Dear Editor,
The ion channel transient receptor potential melastatin-like 7 (TRPM7) has a serine-threonine α-kinase domain (M7CK) on its intracellular C-terminal [1, 2]. In cell lines, M7CK is cleaved and translocated to the nucleus to regulate a variety of cellular processes including cell proliferation and survival [3]. In neuroblastoma cells, M7CK interacts with several cytoskeleton-regulating proteins including cofilin [4]. In the mammalian brain, M7CK regulates synaptic density, plasticity, and learning and memory. Evidence suggests that M7CK protects synaptic and cognitive functions by interacting with, and phosphorylating (inhibiting), cofilin in the brain [5]. It remains unclear how M7CK regulates cofilin activity and whether such regulation might result in changes in actin filaments (F-actin) in neural systems. Ras-related C3 botulinum toxin substrate 1 (Rac1) signaling is a key regulator of cytoskeleton dynamics. Rac1-PAK1 (P21 activated kinase 1)-LIMK1 (LIM domain kinase 1) signaling protects synapse structure/remodeling/plasticity by phosphorylating cofilin, an actin depolymerization factor [6, 7]. Here, we found that M7CK is a key member of a novel protein complex composed of Rac1-M7CK-SSH2 (protein phosphatase slingshot homolog 2)-cofilin that regulates cofilin activity and dendritic F-actin in the mouse nervous system.
We generated transgenic mice with brain-specific deletion of TRPM7 in CaMKIIα-positive glutamatergic neurons (CaMKII-T7−/−). By crossing CaMKII-T7−/− mice with Ai3 mice (a Cre-reporter strain, Fig. S1A) and by using immunostaining of TRPM7 using a commercially-available antibody targeting the ion channel part of TRPM7 (see Supplementary Materials and Methods and Fig. S1B for antibody validation), we confirmed that TRPM7 was knocked out in the majority of the glutamatergic neurons (recombination efficiency ~91.5%, Fig. S1A). As a result, quantitative Western blot analysis of TRPM7 protein in homogenized brain tissue using the same antibody showed that the protein was reduced by ~55% in CaMKII-T7−/− mice (two-tailed unpaired t-test, t(10) = 5.113, P <0.001, Fig. 1A, B), which is in line with our previous results [5]. A polyclonal antibody targeting the kinase domain of TRPM7 (amino-acid residues 1300–1539 of mouse TRPM7) was generated (anti-M7CK antibody, see Supplementary Materials and Methods) and validated (Fig. S1C). Using this antibody, we confirmed that M7CK was also significantly reduced in CaMKII-T7−/− mice (two-tailed unpaired t-test, Welch-corrected t(5.24) = 2.689, P = 0.041, Fig. 1A, C). Furthermore, we used the co-detected full-length TRPM7 by anti-M7CK antibody to calculate the M7CK/TRPM7 ratio in Trpm7flox/flox and CaMKII-T7−/− mice in order to check if TRPM7 conditional knockout in glutamatergic neurons influenced the cleavage rate of M7CK in the brain. We found no significant differences between Trpm7flox/flox and CaMKII-T7−/− mice (two-tailed unpaired t-test, t(10) = 0.1522, P = 0.882, Fig. 1A, D). Deletion of TRPM7 reduces cofilin phosphorylation by ~80% without changing LIMK1 or PAK1 activity [5]. In line with these findings, we found that cofilin phosphorylation was reduced by ~70% in CaMKII-T7−/− mice (Mann-Whitney test, U = 5, P = 0.041, Fig. 1A, E, see also Fig. S2A). Next, we tested whether the deletion of TRPM7 affected the activity of other proteins known to regulate cofilin activity. The SSH2 regulates actin filament dynamics by dephosphorylating (activating) cofilin [8]. Phosphorylation of SSH2 (pSSH2) at serine residues S21 and S32 inhibits the phosphatase activity of SSH2 [9]. We quantified the phosphorylation level of SSH2 and found that CaMKII-T7−/− mice had significantly lower levels of pSSH2 at S21 (two-tailed unpaired t-test, Welch-corrected t(5.045) = 4.311, P = 0.007, Fig. 1A, F, see also Fig. S2B) and S32 (two-tailed unpaired t-test, Welch-corrected t(5.359) = 3.039, P = 0.012, Fig. 1A, G, see also Fig. S2C).
Fig. 1.
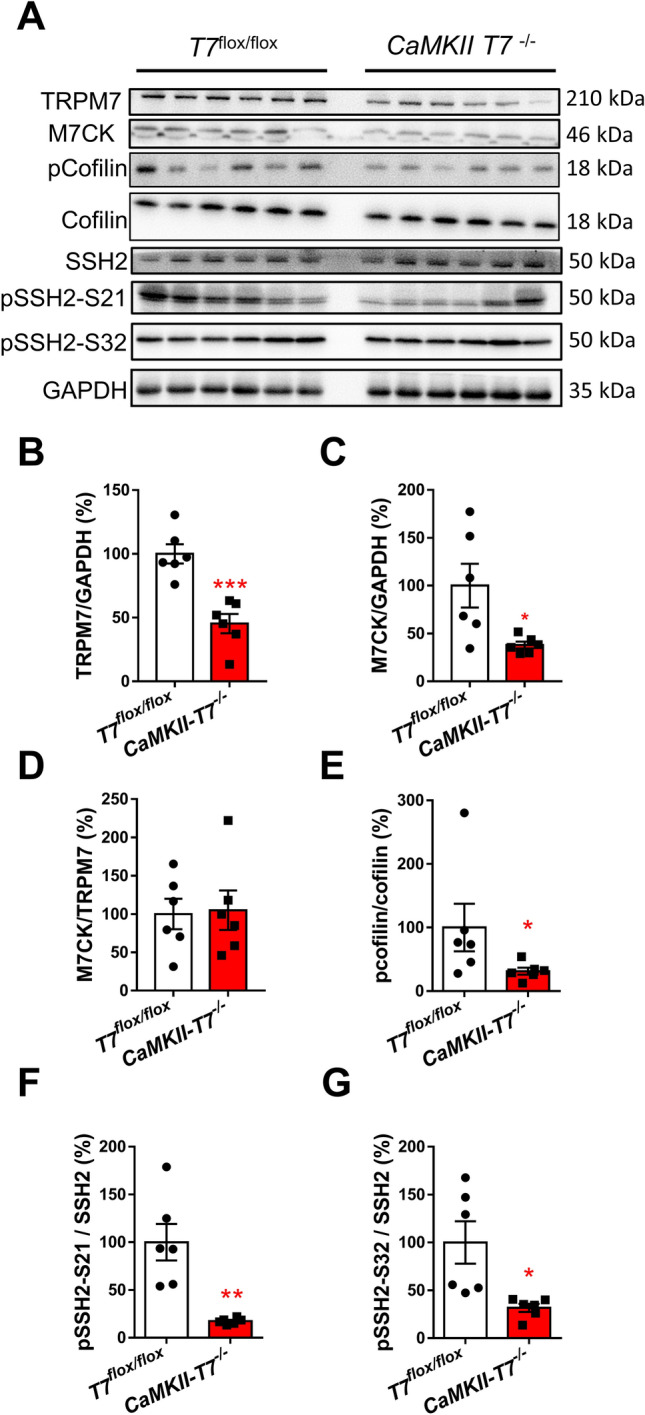
The phosphorylation levels of cofilin and SSH2 in the hippocampus of TRPM7-knockout mice. A Western blot images of protein bands from all the mice that were used for analysis. The following proteins are detected in TRPM7flox/flox or CaMKII-T7−/− mice: TRPM7, M7CK, cofilin, pcofilin, SSH2, pSSH2-S21, and pSSH2-S32. B–G Analysis of the expression levels of TRPM7 (B) and M7CK (C), the M7CK/TRPM7 ratio (D), and the phosphorylation levels of cofilin (E), pSSH2-S21 (F), and pSSH2-S32 (G) calculated as a ratio of the total corresponding protein and presented as percentages of the control TRPM7flox/flox. The co-detection of GAPDH bands serves as the loading control. For B–G, data are presented as the mean ± SEM, n = 6 per group, two-tailed unpaired t-test (B, C, D, F, G), Mann-Whitney test (E), *P <0.05; **P <0.01. ***P <0.001.
We speculated that M7CK might regulate cofilin activity directly by phosphorylating it and indirectly by preventing its dephosphorylation via inhibiting SSH2. If so, then M7CK should interact with, and phosphorylate both proteins. We used the anti-M7CK antibody to perform all co-immunoprecipitation and Western blotting experiments on the kinase domain of TRPM7. We found that the cleaved kinase domain was co-immunoprecipitated with both cofilin and SSH2 in human embryonic kidney (HEK) cells (Fig. 2A). No actin was detected in this complex (Fig. S3A), suggesting that the M7CK-Cofilin-SSH2 interaction is specific and not indirectly induced by actin-linking. We also tested whether M7CK can be detected in the proximity of cofilin and SSH2 in neuronal cells using co-immunostaining. In hippocampal neuronal cultures, we found that M7CK puncta co-localized with cofilin (Fig. S3B) and SSH2 (Fig. S3C) puncta in different areas within the cytoplasm. We examined whether SSH2 and cofilin could be phosphorylated by M7CK. We conducted an enzymatic kinase activity assay using cofilin and/or SSH2 as substrates for M7CK. Negative controls (no M7CK and/or no substrate was added), as well as a positive control (using myelin basic protein, a well-known substrate detecting M7CK activity [10]), were included in those experiments (Fig. 2B). We found that the signal of kinase activity was significantly increased when myelin basic protein, SSH2, or cofilin was used as the substrate (one-way ANOVA, F(3, 20) = 14.86, P <0.001, Fig. 2B). Bonferroni’s post hoc test showed that the signals in +SSH2 (P = 0.032), +cofilin (P <0.001), and +myelin basic protein (MBP, P <0.001) reactions were significantly higher than that in the control reaction containing M7CK but no substrate (Fig. 2B).
Fig. 2.
M7CK plays a key role in the Rac1-M7CK-SSH2-cofilin signaling pathway. A Co-immunoprecipitation of SSH2, cofilin, and M7CK by using an anti-M7CK antibody (targeting the kinase domain). Beads and IgG are loaded and used as control. B Analysis of M7CK kinase activity (expressed as the ratio of the control group) without substrate (negative control), or after adding SSH2, cofilin, and/or MBP (serves as a positive control) as substrates. Negative control (no M7CK and no substrate are added) is also presented (but not included in the statistical analysis). n = 6 per group. C Co-immunoprecipitation of Rac1 and M7CK using anti-M7CK antibody; beads and IgG are loaded and used as control. D Analysis of Rac1 activity (expressed as the ratio of the control group) without substrate (negative control), or after adding SSH2 and/or M7CK as substrates. The Rac1 + M7CK reaction activity is diminished by adding the Rac1 activity-neutralizing antibody. Negative control (no Rac1 and no substrate) is also presented (but not included in the statistical analysis). n = 6 per group. E Left: Western blot images of protein bands from hippocampal neuronal cultures treated with solvent (Control) or NSC23766 (Rac1 inhibitor, n = 6 per group). Right: Analysis of the phosphorylation level of M7CK or SSH2 (at serine 21, S21) calculated as a ratio of the total corresponding protein and presented as percentages of the Control. The co-detection of GAPDH bands serves as a loading control. F Left: Representative fluorescent images of dendrites stained with phalloidin in primary cell cultures transfected with AAV-Controlscrambled-shRNA-tdTomato virus (ControlSCR) or AAV-Trpm7-shRNA-tdTomato virus (Trpm7shRNA). Below, a representative image of RT-PCR confirms the knockdown of Trpm7 mRNA. Right: Analysis of dendritic/synaptic phalloidin puncta per 100 μm dendrite. n = 6 independent cultures per group. G Representative fluorescent images of hippocampal CA1 apical dendrites in the stratum radiatum stained with phalloidin in brain sections from Trpm7flox/flox and CaMKII-T7−/− mice. Right: Analysis of dendritic phalloidin puncta per 1000 μm2 area. n = 7 per group. H Left: Representative fluorescent images of dendrites stained with phalloidin Trpm7shRNA transduced cultures following knock-in of EGFP (control), TRPM7ΔK-EGFP, or M7CK-EGFP. Right: Analysis of dendritic/synaptic phalloidin puncta per 100 μm dendrite showing that the kinase domain, but not the ion channel part, rescues phalloidin puncta in neuronal cultures following knockdown of TRPM7 (the ControlSCR group is the same as in F; it is presented for comparison purposes). n = 6 independent cultures per group. All phalloidin puncta are detected at 647 nm and presented in green for clarity. Data are presented as the mean ± SEM, one-way ANOVA followed by Bonferroni post hoc test (B, D, H), two-tailed unpaired t-test (E, F), Mann-Whitney test (G), *P <0.05; **P <0.01; ***P <0.001.
Rac1 signaling regulates spine remodeling, synapse density, and plasticity by regulating cofilin activity and F-actin dynamics [6, 11, 12]. We checked if Rac1 interacted with M7CK and whether the kinase might be a phosphorylation target for Rac1. Interestingly, we found that M7CK co-immunoprecipitated with Rac1 in HEK cells (Fig. 2C). Furthermore, enzyme activity assays showed a significant increase in Rac1 activity after adding M7CK but not SSH2 as the substrate (one-way ANOVA: F(3, 20) = 421.8, P <0.001; Bonferroni’s post hoc test, compared with Rac1 with no substrate control group: +SSH2, P >0.999, +M7CK, P <0.001, Fig. 2D). Thus, M7CK appears to be a phosphorylation target of Rac1. When Rac1 activity-neutralizing antibody was added, the activity in the Rac1-M7CK reaction was reduced to levels similar to those measured in the blank negative control (Bonferroni’s post hoc test, compared with Rac1+M7CK: M7CK+Rac1+Rac1 neutralizing antibody, P <0.001, Fig. 2D) indicating that the activity was due to Rac1 activity and/or the phosphorylation of M7CK by Rac1, not the opposite. We also found that inhibition of Rac1 in neuronal cell cultures resulted in reductions in the phosphorylation levels of M7CK (two-tailed unpaired t-test, t(10) = 2.957, P = 0.014, Fig. 2E) and SSH2 (two-tailed unpaired t-test, t(10) = 3.73, P = 0.004, Fig. 2E).
Finally, to demonstrate the functional effects of this new complex on F-actin, we quantified F-actin puncta within dendritic areas following the suppression and/or deletion of TRPM7 and its kinase in vitro and/or in vivo (respectively). We used phalloidin staining, a well-known molecule that binds to F-actin colocalized with synaptic proteins in the dendritic spines of excitatory synapses [13]. In hippocampal neuronal cultures, we found that knockdown of TRPM7 by shRNA (Trpm7shRNA) resulted in a significant reduction in the numbers of F-actin puncta on the dendrites (two-tailed unpaired t-test, t(10) = 2.472, P = 0.033, Fig. 2F). In brain sections, we found that CaMKII-T7−/− mice had a significantly lower density of F-actin puncta in hippocampal CA1 dendritic areas (Mann-Whitney test, U = 7, P = 0.026, Fig. 2G). Thus, deletion or suppression of TRPM7 (ion channel and the kinase domain) resulted in reductions in F-actin in dendritic areas. To investigate which part of TRPM7 can rescue F-actin puncta, we overexpressed the kinase domain (M7CK-EGFP) or a truncated ion channel (at D1510, TRPM7ΔK-EGFP that has been shown to be functional [14]) in neuronal cultures following knockdown of TRPM7. We found that expression of the kinase domain, but not the truncated ion channel, was sufficient to significantly increase the F-actin puncta in dendritic areas (one-way ANOVA: F(3, 20) = 18.67, P <0.001; Bonferroni’s post hoc test, in comparison with TRPM7shRNA EGFP control: M7CK-EGFP, P = 0.007, TRPM7ΔK-EGFP, P = 0.678, Fig. 2H). It is worth noting that complexes of cofilin and actin (known as cofilactin) are prominent in neuronal cells. Such complexes are not easily immunolabeled for cofilin or phalloidin after Triton permeabilization [15, 16]. Thus, we cannot exclude the possibility that changes in cofilactin abundance induced by TRPM7 suppression/deletion might contribute to the decline in phalloidin puncta.
In this study, we discovered a novel signaling pathway; namely Rac1-M7CK-SSH2-cofilin and found that it plays a key role in controlling cofilin activity and regulating F-actin in the nervous system (see schematic illustration, Fig. S4). Furthermore, we found that M7CK is a central component within this novel pathway. Suppression or deletion of the kinase expression critically activated cofilin and reduced F-actin.
The Rac1-cofilin signaling pathway is important for maintaining the normal F-actin dynamics necessary for cell growth, proliferation, and migration. In the mammalian brain, dysfunction in the Rac1-cofilin signaling pathway results in deficits in learning and memory, impairments in synaptic plasticity and remodeling, and reductions in synapse density [11, 12]. These abnormalities are explained by possible impairment in the Rac1-PAK1-LIMK1-cofilin signaling pathway, as it is considered the major pathway controlling cofilin activity and F-actin dynamics [12]. However, PAK1-knockout mice have no reduction in cofilin phosphorylation [17]. LIMK1-knockout mice have ~50% reduction in cofilin phosphorylation [18]. Meanwhile, Rac1 mutant mice have >70% reduction in cofilin phosphorylation and significant disruption in F-actin dynamics [11]. These studies support the conclusion that Rac1 is a major regulator of cofilin activity and F-actin dynamics, but evidence also suggests that PAK1-LIMK1 might not be the only signaling molecules mediating the effects of Rac1 on cofilin. In the current study, we found that M7CK is a downstream target of Rac1. The kinase inhibits cofilin directly by phosphorylation), and indirectly by preventing its dephosphorylation. Therefore, it is possible that M7CK-SSH2-cofilin is another major signaling pathway used by Rac1 to maintain tight regulation of cofilin activity and F-actin in the nervous system.
Finally, M7CK has been shown to be cleaved from TRPM7 via proteolytic mechanisms in different cell lines and tissues [3]. Studies in the mammalian brain have also shown that the kinase domain alone without the ion channel component is sufficient to maintain normal synaptic and cognitive functions [5] indicating that M7CK is also cleaved in the nervous system. The mechanisms underlying the cleavage and translocation of M7CK as well as the signaling pathways interacting with it in the mammalian brain remain to be determined.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31970942 and 81573408), the Shanghai Municipal Science and Technology Major Project (2018SHZDZX01), the ZJ Lab, and Shanghai Center for Brain Science and Brain-Inspired Technology.
Conflict of interest
The authors declare no conflict of interest related to this work.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Abumaria N, Li W, Clarkson AN. Role of the chanzyme TRPM7 in the nervous system in health and disease. Cell Mol Life Sci. 2019;76:3301–3310. doi: 10.1007/s00018-019-03124-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Abumaria N, Li W, Liu Y. TRPM7 functions in non-neuronal and neuronal systems: Perspectives on its role in the adult brain. Behav Brain Res. 2018;340:81–86. doi: 10.1016/j.bbr.2016.08.038. [DOI] [PubMed] [Google Scholar]
- 3.Krapivinsky G, Krapivinsky L, Manasian Y, Clapham DE. The TRPM7 chanzyme is cleaved to release a chromatin-modifying kinase. Cell. 2014;157:1061–1072. doi: 10.1016/j.cell.2014.03.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Middelbeek J, Vrenken K, Visser D, Lasonder E, Koster J, Jalink K, et al. The TRPM7 interactome defines a cytoskeletal complex linked to neuroblastoma progression. Eur J Cell Biol. 2016;95:465–474. doi: 10.1016/j.ejcb.2016.06.008. [DOI] [PubMed] [Google Scholar]
- 5.Liu Y, Chen C, Liu Y, Li W, Wang Z, Sun Q, et al. TRPM7 is required for normal synapse density, learning, and memory at different developmental stages. Cell Rep. 2018;23:3480–3491. doi: 10.1016/j.celrep.2018.05.069. [DOI] [PubMed] [Google Scholar]
- 6.Li LZ, Yin N, Li XY, Miao Y, Cheng S, Li F, et al. Rac1 modulates excitatory synaptic transmission in mouse retinal ganglion cells. Neurosci Bull. 2019;35:673–687. doi: 10.1007/s12264-019-00353-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Fan XC, Ma CN, Song JC, Liao ZH, Huang N, Liu X, et al. Rac1 signaling in amygdala astrocytes regulates fear memory acquisition and retrieval. Neurosci Bull. 2021;37:947–958. doi: 10.1007/s12264-021-00677-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhang T, Cheng D, Wu C, Wang X, Ke Q, Lou H, et al. Lysosomal hydrolase cathepsin D non-proteolytically modulates dendritic morphology in Drosophila. Neurosci Bull. 2020;36:1147–1157. doi: 10.1007/s12264-020-00479-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tang W, Zhang Y, Xu W, Harden TK, Sondek J, Sun L, et al. A PLCβ/PI3Kγ-GSK3 signaling pathway regulates cofilin phosphatase slingshot2 and neutrophil polarization and chemotaxis. Dev Cell. 2011;21:1038–1050. doi: 10.1016/j.devcel.2011.10.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ryazanova LV, Dorovkov MV, Ansari A, Ryazanov AG. Characterization of the protein kinase activity of TRPM7/ChaK1, a protein kinase fused to the transient receptor potential ion channel. J Biol Chem. 2004;279:3708–3716. doi: 10.1074/jbc.M308820200. [DOI] [PubMed] [Google Scholar]
- 11.Pyronneau A, He Q, Hwang JY, Porch M, Contractor A, Zukin RS. Aberrant Rac1-cofilin signaling mediates defects in dendritic spines, synaptic function, and sensory perception in fragile X syndrome. Sci Signal. 2017;10:eaan0852. doi: 10.1126/scisignal.aan0852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lai KO, Ip NY. Structural plasticity of dendritic spines: The underlying mechanisms and its dysregulation in brain disorders. Biochim Biophys Acta BBA Mol Basis Dis. 2013;1832:2257–2263. doi: 10.1016/j.bbadis.2013.08.012. [DOI] [PubMed] [Google Scholar]
- 13.Park CE, Cho Y, Cho I, Jung H, Kim B, Shin JH, et al. Super-resolution three-dimensional imaging of actin filaments in cultured cells and the brain via expansion microscopy. ACS Nano. 2020;14:14999–15010. doi: 10.1021/acsnano.0c04915. [DOI] [PubMed] [Google Scholar]
- 14.Desai BN, Krapivinsky G, Navarro B, Krapivinsky L, Carter BC, Febvay S, et al. Cleavage of TRPM7 releases the kinase domain from the ion channel and regulates its participation in fas-induced apoptosis. Dev Cell. 2012;22:1149–1162. doi: 10.1016/j.devcel.2012.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.McGough A, Pope B, Chiu W, Weeds A. Cofilin changes the twist of F-actin: Implications for actin filament dynamics and cellular function. J Cell Biol. 1997;138:771–781. doi: 10.1083/jcb.138.4.771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hylton RK, Heebner JE, Grillo MA, Swulius MT. Cofilactin filaments regulate filopodial structure and dynamics in neuronal growth cones. Nat Commun. 2022;13:2439. doi: 10.1038/s41467-022-30116-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Asrar S, Meng Y, Zhou Z, Todorovski Z, Huang WW, Jia Z. Regulation of hippocampal long-term potentiation by p21-activated protein kinase 1 (PAK1) Neuropharmacology. 2009;56:73–80. doi: 10.1016/j.neuropharm.2008.06.055. [DOI] [PubMed] [Google Scholar]
- 18.Meng Y, Zhang Y, Tregoubov V, Janus C, Cruz L, Jackson M, et al. Abnormal spine morphology and enhanced LTP in LIMK-1 knockout mice. Neuron. 2002;35:121–133. doi: 10.1016/S0896-6273(02)00758-4. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.



