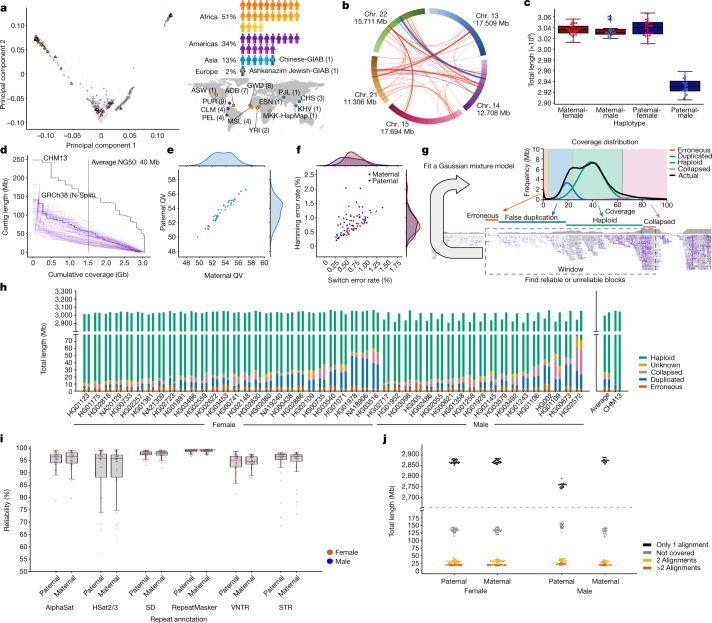
Fig. 1. Presenting 47 accurate and near-complete diverse diploid human genome assemblies.
a, Selecting the HPRC samples. Left, the first two principal components of 1KG samples showing HPRC (triangles) samples, excluding HG002, HG005 and NA21309. Right, summary of the HPRC sample subpopulations (three letter abbreviations) on a map of Earth as defined by the 1KG. ACB, African Caribbean in Barbados; ASW, African Ancestry in Southwest US; CHS, Han Chinese South; CLM, Colombian in Medellin, Colombia; ESN, Esan in Nigeria; GWD, Gambian in Western Division; KHV, Kinh in Ho Chi Minh City, Vietnam; MKK, Maasai in Kinyawa, Kenya; MSL, Mende in Sierra Leone; PEL, Peruvian in Lima, Peru; PJL, Punjabi in Lahore, Pakistan; PUR, Puerto Rican in Puerto Rico; YRI, Yoruba in Ibadan, Nigeria. b, Interchromosomal joins between acrocentric chromosome short arms. Red, the join is on the same strand; blue, otherwise. c, Total assembled sequence per haploid phased assembly. d, Assembly contiguity shown as a NGx plot. T2T-CHM13 and GRCh38 contigs are included for comparison. e, Assembly QVs showing the base-level accuracy of the maternal and paternal assembly for each sample. f, Yak-reported phasing accuracy showing the switch error percentage versus Hamming error percentage. g, Flagger read-based assembly evaluation pipeline. Coverage is calculated across the genome and a mixture model is fit to account for reliably assembled haploid sequence and various classes of unreliably assembled sequence. For each coverage block, a label is assigned according to the most probable mixture component to which it belongs: erroneous, falsely duplicated, (reliable) haploid, collapsed, and unknown. h, Reliability of the 47 HPRC assemblies using read mapping. For each sample, the left bar is the paternal and the right bar is the maternal haplotype. Regions flagged as haploid are reliable (green), constituting more than 99% on average of each assembly. The y axis is broken to show the dominance of the reliable haploid component and the stratification of the unreliable blocks. i, Assembly reliability of six types of repeats. AlphaSat, alpha satellites; HSat2/3, human satellites 2 and 3. j, Completeness of the HPRC assemblies relative to T2T-CHM13. The number of reference bases covered by none, by one, by two or by more than two alignments are included.