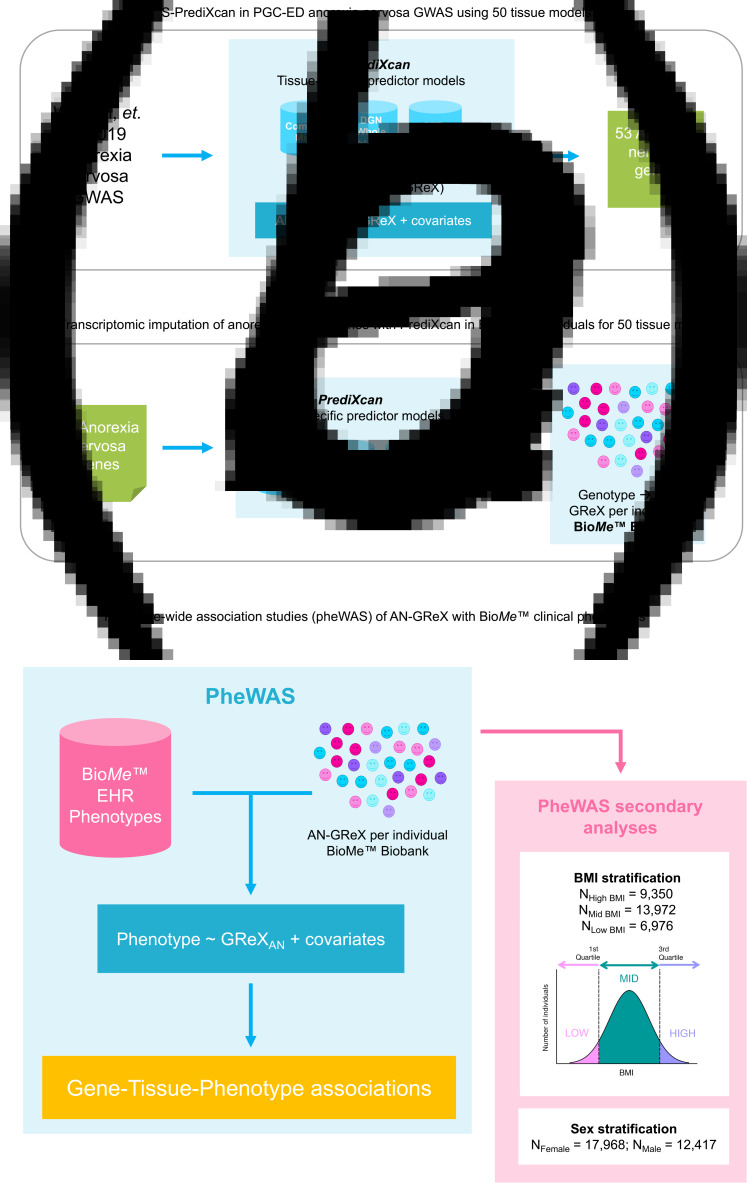
Fig. 1.
Graphical depiction of S-PrediXcan and PrediXcan TI and pheWAS analyses. (a) We used S-PrediXcan predictor models for 50 different tissue types to impute GReX in Watson et al. (2019) AN GWAS and found 53 genes whose GReX was associated with AN. We then (b) imputed GReX for those 53 AN-genes in our BioMe™ cohort and (c) performed a pheWAS across available EHR phenotypes. The pheWAS analyses were run within each ancestral population and then meta-analyzed using an inverse-variance approach in METAL. Secondary analyses included stratifying individuals in BioMe™ by BMI and sex, and running the pheWAS analyses within each stratification group.