1 puncta, suggest that TAN1 directly or indirectly promotes both microtubule plus end capture and shrinkage at the division site.
Interaction of cortical-telophase microtubule arrays with the phragmoplast
We hypothesized that when the phragmoplast reached the cortex, it would interact with cortical-telophase microtubules. To examine how cortical-telophase microtubule arrays interacted with the phragmoplast, we used time-lapse imaging. Cortical-telophase microtubules were typically added into the phragmoplast by parallel bundling, as described below. To assess how individual microtubules from the cortical-telophase array interacted with the phragmoplast, we first identified sites of microtubule nucleation at the cortex from cortical-telophase microtubules that were clearly distinct from the phragmoplast. Next, we determined how these microtubules interacted with the phragmoplast. An annotated micrograph describes the terms used here, such as leading and lagging edges (Supplemental Fig. S5). When cortical-telophase microtubules contacted the phragmoplast, most (78%, n = 197/252 microtubules from 5 cells from 3 plants) were incorporated into the leading edge of the phragmoplast by parallel bunding (Fig. 4G, see Movies 3 and 4). After bundling into the phragmoplast, the microtubules would either remain connected to the original cortical-telophase array during the 252 s time lapse (41%, n = 103/252, Fig. 4G) or become fully incorporated into the phragmoplast by severing the connection between the cortical-telophase array and the phragmoplast (37%, n = 94/251, Fig. 4E; see Movie 5). We speculate that severing was performed by the microtubule-severing protein KATANIN, which localizes to the distal phragmoplast (Nakamura et al. 2010; Panteris et al. 2011), possibly via the microtubule-binding protein MACET4/CORD4 (Sasaki et al. 2019; Schmidt and Smertenko 2019). The remaining cortical-telophase microtubules that contacted the phragmoplast underwent catastrophe after touching the phragmoplast (22%, n = 55/252, Fig. 4F, see Movie 6). Most (66%, n = 166/252) cortical-telophase microtubules interacted with the leading edge, although others interacted with the lagging edge (n = 86/252) and then primarily were incorporated into the phragmoplast by low-angle parallel bunding (<45 degrees, Table 1).
Figure 4.
Time-lapse images of cortical-telophase microtubules interacting with the phragmoplast using YFP-TUBULIN to label microtubules. A) A single early snapshot of a maize dividing cell during telophase with surrounding interphase cells. B) Color-coded time projection showing the movement of the phragmoplast and cortical-telophase microtubules of the cell in A). C) Time projection of cell in F), D) time projection of cell in G). E) The cell shown in A) at a later time point. Representative example of severing leading to the incorporation of a cortical-telophase microtubule into the phragmoplast. Microtubules of interest are indicated with an adjacent blue line; red asterisks indicate the cortical-telophase microtubule minus ends and red pluses indicate the microtubule plus end. Red arrowheads show severing followed by depolymerization. The orange square marks the new microtubule minus end after severing. F) Representative example of depolymerization of a cortical-telophase microtubule following contact with the phragmoplast. G) Representative bundling of a cortical-telophase microtubule into the phragmoplast. Orange arrowheads show a cortical-telophase microtubule incorporated into the phragmoplast by parallel bundling. Bars are 5 µm, Time (s) rounded to a 10th of a second. Images were acquired using a Zeiss LSM 880 equipped with Airyscan with a 100× (NA = 1.46) lens.
Movie 3.
Time-lapse imaging of a YFP-TUBULIN labeled cortical-telophase microtubule bundling into the phragmoplast.
Movie 4.

Time-lapse imaging of a YFP-TUBULIN labeled cortical-telophase microtubule bundling into the phragmoplast.
Movie 5.

Time-laspse imaging of a YFP-TUBULIN labeled cortical-telophase microtubule incorporation into the phragmoplast and severing.
Movie 6.

Time-lapse imaging of a YFP-TUBULIN labeled cortical-telophase microtubule contacting the phragmoplast, then depolymerizing.
Cortical-telophase microtubules, when present, were also added into tan1 mutant phragmoplasts. Similar to wild-type microtubules, tan1 cortical-telophase microtubules were incorporated into the phragmoplast, although relatively more microtubules interacted with the lagging edge of the phragmoplast (Table 1). Proportionally more of the microtubules that interacted with the phragmoplast were eventually incorporated in tan1 phragmoplasts (90% n = 147/163% versus 78% in wild-type cells n = 197/252; Table 1). These data indicate that cortical-telophase microtubules in close contact with the phragmoplast were primarily added to the leading edge in both wild-type and tan1 cells. Although cortical-telophase microtubules interacted similarly with the phragmoplast in wild-type and tan1 cells, the abundance, orientation, and relative accumulation of cortical-telophase microtubules were more variable in tan1 cells (Fig. 2, A and D).
Effects of cortical-telophase microtubule accumulation on the trajectory of phragmoplast expansion
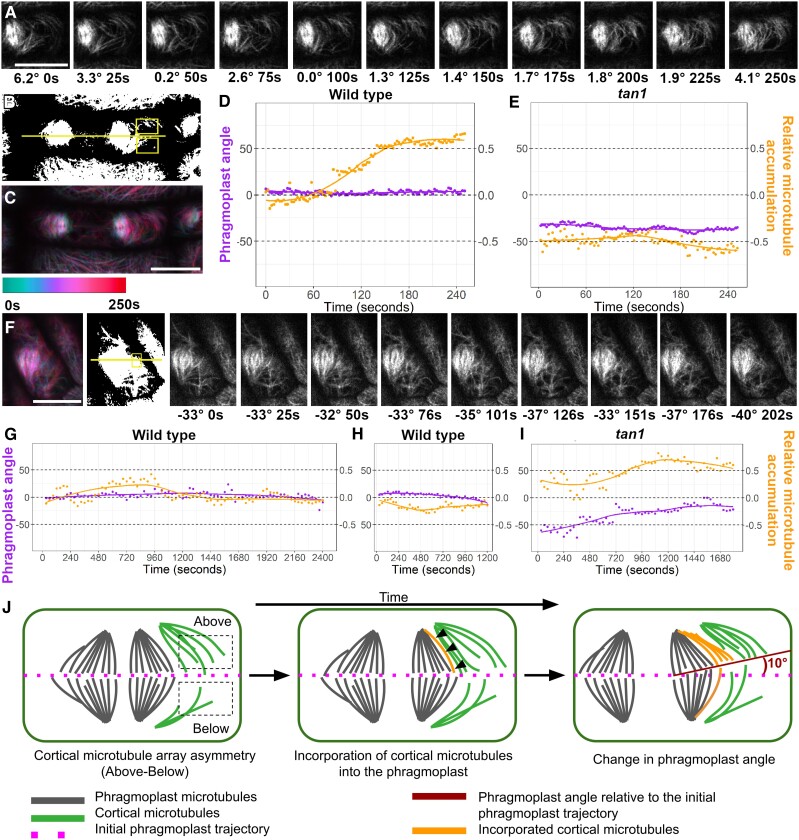
We hypothesized that the addition of microtubules from the cortex into the phragmoplast would alter the direction of phragmoplast expansion. Using time-lapse imaging, we measured the movement of the phragmoplast over time and also measured the corresponding cortical-telophase microtubule array. If more cortical-telophase microtubules accumulated on one side of the phragmoplast, the phragmoplast moved toward the same direction within ∼15 min (960 s, n = 6 cells). Terms describing phragmoplasts are defined in the model (Supplemental Fig. S5; Fig. 5J). We compared the phragmoplast trajectory with the relative accumulation of cortical-telophase microtubules “above” and “below” the division plane (Fig. 5, A–C). The phragmoplast trajectory was measured as an angle parallel to the division site, defined in Fig. 5A: if the angle is positive, it indicates that the phragmoplast angle moved above the division plane. If the angle is negative, the phragmoplast angle moved down below the division plane. We selected 2 equally sized region-of-interest boxes (Fig. 5B) above and below the phragmoplast to measure the relative cortical-telophase microtubule accumulation in front of the expanding phragmoplast. Relative cortical-telophase microtubule accumulation was measured by subtracting the microtubule coverage below from above. Positive values indicate that more microtubules accumulate above the phragmoplast.
Figure 5.
Long-term uneven accumulation of cortical-telophase microtubules is correlated with changes in phragmoplast direction. A) to D) A wild-type phragmoplast: A) Time-lapse imaging with phragmoplast angle relative to the division site and time (s) indicated below. Time-lapse images were acquired using a Zeiss LSM 880 with Airyscan (100×, NA = 1.46) or a Yokogawa spinning disk with a Nikon stand (100×, NA 1.45). B) Thresholded image with ROI (yellow rectangles) selected to measure relative cortical-telophase microtubule accumulation above and below the phragmoplast. The phragmoplast trajectory is indicated by a yellow line. C) Time projection with time-color legend. D) Graph comparing changes in phragmoplast angle over time (purple) and relative cortical-telophase microtubule accumulation (orange) above or below the phragmoplast. E) and F) A tan1 phragmoplast E) Graph of changes in phragmoplast angle and cortical-telophase microtubule accumulation in tan1 over time. F) Time-lapse imaging of tan1; phragmoplast angle and time are shown below. G-I) Longer time lapses: G) A wild-type cell with little overall phragmoplast movement. H) Wild-type cell with consistent cortical-telophase microtubule accumulation below the phragmoplast and downward phragmoplast angle movement. I) tan1 cell with consistent cortical-telophase microtubule accumulation above the phragmoplast with phragmoplast angle movement towards the top. Bars = 10 µm. J) Model of the cell cortex of maize epidermal cells showing cortical-telophase microtubule accumulation, incorporation into the phragmoplast, and changes in the trajectory of the phragmoplast over time.
The direction of phragmoplast expansion in wild-type cells typically followed a flat trajectory within 5 min, with <10 degrees overall change (n = 5, Fig. 5D, Supplemental Fig. S6). During longer timeframes (18–30 min), wild-type phragmoplast trajectories were more variable, but overall, they did not persistently change direction (n = 2/4, Supplemental Fig. S7), which is consistent with previous time-lapse observations (Martinez et al. 2017). In wild-type cells with little overall phragmoplast angle displacement, cortical-telophase microtubule accumulation varied over time but did not maintain uneven accumulation (Fig. 5G, Supplemental Figs. S6D and S7B). By contrast, sustained accumulation of cortical-telophase microtubules either above or below was correlated with phragmoplast movement in the same direction (Fig. 5H, Supplemental Fig. S7, A and C).
In tangled1 mutants, both phragmoplast expansion direction and cortical-telophase microtubule array accumulation were more variable than in wild-type plants, but the relationship between asymmetric cortical-telophase microtubule accumulation and changes in phragmoplast direction was the same (Supplemental Fig. S8). Over longer timeframes, sustained asymmetric cortical-telophase microtubule accumulation in tan1 mutants also correlated with changes in phragmoplast trajectories in the same direction (Fig. 5I, Supplemental Fig. S9). Therefore, in both wild-type and tan1 mutants, cortical-telophase microtubule accumulation preceded changes in the direction of the phragmoplast. Cortical-telophase microtubules interacted less with the division site in tan1 mutants, often passing through without any pause or change in trajectory. We speculate that this in turn leads to disorganized and asymmetric cortical-telophase microtubule arrays (Fig. 2). These asymmetric cortical-telophase arrays are then added into the phragmoplast (Fig. 4), leading to defects in phragmoplast guidance observed as changes in phragmoplast direction over time in the tan1 mutant (Fig. 5). Changes in phragmoplast direction mediated by cortical-telophase also occurred in wild-type cells, albeit at a lower frequency.
Discussion
We showed that cortical-telophase microtubule arrays accumulate and interact with the division site in maize during telophase before the phragmoplast reaches the cell cortex. Cortical-telophase microtubules that nucleated directly at the cortex were our focus, although some cortical-telophase microtubules may also come from the nucleus. Cortical-telophase microtubule nucleation is reminiscent of branching clusters of newly regenerating interphase cortical microtubules that form after the removal of microtubule-depolymerizing drugs (Wasteneys and Williamson 1989). Previous reports of microtubules stabilized with Taxol showed that cortical-telophase microtubules nucleated directly on the cell cortex in the monocot durum wheat (Triticum durum) (Panteris et al. 1995), while they may have originated from the nuclear envelope in tobacco (Nicotiana tabacum) cultured cells (Van Damme and Geelen 2008). Both cortical-telophase microtubules and nuclear-envelope-nucleated microtubules accumulate at the cortex in Arabidopsis cotyledon cells (Lucas 2021). Whether cortical-telophase microtubules primarily originate at the cortex or the nucleus may depend on the species.
We showed that cortical-telophase microtubules often orient toward the division site due to increased microtubule plus end pausing or capture at the division site (Fig. 3) and that cortical-telophase microtubules are most often added into the phragmoplast by parallel bundling at low contact angles (Table 1). Cortical-telophase microtubules are bundled into the phragmoplast leading edge, perhaps similar to previously described “mini-phragmoplasts,” which are preassembled phragmoplast modules that are added to the phragmoplast by parallel bundling during endosperm cellularization (Otegui and Staehelin 2000; Lee and Liu 2013).
Uneven or asymmetric cortical-telophase microtubule accumulation was correlated with changes in phragmoplast trajectories over time (Fig. 5J). While it is possible that asymmetric accumulation of cortical-telophase microtubules and changes in phragmoplast angles over time both respond independently to some yet unknown cue, we propose that cortical-telophase microtubules, which are incorporated into the phragmoplast by parallel bunding, fine-tune the positioning of the phragmoplast so it reaches the exact division site at the cell cortex. The localized addition of preloaded and properly oriented microtubules also provides a plausible mechanism to achieve phragmoplast insertion at the cell cortex in cells with polarized cytokinesis. In maize, as well as other model systems, most divisions are polarized: the phragmoplast contacts the cortex at one location, then expands outward at the cortex to complete division. Highly polarized cytokinesis occurs during periclinal cambial divisions, in which phragmoplasts traverse tens to hundreds of microns to complete division (Kajala et al. 2014; Fischer et al. 2019). Other examples include Arabidopsis epidermal cells (Cutler and Ehrhardt 2002; Lucas and Sack 2012), cultured cells (Chan et al. 2005), and vacuolated Nautilocalyx cells (Venverloo and Libbenga 1987). A guiding mechanism provided by local cortical microtubules, directly incorporated into the phragmoplast as it expands at the cell cortex, would provide the necessary specificity in targeting. The direct addition of cortical-telophase microtubules into the phragmoplast to alter its local position occurs when the phragmoplast is within the micron range of the cell cortex. We suspect that this phragmoplast zippering event occurs when the addition of vesicles to the cell plate becomes slow and variable (van Oostende-Triplet et al. 2017).
We showed that cortical-telophase microtubule plus ends are stabilized at the division site near TAN1 puncta. In vitro, when TAN1 is added to microtubule dynamic assays, it both decreases shrinkage rates and slows microtubule growth compared to microtubules without TAN1 addition (Martinez et al. 2020). This is consistent with the notion that TAN1 potentially stabilizes microtubules. However, in vivo, tan1 mutants also have slower microtubule growth and shrinkage rates than wild type (WT). It is still unclear whether TAN1 could also play a role in nucleating microtubules or other functions. Given the intriguing contact angle-independent in vitro microtubule interactions observed during co-incubation with HIS-TAN1 (Martinez et al. 2020), we speculate that in vivo, TAN1 may block microtubules from passing through the division site by capturing the microtubule plus ends. Our hypothesis is that high contact angle microtubule interactions with TAN1 stabilize their plus ends to increase microtubule pause or capture times at the division site. An alternate hypothesis is that other division site-localized proteins in close proximity to TAN1 may mediate this activity. However, no other end-on microtubule-interacting proteins in plants have yet been shown to localize to the division site (Rasmussen and Bellinger 2018; Livanos and Müller 2019).
In addition to TAN1, other candidate MAPs might also contribute to the stabilization of cortical microtubules at the division site during telophase. The kinesin-like calmodulin-binding protein (KCBP), a processive minus end-directed kinesin-14 that localizes to the division site in moss and Arabidopsis, is a highly plausible candidate (Miki et al. 2014; Buschmann et al. 2015; Yamada et al. 2017). Analogous minus end-directed motor proteins in animals and yeast (Saccharomyces cerevisiae), dyneins, capture and stabilize microtubule plus ends at the cell cortex during division. Dyneins play a critical role in division plane positioning by pulling on astral spindle microtubules at the cell cortex to adjust the position of the spindle (Busson et al. 1998; Hendricks et al. 2012; Laan et al. 2012). In Arabidopsis, kcbp mutants do not have defects in division plane positioning, possibly due to a redundant function of a yet unknown minus end-directed kinesin at the division site. Whether KCBP puncta localized at the division site interact directly with microtubules is also unknown (Buschmann et al. 2015).
KCBP interacts with a transient division site-localized protein called AUXIN INDUCED IN ROOT CULTURES9 (AIR9) (Buschmann et al. 2015). AIR9 localizes to the division site as the phragmoplast reaches the cortex but is not at the division site from metaphase until the end of telophase in tobacco cultured cells. Therefore, AIR9 is not at the division site when the cortical-telophase microtubules originate, or when interactions between cortical-telophase microtubules and division site-localized proteins begin (Buschmann et al. 2006). The lack of localization of AIR9 during the majority of telophase suggests that it is unlikely to be a major player in plus end cortical-telophase microtubule stabilization at the division site. Another candidate is the antiparallel microtubule-bundling protein MAP65-4, which localizes to the division site (Li et al. 2017). However, since most cortical-telophase microtubules are not bundled into antiparallel microtubule arrays, but instead interact directly with the division site, it is more likely that cortical-telophase microtubules interact with other classes of microtubule-binding proteins.
POK1 and POK2, which are plus end-directed kinesin-12 motor proteins that localize to the division site (Lipka et al. 2014; Chugh et al. 2018; Herrmann et al. 2018), might stabilize cortical-telophase microtubules. Since POK1 and POK2 directly interact with TAN1 (Müller et al. 2006; Rasmussen et al. 2011; Lipka et al. 2014; Mills et al. 2022), they might also function together with TAN1 at the division site to capture microtubules. POK1 and POK2 may capture cortical-telophase microtubules at the division site just behind their plus ends and then move toward the plus ends, effectively pushing the minus ends away from the division site. This idea is consistent with microtubule buckling observed following contact with the division site.
Interactions of microtubules and microfilaments with division site-localized proteins such as MYOSINVIII are likely broadly conserved features of division plane positioning in plants. MYOSINVIII interacts with both actin and microtubules: their combined interaction, which is mediated by MYOSINVIII, guides the phragmoplast toward the proper division site (Wu and Bezanilla 2014). Fascinatingly, both actin and MYOSIN VIII participate in incorporating peripheral microtubules, defined as microtubules that nucleate from the phragmoplast and grow outward toward the cortex, back into the phragmoplast. Drug treatments that block myosin activity cause defects in phragmoplast guidance toward the cortex (Molchan et al. 2002), as do drugs that disrupt actin filaments (Yoneda et al. 2004; Wu and Bezanilla 2014). MYOSINXI also promotes proper division plane positioning in both maize and Arabidopsis (Abu-Abied et al. 2018; Nan et al. 2021). Cytoskeleton-mediated (actin-based) division plane corrections also occur during telophase in mouse (Mus musculus) epithelial cells, suggesting that analogous mechanisms occur in other eukaryotes (Lough et al. 2019).
Materials and methods
Plant growth and imaging conditions
Maize (Zea mays) plants were grown in 1-L pots in standard greenhouse conditions, with the temperature setpoint ∼32 °C with a photoperiod of 14 h of light (∼400 μE*m−2 s−1)/10 h of dark. Supplemental lighting was provided by 1,000-w high-pressure sodium bulbs (Gravita Pro Plus 1000W EL). Plants were grown in soil containing 20% peat, 50% bark, 10% perlite, and 20% medium vermiculite with calcium nitrate (90 ppm CA 75 ppm N), magnesium nitrate (45 ppm Mg, 50 ppm N), and further supplemented with Osmocote (NPK 14-14-14). Maize plants between 3 and 5 wk old containing YFP-TUBULIN, CFP-TUBULIN, TANGLED1-YFP (Mohanty et al. 2009; Wu et al. 2013), or the tangled1 mutant were used for imaging and identified by microscopy or by genotyping as previously described (Martinez et al. 2017). The primers used for genotyping were as follows: for TANGLED1-YFP (TAN LSP1 5′ ACGACCGTTAGCACAGAACC and GFP5REV 5′ CTGAACTTGTGGCCGTTTACGTCGC); for YFP-TUBULIN (TubAlpha Rp1 5′ GGTTTCGGGTGATCCCTATT and TubalphaFp1 5′ GCAAGGTTTCGATTTCCGTA); and for CFP-TUBULIN (BTUBR3187 5′ GACAGGCGGGCATAAGATCC and TUBbeta FP 5′ CGAATTTTCGAATCCTCAGC). Leaves were removed from plants until the ligule height was <2 mm. Abaxial symmetrically dividing leaf blade samples were mounted in water between cover glass and glass slides (Fisherbrand) or in a Rose chamber, as previously described (Rasmussen 2016). For FM4-64 staining, leaf samples were mounted in 50 μM FM-464 and placed in a Rose chamber for imaging. Three or more plants per genotype were analyzed. Room temperatures during imaging were between 21 and 24 °C.
Arabidopsis thaliana seedlings were grown on ½ strength Murashige and Skoog (MS) medium solidified with 0.8% agar. Plates were sealed with surgical tape (3 M) and grown vertically in a growth chamber (Percival) with 16-h white light ∼111 µE*m−2 s−1 [F17T8/TL741 Fluorescent Tube (Philips)/8-h dark cycles with a 22 °C temperature set point]. Arabidopsis plants containing CFP-TUBULIN (identified by microscopy) were imaged between 3 and 5 d after germination. Seedlings were mounted in water and covered with a cover slip. Root epidermal cells from the meristematic zone were imaged at 23 °C.
Confocal microscopy
Micrographs and short time-lapse images were taken with a Zeiss LSM 880 confocal laser scanning microscope equipped with Airyscan with a 100×, NA = 1.46, oil immersion objective lens. A 514-excitation laser with bandpass (BP) filters 465–505 with long-pass (LP) 525 filter was used with Airyscan super resolution mode. Images captured using the Zeiss LSM 880 were processed using default Airyscan settings with ZEN software (Zeiss). For longer time-lapse imaging, 30 s intervals were used to capture images of microtubules at the cortex to measure both cortical-telophase microtubule accumulation and the orientation of the phragmoplast (data used in Fig. 5, G–I) with a Yokogawa W1 spinning disk microscope with an EM-CCD camera (Hamamatsu 9100c) and Nikon Eclipse TE inverted stand with a 100×, NA 1.45, oil immersion objective lens controlled with Micromanager-1.4 with an ASI Piezo Z stage and a 3 axis DC servo motor controller. Solid-state lasers (Obis) between 40 and 100 mW were used with standard emission filters from Chroma Technology. For YFP-TUBULIN or TANGLED1-YFP, a 514 laser with emission filter 540/30 was used. For CFP-TUBULIN, a 445 laser with emission filter 480/40 was used. For the membrane dye FM4-64, a 516 nm laser with emission filter 620/60 was used.
Telophase cells were identified by the presence of a phragmoplast, and cortical-telophase microtubules were imaged on the cortical edges of epidermal cells. Two-dimensional projections, time projections, and 3-dimensional reconstructions of z stacks and time-lapse images were generated in FIJI (ImageJ, http://rsb.info.nih.gov/ij/). Image brightness and contrast were altered using the linear levels option, and figures were assembled with FIJI and Gnu Image Manipulation Program (GIMP https://www.gimp.org/downloads/). Drift during time-lapse imaging was corrected with StackReg https://imagej.net/StackReg using the translation option (Thévenaz 1998).
Quantification of microtubule array organization and coverage
Maize lines expressing YFP-TUBULIN were used to examine the microtubule cytoskeleton. To measure anisotropy (Fig. 2B) and orientation (Fig. 2D), TIFF image files were converted to PNG files using FIJI software and processed with the FibrilTool plugin (Boudaoud et al. 2014). For wild-type plants, 38 arrays from 19 transverse cell divisions during telophase were measured from 5 plants with median anisotropy 0.11 ± 0.01 A.U. For tan1 mutants, 50 arrays from 25 transverse cell divisions during telophase were measured from 9 plants (0.07 ± 0.01 A.U.).
To measure the percent microtubule coverage in Fig. 2C, image files were made binary and thresholded using mean fluorescence and processed using the Area/Volume fraction function in the BoneJ plugin (https://imagej.net/BoneJ;Doube et al. 2010). The median value for wild-type cells (n = 38 arrays, coverage fraction 0.33 ± 0.02) is significantly different from the median value for tan1 mutant cells (n = 54 arrays, coverage fraction 0.20 ± 0.01, Mann–Whitney test, P < 0.0001).
Measuring microtubule dynamics
Time-lapse imaging was used to measure microtubule interactions at the division site, near TAN1 puncta, and with the phragmoplast. The division site was defined as a location typically parallel or perpendicular to the long axis of the cell and corresponding to the position of the phragmoplast midplane or based on the accumulation of the membrane dye FM4-64 at the cell plate. In tan1 mutants, the “division site” was defined the same way unless the phragmoplast was misoriented, in which case the “division site” was defined as the midplane of the phragmoplast or the cell plate location defined using the membrane dye FM4-64. Individual microtubule movements were measured using the Dynamic Kymograph plugin (https://imagej.net/plugins/dynamic-kymograph) in FIJI and binned into categories (for phragmoplast interactions). Growth, pause, and shrinkage rates were measured by tracing the outlines of dynamic kymographs using regions of interest (ROIs) in FIJI. Pauses were defined as the presence of a microtubule plus end in a region ± 3 pixels.
Time-lapse imaging was used to compare the abundance of cortical-telophase microtubules and the phragmoplast angle over time. The phragmoplast angle at each time point was measured in FIJI and saved in Google sheets or Excel (Microsoft Office). Time-lapse image files were first processed to remove drift using the transformation selection within StackReg (Thévenaz 1998) and to correct for photobleaching using bleach correction (exponential fit) in FIJI. Next, images were made binary and thresholded using mean fluorescence and processed using the Area/Volume fraction function in the BoneJ plugin (Doube et al. 2010). Two equally sized ROIs were selected above and below the phragmoplast, such that the ROIs captured cortical-telophase microtubule accumulation near the phragmoplast but not touching the phragmoplast at any time frame. The bottom ROI was subtracted from the top ROI. A positive value indicated more microtubule density or accumulation on the top half of the cell. Both phragmoplast angle and relative cortical-telophase microtubule accumulation were graphed together by time using R, RStudio (Version 1.3.1093), and ggplot2 (Wickham et al. 2008; R Core Team 2013). Figures were made using the Gnu Image Manipulation Program (Gimp, versions 2.10.22-2.10.32) with no interpolation during scaling and linear adjustments to levels.
Statistical analysis
Microtubule anisotropy measurements were made for wild-type and tan1 mutant plants described above. Differences in anisotropy were analyzed with GraphPad Prism, and statistical significance was determined with a Mann–Whitney U test, P = 0.0054. Microtubule coverage of the cortex during telophase was measured in wild-type and tan1 mutant plants. Differences in coverage were analyzed with GraphPad Prism, and significance was determined with a Mann–Whitney U test, P < 0.0001. Microtubule dynamics data were graphed in GraphPad Prism and statistically analyzed with R using the Kruskal–Wallis test followed by Dunn's test (P-values adjusted with Bonferroni correction). More information is available in Supplemental Data Set S1.
Accession numbers
Sequence data from this article can be found in the GenBank/EMBL libraries under the following accession number: TANGLED1: NP_001105636.1
Supplementary Material
Acknowledgments
We thank Professors Henrik Buschmann (Osnabrück University) and Laurie Smith (UCSD) and the reviewers for their helpful comments. We also thank Professor Meng Chen (UCR) for procuring funds for the Zeiss 880 (NIH 3R01GM087388-06S2) and Dr. David Carter (UCR) for Zeiss 880 training. Funding to C.G.R.: NSF-CAREER 1942734, NSF 1716972, and USDA CA-R-BPS-5108-H; funding to A.N.U.: NSF1922642; funding to M.A.B.: Department of Education GAANN #P200A150300.
Contributor Information
Marschal A Bellinger, Department of Botany and Plant Sciences, Center for Plant Cell Biology, Institute for Integrative Genome Biology, University of California, Riverside, CA 92521, USA.
Aimee N Uyehara, Department of Botany and Plant Sciences, Center for Plant Cell Biology, Institute for Integrative Genome Biology, University of California, Riverside, CA 92521, USA.
Lindy Allsman, Department of Botany and Plant Sciences, Center for Plant Cell Biology, Institute for Integrative Genome Biology, University of California, Riverside, CA 92521, USA.
Pablo Martinez, Biochemistry Graduate Group, University of California, Riverside, CA 92508, USA.
Michael C McCarthy, Radical Research, LLC, Riverside, CA 92521, USA.
Carolyn G Rasmussen, Department of Botany and Plant Sciences, Center for Plant Cell Biology, Institute for Integrative Genome Biology, University of California, Riverside, CA 92521, USA; Biochemistry Graduate Group, University of California, Riverside, CA 92508, USA.
Author contributions
M.A.B.: formal analysis, investigation, writing—original draft, visualization, and funding acquisition. A.N.U.: formal analysis, investigation, writing—original draft writing—review and editing, visualization, and funding acquisition. P.M.: investigation, writing—review and editing, and funding acquisition. M.C.M.: resources and visualization. L.A.: investigation and visualization. C.G.R.: formal analysis, investigation, writing—original draft, writing—review and editing, visualization, supervision, project administration, and funding acquisition.
Supplemental data
The following materials are available in the online version of this article.
Supplemental Figure S1 . Cortical-telophase microtubules in wild-type maize epidermal cells, Arabidopsis root cells, and very sparse cortical-telophase microtubules in maize tan1 mutant epidermal cells.
Supplemental Figure S2 . Propyzamide treatment of maize epidermal cells depolymerizes cortical-telophase microtubules but does not depolymerize phragmoplast microtubules.
Supplemental Figure S3 . Wild-type and tangled1 microtubule dynamics during telophase.
Supplemental Figure S4 . Examples of microtubule dynamics measurements using the Dynamic Kymograph plugin tool in FIJI.
Supplemental Figure S5 . Description of the features of cells in telophase extracted for analysis.
Supplemental Figure S6 . Short time lapses (<5 min) of 4 different wild-type phragmoplasts showing little change in direction.
Supplemental Figure S7 . Longer time lapses (>8 min) of wild-type phragmoplasts with >10-degre changes in the direction of movement (A, C) and (B, D) with < 10-degre changes in the direction of movement.
Supplemental Figure S8 . Short time lapses (5 min or less) of 5 different tan1 phragmoplasts showing little change in direction, but frequently unevenly distributed cortical-telophase array.
Supplemental Figure S9 . Longer time lapses (>12 min) of tan1 phragmoplast angle measurements compared to relative cortical microtubule accumulation.
Supplemental Table S1 . Summary of microtubule dynamics in wild-type and tangled1 cells across different locations at the cell cortex.
Supplemental Data Set S1 . Results of Kruskal–Wallis followed by Dunn's test.
Funding
National Institute of Health NIH 3R01GM087388-06S2, National Science Foundation NSF-CAREER 1942734, NSF 1716972, NSF1922642, United States Department of Agriculture USDA CA-R-BPS-5108-H, Department of Education GAANN #P200A150300.
Data availability
Maize lines are available at the Maize Cooperative (http://maizecoop.cropsci.uiuc.edu/) or upon request.
References
- Abu-Abied M, Belausov E, Hagay S, Peremyslov V, Dolja V, Sadot E. Myosin XI-K is involved in root organogenesis, polar auxin transport, and cell division. J Exp Bot. 2018:69(12):2869–2881. 10.1093/jxb/ery112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boruc J, Weimer AK, Stoppin-Mellet V, Mylle E, Kosetsu K, Cedeño C, Jaquinod M, Njo M, De Milde L, Tompa P, et al. Phosphorylation of MAP65-1 by Arabidopsis Aurora kinases is required for efficient cell cycle progression. Plant Physiol. 2017:173(1):582–599. 10.1104/pp.16.01602 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boudaoud A, Burian A, Borowska-Wykręt D, Uyttewaal M, Wrzalik R, Kwiatkowska D, Hamant O. Fibriltool, an ImageJ plug-in to quantify fibrillar structures in raw microscopy images. Nat Protoc. 2014:9(2):457–463. 10.1038/nprot.2014.024 [DOI] [PubMed] [Google Scholar]
- Buschmann H, Chan J, Sanchez-Pulido L, Andrade-Navarro MA, Doonan JH, Lloyd CW. Microtubule-associated AIR9 recognizes the cortical division site at preprophase and cell-plate insertion. Curr Biol. 2006:16(19):1938–1943. 10.1016/j.cub.2006.08.028 [DOI] [PubMed] [Google Scholar]
- Buschmann H, Dols J, Kopischke S, Peña EJ, Andrade-Navarro MA, Heinlein M, Szymanski DB, Zachgo S, Doonan JH, Lloyd CW. Arabidopsis KCBP interacts with AIR9 but stays in the cortical division zone throughout mitosis via its MyTH4-FERM domain. J Cell Sci. 2015:128(11):2033–2046. 10.1242/jcs.156570 [DOI] [PubMed] [Google Scholar]
- Busson S, Dujardin D, Moreau A, Dompierre J, De Mey JR. Dynein and dynactin are localized to astral microtubules and at cortical sites in mitotic epithelial cells. Curr Biol. 1998:8(9):541–544. 10.1016/S0960-9822(98)70208-8 [DOI] [PubMed] [Google Scholar]
- Chan J, Calder G, Fox S, Lloyd C. Localization of the microtubule end binding protein EB1 reveals alternative pathways of spindle development in Arabidopsis suspension cells. Plant Cell. 2005:17(6):1737–1748. 10.1105/tpc.105.032615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chugh M, Reißner M, Bugiel M, Lipka E, Herrmann A, Roy B, Müller S, Schäffer E. Phragmoplast orienting kinesin 2 is a weak motor switching between processive and diffusive modes. Biophys J. 2018:115(2):375–385. 10.1016/j.bpj.2018.06.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cleary AL, Smith LG. The Tangled1 gene is required for spatial control of cytoskeletal arrays associated with cell division during maize leaf development. Plant Cell. 1998:10(11):1875–1888. 10.1105/tpc.10.11.1875 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cutler SR, Ehrhardt DW. Polarized cytokinesis in vacuolate cells of Arabidopsis. Proc Natl Acad Sci U S A. 2002:99(5):2812–2817. 10.1073/pnas.052712299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doube M, Kłosowski MM, Arganda-Carreras I, Cordelières FP, Dougherty RP, Jackson JS, Schmid B, Hutchinson JR, Shefelbine SJ. Bonej: free and extensible bone image analysis in ImageJ. Bone. 2010:47(6):1076–1079. 10.1016/j.bone.2010.08.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fischer U, Kucukoglu M, Helariutta Y, Bhalerao RP. The dynamics of cambial stem cell activity. Annu Rev Plant Biol. 2019:70(1):293–319. 10.1146/annurev-arplant-050718-100402 [DOI] [PubMed] [Google Scholar]
- Flanders DJ, Rawlins DJ, Shaw PJ, Lloyd CW. Re-establishment of the interphase microtubule array in vacuolated plant cells, studied by confocal microscopy and 3-D imaging. Development. 1990:110(3):897–904. 10.1242/dev.110.3.897 [DOI] [Google Scholar]
- Gunning BE, Hardham AR, Hughes JE. Evidence for initiation of microtubules in discrete regions of the cell cortex in Azolla root-tip cells, and an hypothesis on the development of cortical arrays of microtubules. Planta. 1978:143(2):161–179. 10.1007/BF00387788 [DOI] [PubMed] [Google Scholar]
- Hamada T, Igarashi H, Itoh TJ, Shimmen T, Sonobe S. Characterization of a 200 kDa microtubule-associated protein of tobacco BY-2 cells, a member of the XMAP215/MOR1 family. Plant Cell Physiol. 2004:45(9):1233–1242. 10.1093/pcp/pch145 [DOI] [PubMed] [Google Scholar]
- Hendricks AG, Lazarus JE, Perlson E, Gardner MK, Odde DJ, Goldman YE, Holzbaur ELF. Dynein tethers and stabilizes dynamic microtubule plus ends. Curr Biol. 2012:22(7):632–637. 10.1016/j.cub.2012.02.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrmann A, Livanos P, Lipka E, Gadeyne A, Hauser M-T, Van Damme D, Müller S. Dual localized kinesin-12 POK2 plays multiple roles during cell division and interacts with MAP65-3. EMBO Rep. 2018:19(9):e46085. 10.15252/embr.201846085 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hotta T, Kong Z, Ho C-MK, Zeng CJT, Horio T, Fong S, Vuong T, Lee Y-RJ, Liu B. Characterization of the Arabidopsis augmin complex uncovers its critical function in the assembly of the acentrosomal spindle and phragmoplast microtubule arrays. Plant Cell. 2012:24(4):1494–1509. 10.1105/tpc.112.096610 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kajala K, Ramakrishna P, Fisher A, Bergmann DC, De Smet I, Sozzani R, Weijers D, Brady SM. Omics and modelling approaches for understanding regulation of asymmetric cell divisions in Arabidopsis and other angiosperm plants. Ann Bot. 2014:113(7):1083–1105. 10.1093/aob/mcu065 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawamura E, Himmelspach R, Rashbrooke MC, Whittington AT, Gale KR, Collings DA, Wasteneys GO. MICROTUBULE ORGANIZATION 1 regulates structure and function of microtubule arrays during mitosis and cytokinesis in the Arabidopsis root. Plant Physiol. 2006:140(1):102–114. 10.1104/pp.105.069989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong Z, Hotta T, Lee YR, Horio T, Liu B. The {gamma}-tubulin complex protein GCP4 is required for organizing functional microtubule arrays in Arabidopsis thaliana. Plant Cell. 2010:22(1):191–204. 10.1105/tpc.109.071191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laan L, Pavin N, Husson J, Romet-Lemonne G, van Duijn M, López MP, Vale RD, Jülicher F, Reck-Peterson SL, Dogterom M. Cortical dynein controls microtubule dynamics to generate pulling forces that position microtubule asters. Cell. 2012:148(3):502–514. 10.1016/j.cell.2012.01.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Y-RJ, Hiwatashi Y, Hotta T, Xie T, Doonan JH, Liu B. The mitotic function of augmin is dependent on its microtubule-associated protein subunit EDE1 in Arabidopsis thaliana. Curr Biol. 2017:27(24):3891–3897.e4. 10.1016/j.cub.2017.11.030 [DOI] [PubMed] [Google Scholar]
- Lee Y-RJ, Liu B. The rise and fall of the phragmoplast microtubule array. Curr Opin Plant Biol. 2013:16(6):757–763. 10.1016/j.pbi.2013.10.008 [DOI] [PubMed] [Google Scholar]
- Lee Y-RJ, Liu B. Microtubule nucleation for the assembly of acentrosomal microtubule arrays in plant cells. New Phytol. 2019:222(4):1705–1718. 10.1111/nph.15705 [DOI] [PubMed] [Google Scholar]
- Li H, Sun B, Sasabe M, Deng X, Machida Y, Lin H, Julie Lee Y-R, Liu B. Arabidopsis MAP65-4 plays a role in phragmoplast microtubule organization and marks the cortical cell division site. New Phytol. 2017:215(1):187–201. 10.1111/nph.14532 [DOI] [PubMed] [Google Scholar]
- Lipka E, Gadeyne A, Stöckle D, Zimmermann S, De Jaeger G, Ehrhardt DW, Kirik V, Van Damme D, Müller S. The phragmoplast-orienting kinesin-12 class proteins translate the positional information of the preprophase band to establish the cortical division zone in Arabidopsis thaliana. Plant Cell. 2014:26(6):2617–2632. 10.1105/tpc.114.124933 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu B, Joshi HC, Palevitz BA. Experimental manipulation of gamma-tubulin distribution in Arabidopsis using anti-microtubule drugs. Cell Motil Cytoskeleton. 1995:31(2):113–129. 10.1002/cm.970310204 [DOI] [PubMed] [Google Scholar]
- Livanos P, Müller S. Division plane establishment and cytokinesis. Annu Rev Plant Biol. 2019:70(1):239–267. 10.1146/annurev-arplant-050718-100444 [DOI] [PubMed] [Google Scholar]
- Lough KJ, Byrd KM, Descovich CP, Spitzer DC, Bergman AJ, Beaudoin GM, Reichardt LF, Williams SE. Telophase correction refines division orientation in stratified epithelia. Elife. 2019:8:e49249. 10.7554/eLife.49249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lucas JR. Appearance of microtubules at the cytokinesis to interphase transition in Arabidopsis thaliana. Cytoskeleton. 2021:78(7):361–371. 10.1002/cm.21689 [DOI] [PubMed] [Google Scholar]
- Lucas JR, Sack FD. Polar development of preprophase bands and cell plates in the Arabidopsis leaf epidermis. Plant J. 2012:69(3):501–509. 10.1111/j.1365-313X.2011.04809.x [DOI] [PubMed] [Google Scholar]
- Martinez P, Dixit R, Balkunde RS, Zhang A, O′Leary SE, Brakke KA, Rasmussen CG. TANGLED1 mediates microtubule interactions that may promote division plane positioning in maize. J Cell Biol. 2020:219(8):e201907184. 10.1083/jcb.201907184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinez P, Luo A, Sylvester A, Rasmussen CG. Proper division plane orientation and mitotic progression together allow normal growth of maize. Proc Natl Acad Sci U S A. 2017:114(10):2759–2764. 10.1073/pnas.1619252114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miki T, Naito H, Nishina M, Goshima G. Endogenous localizome identifies 43 mitotic kinesins in a plant cell. Proc Natl Acad Sci U S A. 2014:111(11):E1053–E1061. 10.1073/pnas.1311243111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mills AM, Morris VH, Rasmussen CG. Localization of PHRAGMOPLAST ORIENTING KINESIN1 at the division site depends on the microtubule binding proteins TANGLED1 and AUXIN-INDUCED IN ROOT CULTURES9 in Arabidopsis. Plant Cell. 2022:34(11):4583–4599. 10.1093/plcell/koac266 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohanty A, Luo A, DeBlasio S, Ling X, Yang Y, Tuthill DE, Williams KE, Hill D, Zadrozny T, Chan A, et al. Advancing cell biology and functional genomics in maize using fluorescent protein-tagged lines. Plant Physiol. 2009:149(2):601–605. 10.1104/pp.108.130146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molchan TM, Valster AH, Hepler PK. Actomyosin promotes cell plate alignment and late lateral expansion in Tradescantia stamen hair cells. Planta. 2002:214(5):683–693. 10.1007/s004250100672 [DOI] [PubMed] [Google Scholar]
- Müller S, Han S, Smith LG. Two kinesins are involved in the spatial control of cytokinesis in Arabidopsis thaliana. Curr Biol. 2006:16(9):888–894. 10.1016/j.cub.2006.03.034 [DOI] [PubMed] [Google Scholar]
- Murata T, Sano T, Sasabe M, Nonaka S, Higashiyama T, Hasezawa S, Machida Y, Hasebe M. Mechanism of microtubule array expansion in the cytokinetic phragmoplast. Nat Commun. 2013:4(1):1967. 10.1038/ncomms2967 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamura M, Ehrhardt DW, Hashimoto T. Microtubule and katanin-dependent dynamics of microtubule nucleation complexes in the acentrosomal Arabidopsis cortical array. Nat Cell Biol. 2010:12(11):1064–1070. 10.1038/ncb2110 [DOI] [PubMed] [Google Scholar]
- Nakaoka Y, Miki T, Fujioka R, Uehara R, Tomioka A, Obuse C, Kubo M, Hiwatashi Y, Goshima G. An inducible RNA interference system in Physcomitrella patens reveals a dominant role of augmin in phragmoplast microtubule generation. Plant Cell. 2012:24(4):1478–1493. 10.1105/tpc.112.098509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nan Q, Liang H, Mendoza J, Liu L, Fulzele A, Wright A, Bennett EJ, Rasmussen CG, Facette MR. The OPAQUE1/DISCORDIA2 myosin XI is required for phragmoplast guidance during asymmetric cell division in maize. bioRxiv 458084. 10.1101/2021.08.29.458084, 4 September2021, preprint: not peer reviewed. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otegui M, Staehelin LA. Cytokinesis in flowering plants: more than one way to divide a cell. Curr Opin Plant Biol. 2000:3(6):493–502. 10.1016/S1369-5266(00)00119-9 [DOI] [PubMed] [Google Scholar]
- Panteris E, Adamakis I-DS, Voulgari G, Papadopoulou G. A role for katanin in plant cell division: microtubule organization in dividing root cells of fra2 and lue1 Arabidopsis thaliana mutants. Cytoskeleton. 2011:68(7):401–413. 10.1002/cm.20522 [DOI] [PubMed] [Google Scholar]
- Panteris E, Apostolakos P, Galatis B. Telophase-interphase transition in taxol-treated Triticum root cells: cortical microtubules appear without the prior presence of a radial perinuclear array. Protoplasma. 1995:188(1–2):78–84. 10.1007/BF01276798 [DOI] [Google Scholar]
- Rasmussen CG, Bellinger M. An overview of plant division-plane orientation. New Phytol. 2018:219:505–512. 10.1111/nph.15183 [DOI] [PubMed] [Google Scholar]
- Rasmussen CG. Using live-cell markers in maize to analyze cell division orientation and timing, eds, Plant cell division: Methods and Protocols. Vol. 1370. New York (NY): Humana Press; 2016. p. 209–225. 10.1111/nph.15183 [DOI] [PubMed] [Google Scholar]
- Rasmussen CG, Sun B, Smith LG. Tangled localization at the cortical division site of plant cells occurs by several mechanisms. J Cell Sci. 2011:124(2):270–279. 10.1242/jcs.073676 [DOI] [PubMed] [Google Scholar]
- R Core Team . R: a language and environment for statistical computing. Vienna: R Core Team; 2013. [Google Scholar]
- Sasabe M, Machida Y. Regulation of organization and function of microtubules by the mitogen-activated protein kinase cascade during plant cytokinesis. Cytoskeleton. 2012:69(11):913–918. 10.1002/cm.21072 [DOI] [PubMed] [Google Scholar]
- Sasabe M, Soyano T, Takahashi Y, Sonobe S, Igarashi H, Itoh TJ, Hidaka M, Machida Y. Phosphorylation of NtMAP65-1 by a MAP kinase down-regulates its activity of microtubule bundling and stimulates progression of cytokinesis of tobacco cells. Genes Dev. 2006:20(8):1004–1014. 10.1101/gad.1408106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasaki T, Tsutsumi M, Otomo K, Murata T, Yagi N, Nakamura M, Nemoto T, Hasebe M, Oda Y. A novel katanin-tethering machinery accelerates cytokinesis. Curr Biol. 2019:29(23):4060–4070.e3. 10.1016/j.cub.2019.09.049 [DOI] [PubMed] [Google Scholar]
- Schmidt S, Smertenko A. Identification and characterization of the land-plant-specific microtubule nucleation factor MACET4. J Cell Sci. 2019:132(11):jcs232819. 10.1242/jcs.232819 [DOI] [PubMed] [Google Scholar]
- Smertenko A, Assaad F, Baluška F, Bezanilla M, Buschmann H, Drakakaki G, Hauser M-T, Janson M, Mineyuki Y, Moore I, et al. Plant cytokinesis: terminology for structures and processes. Trends Cell Biol. 2017:27(12):885–894. 10.1016/j.tcb.2017.08.008 [DOI] [PubMed] [Google Scholar]
- Thévenaz P. StackReg: an ImageJ plugin for the recursive alignment of a stack of images. Vol. 2012. Lausanne: Biomedical Imaging Group, Swiss Federal Institute of Technology; 1998. [Google Scholar]
- Van Damme D, Geelen D. Demarcation of the cortical division zone in dividing plant cells. Cell Biol Int. 2008:32(2):178–187. 10.1016/j.cellbi.2007.10.010 [DOI] [PubMed] [Google Scholar]
- van Oostende-Triplet C, Guillet D, Triplet T, Pandzic E, Wiseman PW, Geitmann A. Vesicle dynamics during plant cell cytokinesis reveals distinct developmental phases. Plant Physiol. 2017:174(3):1544–1558. 10.1104/pp.17.00343 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vavrdová T, Šamajová O, Křenek P, Ovečka M, Floková P, Šnaurová R, Šamaj J, Komis G. Multicolour three dimensional structured illumination microscopy of immunolabeled plant microtubules and associated proteins. Plant Methods. 2019:15(1):22. 10.1186/s13007-019-0406-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venverloo CJ, Libbenga KR. Regulation of the plane of cell division in vacuolated cells I. The function of nuclear positioning and phragmosome formation. J Plant Physiol. 1987:131(3–4):267–284. 10.1016/S0176-1617(87)80166-9 [DOI] [Google Scholar]
- Walker KL, Müller S, Moss D, Ehrhardt DW, Smith LG. Arabidopsis TANGLED identifies the division plane throughout mitosis and cytokinesis. Curr Biol. 2007:17(21):1827–1836. 10.1016/j.cub.2007.09.063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wasteneys GO, Williamson RE. Reassembly of microtubules in Nitella tasmanica: assembly of cortical microtubules in branching clusters and its relevance to steady-state microtubule assembly. J Cell Sci. 1989:93(4):705–714. 10.1242/jcs.93.4.705 [DOI] [Google Scholar]
- Wick SM. Immunofluorescence microscopy of tubulin and microtubule arrays in plant cells. III. Transition between mitotic/cytokinetic and interphase microtubule arrays. Cell Biol Int Rep. 1985:9(4):357–371. 10.1016/0309-1651(85)90031-1 [DOI] [PubMed] [Google Scholar]
- Wickham H, Chang W, et al. ggplot2: an implementation of the grammar of graphics. R package version 0.7. http://CRAN.R-project.org/package=ggplot2. 2008. 3 [Google Scholar]
- Wu S-Z, Bezanilla M. Myosin VIII associates with microtubule ends and together with actin plays a role in guiding plant cell division. Elife. 2014:3:e03498. 10.7554/eLife.03498 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Q, Luo A, Zadrozny T, Sylvester A, Jackson D. Fluorescent protein marker lines in maize: generation and applications. Int J Dev Biol. 2013:57(6–8):535–543. 10.1387/ijdb.130240qw [DOI] [PubMed] [Google Scholar]
- Yamada M, Tanaka-Takiguchi Y, Hayashi M, Nishina M, Goshima G. Multiple kinesin-14 family members drive microtubule minus end–directed transport in plant cells. J Cell Biol. 2017:216(6):1705–1714. 10.1083/jcb.201610065 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yasuhara H, Muraoka M, Shogaki H, Mori H, Sonobe S. TMBP200, a microtubule bundling polypeptide isolated from telophase tobacco BY-2 cells is a MOR1 homologue. Plant Cell Physiol. 2002:43(6):595–603. 10.1093/pcp/pcf074 [DOI] [PubMed] [Google Scholar]
- Yoneda A, Akatsuka M, Kumagai F, Hasezawa S. Disruption of actin microfilaments causes cortical microtubule disorganization and extra-phragmoplast formation at M/G1 interface in synchronized tobacco cells. Plant Cell Physiol. 2004:45(6):761–769. 10.1093/pcp/pch091 [DOI] [PubMed] [Google Scholar]
- Young DH, Lewandowski VT. Covalent binding of the benzamide RH-4032 to tubulin in suspension-cultured tobacco cells and its application in a cell-based competitive-binding assay. Plant Physiol. 2000:124(1):115–124. 10.1104/pp.124.1.115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou R, Liu H, Ju T, Dixit R. Quantifying the polymerization dynamics of plant cortical microtubules using kymograph analysis. Methods Cell Biol. 2020:160:281–293. 10.1016/bs.mcb.2020.04.006 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Maize lines are available at the Maize Cooperative (http://maizecoop.cropsci.uiuc.edu/) or upon request.