Abstract
Background
This study used admixture mapping to prioritize the genetic regions associated with Alzheimer disease (AD) in African American (AA) individuals, followed by ancestry-aware regression analysis to fine-map the prioritized regions.
Methods
We analyzed 10,271 individuals from 17 different AA datasets. We performed admixture mapping and meta-analyzed the results. We then used regression analysis, adjusting for local ancestry main effects and interactions with genotype, to refine the regions identified from admixture mapping. Finally, we leveraged in silico annotation and differential gene expression data to prioritize AD-related variants and genes.
Results
Admixture mapping identified two genome-wide significant loci on chromosomes 17p13.2 (p=2.2 ×10−5) and 18q21.33 (p=1.2×10−5). Our fine mapping of the chromosome 17p13.2 and 18q21.33 regions revealed several interesting genes such as the MINK1, KIF1C, and BCL2.
Discussion
Our ancestry-aware regression approach showed that AA individuals have a lower risk of AD if they inherited African ancestry admixture block at the 17p13.2 locus.
Keywords: ancestry, admixture, admixture mapping, African American, local ancestry, global ancestry
1. BACKGROUND
Alzheimer disease (AD) is the most common form of dementia and is characterized by the loss of memory and other cognitive functions. AD studies showed that disease affects individuals of all races and ethnicities with the higher prevalence and incidence in African American (AA) and Hispanic populations than non-Hispanic whites (NHW)1,2. Large-scale genetic studies have successfully identified more than 30 susceptibility loci for AD3–6. Most of these studies have focused on NHW populations; yet genetic studies in diverse populations have found ethnic-specific variations in several genes influencing disease5,7. The effect of the polymorphisms in the APOE gene on AD is a notable example of how different ethnic backgrounds may lead to differential effects in different populations8. Studies in admixed populations have shown that the effect of the APOE ε4 allele is correlated with the genetic ancestral background and suggested that regions of African ancestry around the APOE gene are likely to harbor protective variant(s) that mitigate the effect of the known APOE risk variant9,10. Another example of ethnic-specific effects is the ABCA7 locus. Genome-wide association studies (GWAS) in AAs found that ABCA7 influences risk in AAs more strongly than in NHW7. Follow-up studies identified a pathogenic 44 bp deletion in ABCA7 associated with AD exclusively in individuals with African ancestral background11. Such findings highlight the need to study diverse populations to understand the differences in genetics effects on disease and discover ethnic-specific variants that would be missed otherwise.
There is an emerging effort to include diverse populations in AD genetic studies; yet the largest AA GWAS to date was performed in case-control study of ~8,500 AA individuals, which is less than one-tenth of the size of the largest GWAS of AD in NHW individuals of ~95,000 individuals3,5. Because of the substantial underrepresentation of AAs in genetic studies of AD relative to NHW, SNP-based GWAS in AAs have reduced statistical power for novel variant discovery and have limited power to assess genetic factors identified in NHW populations.
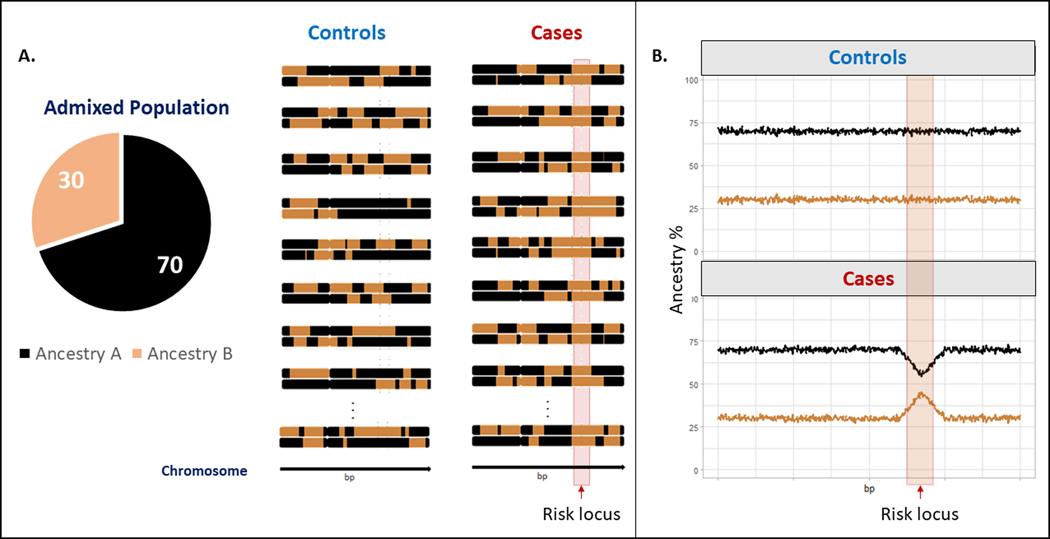
The AA population is a two-way admixed population with genetic ancestry from African and European ancestors. Admixture in AAs creates mosaic chromosomes containing discrete segments of European and African ancestry, referred to as local ancestry. This mosaic pattern varies among the individuals, though the proportion of each ancestry across the genome (global ancestry) tends to be similar. Admixture mapping is a complementary statistical approach to SNP-based GWAS that maps the risk loci in admixed populations by leveraging the local ancestry estimates. This approach is based on the hypothesis that the causal risk alleles with large frequency or effect-size differences between the parental ancestries (African and European for AAs) will result in deviation of the expected local-ancestry proportions at the risk loci. Figure 1A illustrates a two-way admixed population (parental ancestries “A” and “B”) with the excess of “B” local ancestry among cases at the risk locus. This excess results in a spike of “B” ancestry percentage (Figure 1B). Admixture mapping identifies such risk loci by calculating the deviation of the local-ancestry proportions from the genome-wide averages. Moreover, admixture mapping has a lower multiple testing burden than SNP-based GWAS studies, since recent admixture (~20 generations) in AAs created admixture blocks, which are larger than linkage disequilibrium (LD) blocks (Supplemental Figure 1).
Figure 1.
(A) An example of a two-way admixed population (“A” and “B”) with the uniform distribution of the local ancestries in controls and excess of the “B” local ancestry at the risk locus in cases. (B) The proportion of “A” and “B” ancestries in cases and controls across the chromosome with the spike of “B” ancestry at the risk locus in cases.
In the present study, we performed the largest AA admixture-mapping analysis to date with 9,322 total AA individuals. The sample includes the 5,644 participants previously analyzed by Hohman et al12 and increased the number of subjects by 65%. Taking advantage of the large sample size and potential power of admixture mapping, this study aims to identify ancestry-specific genetic loci associated with AD in AAs using twostep approach: (i) admixture mapping to prioritize the genetic regions associated with AD, and (ii) multipronged fine-mapping strategy to translate the findings from associated genetic regions to potential causal gene(s) (Figure 2).
Figure 2.
Schematic illustration of the admixture mapping and fine-mapping analysis.
2. METHODS
2.1. Study samples
This study included 3,166 AD cases and 6,156 controls of self-reported African Americans from 17 datasets: the Adult Changes in Thought (ACT) Study13, the National Institute on Aging (NIA) Alzheimer’s Disease Centers (ADCs)14, the University of Miami/Vanderbilt University (UM/VU)15,16, the Mount Sinai School of Medicine (MSSM) Brain Bank17, the Washington Heights Inwood Columbia Aging Project (WHICAP)18, The African American Alzheimer’s Disease Genetics (AAG) Study19, the MIRAGE Study20, NIA- LOAD/NCRAD21, the Mayo Clinic22, the Rush University Alzheimer’s disease Center23–26 (ROS/MAP, MARS/CORE), the Chicago Health and Aging Project (CHAP)27, the Indianapolis Ibadan Dementia Study (Indianapolis)28, the Genetic and Environmental Risk Factors for Alzheimer’s Disease Among African Americans (GenerAAtions) Study29, the University of Pittsburgh (UP)30, Washington University (WU)31–34 and Research in African American Alzheimer Disease Initiative (REAAADI)35. All participants were consented under IRB protocols approved by the appropriate Institutional Review Board. Participants were assessed by following NINCDS-ADRDA criteria for possible and probable AD36. The detailed description of the study samples, diagnosis of AD, and age of onset are described elsewhere7. Three datasets have family data. The details of the family numbers within these datasets are provided in Supplemental Table 1.
Genotype quality control:
Genome-wide genotyping was performed using different array chips, full details of array genotyping explained elsewhere7. Quality control analyses were performed individually for each dataset using the software PLINK v.237. Genotypes with a call rate less than 95% or deviating from Hardy-Weinberg equilibrium (HWE) (p<1.e-6) were eliminated. The concordance between reported sex and genotype-inferred sex was verified using X-chromosome data. The relationships among individuals within families were confirmed using “identical by descent” (IBD) allele sharing.
Genotype Imputation:
All genotype-array datasets were individually phased and imputed to the TOPMed R2 version panel (build of GRCh38) using the TOPMed Imputation server38. The TOPMed R2 reference panel provides information on ~308 million genetic variants and contains 97,256 samples. Most of samples are predominantly from non-European populations and ~25% of the samples are from African-descent populations.
2.2. Assessment of genetic ancestry
We assessed the global ancestry using the PC-AiR approach, which is robust to known and cryptic relatedness39. We calculated PCs without population reference datasets and performed the Tracy-Widom test to select PCs for covariates in the genome-wide admixture-mapping analysis40.
To assess local ancestry, first we combined our array genotype data with HGDP reference panel using PLINK v2 software37, 41 and phased the data using the SHAPEIT tool ver. 2 with default settings and 1000 Genomes Phase 3 reference panel42,43. Then, we performed RFMix, a discriminative modeling approach, to infer the local ancestry at each locus across the genome44.
2.3. Statistical analysis
Admixture mapping:
We performed admixture mapping separately within each of 17 AA datasets using the GENESIS software. First, we encoded copies of African local-ancestry calls as dosage values (0,1, or 2; number of African haplotypes at a locus). Then, to test for an association between AD and local ancestry at a genomic location, we used a logistic mixed model. The model includes local ancestry () as main and the genetic relationship matrix (GRM) as a random effect to adjust for the sample relatedness, and was adjusted further for age, sex, and genome-wide ancestry (PC1:3) (equation 1).
| (equation 1) |
Meta-analysis:
To meta-analyze the 17 AA datasets, which were separately analyzed using admixture mapping, we first aligned the genetic positions across the datasets, and then we performed meta-analysis using METASOFT software45. To replicate the findings, we conducted a meta-analysis with an independent study of imputed genotype data from 11,424 Caribbean Hispanic (CHI) individuals (full details of array genotyping, imputation, and QC are described elsewhere46). We used the random effects approach proposed by Han and Eskin (RE2) that is more powerful than traditional random-effect models for detecting small-effect heterogeneous genetic associations45. The RE2 model assumes no heterogeneity under the null hypothesis, the average genetic effect () and between-study variance () are zero: vs . (Ancestry and statistical analysis methods are described in detail in Supplemental methods).
2.4. Adjustment for multiple testing and power analysis
Because of the highly correlated tests in admixture mapping analysis, we implemented the STEAM software to estimate the admixture mapping significance threshold and control the family-wise error rate at level 0.0547. To calculate an appropriate significance threshold for admixture mapping STEAM accounts for the number of ancestral populations, generations since admixture, population structure, and density of markers tested47. We performed power analysis by using multi-ancestry admixture mapping (PAMAM) software48.
2.5. Fine Mapping
Admixture mapping identifies regions of interest that may span multiple megabases (mB). We used a multipronged fine-mapping strategy to translate the findings from associated genetic regions to potential causal gene(s). First, to identify risk loci that correlate with ancestral background within the relatively broad regions identified by admixture mapping (p-value < 0.001), we conducted what we refer to as an “ancestry-aware regression analysis”. In this model, affection status is modeled as a function of the risk genotype, LA and an interaction term (LA x genotype). Significance of the interaction term suggests that association between genotype and the risk of AD depends on the ancestral background of the genotype. Specifically, LA modifies the effect of the risk loci such that risk allele has a modified effect on the risk of AD with increasing or decreasing dosage of specific ancestry. This can be an effective fine-mapping tool because conditioning on LA removes signals at variants that are in LD with the causal variant due only to the admixture of the population49.
We conducted ancestry-aware analyses in the AA imputed datasets with the local ancestry and genotype (G) as main effects and the interaction term, also adjusted for age, sex, and genome-wide ancestry (PC1:3) and included the genetic relationship matrix (GRM) as a random effect to adjust for the sample relatedness (equation 2). We used a parametric bootstrap50 to get precise p-value for each test, using a large number of bootstrap permutations that substantially exceeds the number of independent tests in the prioritized genetic regions. An adjusted p-value was calculated by multiplying the p-value obtained through the parametric bootstrap by the number of independent tests. To identify the number of independent tests we implemented a linkage disequilibrium (LD)-based pruning step to remove SNPs in high LD (r2>0.1) with any nearby SNP (< 250 Kilobase), based LD estimates from publicly available 1000 Genomes phase3 ASW (African ancestry in Southwest USA) and YRI (Yoruba in Ibadan, Nigeria) populations data (The code used in this study to perform admixture mapping and ancestry-aware analysis is available here https://github.com/hihg-um/ad-admixture-mapping).
| (equation 2) |
For further prioritization of ancestry-aware analysis results, we used the web-based platform FUMA to prioritize a set of variants and genes using the functional annotation and the LD pattern with the lead variant (r2>0.6)51. All variants were annotated by ANNOVAR, combined annotation dependent depletion (CADD) scores, and RegulomeDB (RDB) scores52–54. We also examined genes for differential expression using results from RNA sequencing of peripheral blood collected from 234 AA (115 AD cases, 119 controls) and 240 NHW (121 AD cases, 119 controls). Analyses and datasets are described elsewhere55. Finally, we validated the prioritized genes using Accelerating Medicines Partnership in Alzheimer’s Disease (AMP-AD) AGORA knowledge portal (https://agora.ampadportal.org/genes), where the data on differential expression of genes across the brain regions from NHW individuals are available. These data are from Mount Sinai Brain Bank Study (MSBB) (number of individuals = 164), Religious Orders Study (ROS) and Memory and Aging Project (MAP) (number of individuals = 241), and Mayo RNASeq Study (number of individuals = 179).
3. RESULTS
3.1. Estimates of Genetic Ancestry
The average proportions of African and European local ancestry across the 17 cohorts, based on the autosomes only, were similar (Supplemental Figure 2, Supplemental Table 2). The average African ancestry proportion across all studies was 0.782 (SD=0.023). We did not observe any significant differences in global ancestry proportions (based on average LA) between cases and controls. The overall African-ancestry proportion in cases was 0.783 (SD = 0.032) and in controls was 0.782 (SD = 0.024).
3.2. Admixture Mapping
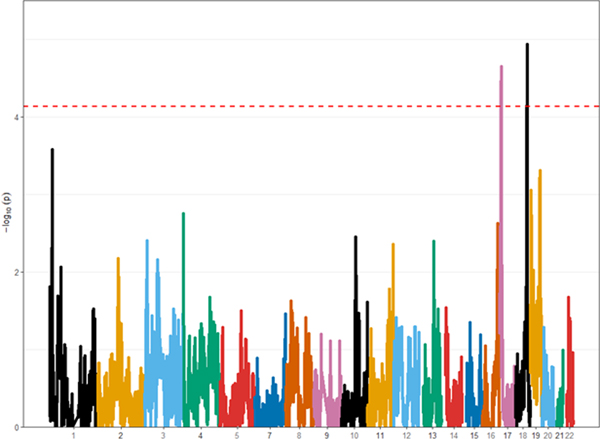
We estimated 7.3e-5 as a threshold for statistical genome-wide significance for this study using the analytic approximation approach in STEAM software. Using this threshold, we identified two loci genome-wide significantly associated with AD: one on chromosome 17p13.2 (p-value = 2.20 × 10−5) and one on 18q21.33 (p-value = 1.20 × 10−5) (Figure 3). Both regions were genome-wide significant in the meta-analysis of our AA results with results from the CHI dataset from Sariya et al.46 (Supplemental Figure 3) (17p13.2: p-value = 2.01 × 10−5; 18q21.33: p-value = 1.96 × 10−5).
Figure 3. Admixture mapping Manhattan plot of meta-analysis African American datasets.
Genomic coordinates are displayed along the X-axis, with each color representing a different chromosome. Negative logarithm of the association p-value for the ancestral allele is displayed on the Y-axis. The red dotted horizontal line represents the significance threshold after multiple testing correction, i.e. those dots falling above the line are considered study-wise significant
3.3. Fine Mapping
The admixture blocks span the large genomic region, so identifying the causal gene(s) from the admixture mapping is challenging. The mean of admixture block sizes in the largest dataset of AA individuals (~2,900) in our study was ~36Mb. Supplementary Figure 4 illustrates the distribution of the admixture block sizes in AAs. In this study, we leveraged several sources of information to prioritize genes and causal variants. We overlapped information from ancestry-aware regression analysis followed by comprehensive in silico fine-mapping, and differential gene expression data to prioritize AD related genes and variants at the loci identified through the admixture mapping.
3.3.1. Ancestry-aware Analysis
We defined two regions of small p-values (p-value < 0.001) around the genome-wide significant results at chromosome 17p13.2 (spanning from ~4.8 Mb to ~8.6 Mb) and at chromosome 18q21.33 (spanning from ~58.0 Mb to ~61.0 Mb) for fine-mapping (build of GRCh37). The ancestry-aware regression model identified a significant interaction term (G x LA) on chromosome 17p13.2 with the genotypes of the lead SNP rs72835045 ( = 0.97; SE = 0.28; p-value=7.93×10−5; parametric bootstrap FDR < 0.05). Figure 4 illustrates the ancestry-aware regression model results across the chromosome 17p13.2 locus for the G X LA interaction term. Significance of the interaction term at the chromosome 17p13.2 locus implies that association between rs72835045 genotype and the risk of AD depends on the LA background of the genotype. Specifically, LA modifies the effect of the rs72835045 genotype risk allele (A) such that allele “A” has a greater effect on the risk of AD with increasing dosage of European ancestry (or conversely decreasing dosage of African ancestry). We did not observe any significant G x LA interactions for the chromosome 18q21.33 locus (Supplement Figure 4).
Figure 4. Region plot of ancestry-aware regression analysis for the 17p13.2 region.
Genomic coordinates are displayed along the X-axis. Negative logarithm of the ancestry-aware regression approach p-value for the interaction term (G x LA) is displayed on the Y-axis.
3.3.2. Functional Mapping and Annotation
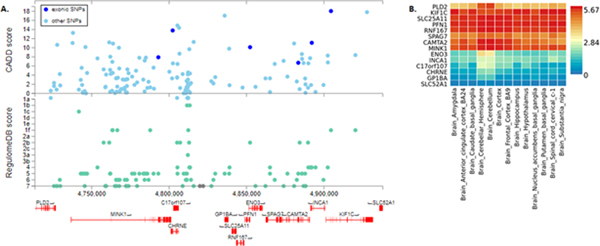
FUMA identified 157 variants, which are in LD (r2>0.6) with the lead variant rs72835045 on chromosome 17q. The resulting output of the interaction model for these variants is shown in Supplemental Table 3. We used pre-calculated LD structure based on 1000G European populations as the European local ancestry were increasing the AD risk in rs72835045_A allele carriers. Figure 5A illustrates the distribution of 157 variants across the rs72835045 genotype LD region. There are 6 exonic variants, two of which are missense variants (rs2283568 in the INCA1 gene, rs74744272 in the PFN1 gene). Both missense variants have CADD score > 10, and missense variant rs2283568 has a regulatory potential with RDB score of 2b. The synonymous exonic variant rs17707385 in KIF1C gene has the highest CADD score of 18.04 and the RDB score 1f. Functional annotation shows that in total 23 variants have a strong regulatory potential with the RDB score , and 18 variants have possible deleterious effect with CADD score . The whole list of annotated variants by ANNOVAR, CADD scores, and RDB scores are presented in the Supplemental Table 4.
Figure 5. Follow up 17p13.2 locus using functional mapping and annotation.
A-Regional plot of the 17p13.2 locus, which includes prioritized genes and variants with CADD and RequlomeDB scores. B-Gene expression heatmaps constructed with GTEX v6 13 brain tissues types. Genes and tissues are ordered by clusters for the GTEX heatmap.
FUMA mapped the variants to 14 genes based on ANNOVAR annotation. The FUMA GENE2FUNC process provided tissue specific expression patterns based on GTEx v6 RNA-seq data. Figure 5B illustrates the expression levels for 14 genes in the heatmap for 13 brain tissue types. Eight genes (PLD2, KIFC1, SLC25A11, PFN1, RNF167, SPAG7, CAMTA2, and MINK1) show consistent high expression pattern among all brain tissue types.
3.3.3. Differential Gene-expression Analysis
Peripheral blood from AA and NHW individuals:
We examined the genes on the chromosome 17p13.2 that were in LD with the rs72835045 genotype and all the genes located on the chromosome 18q21.33 that spans the admixture mapping signal. We identified differential expression (DE) for three genes in the chromosome 17p13.2 locus in the joint AA and NHW gene-expression analyses and DE in one gene in the chromosome 18q21.33 locus in the NHW gene-expression analysis (Table 1). All genes in the chromosome 17p13.2 locus were upregulated in AD and the gene on the chromosome 18q21.33 locus was downregulated in AD. A complete list of expression results for the genes on the chromosome 17p13.2 and 18q21.33 is provided in Supplemental Table 4.
Table 1.
Significantly differentially expressed genes between Alzheimer disease and controls in non-Hispanic Whites (NHW), non-Hispanic Whites and African Americans joint analysis (NHW+AA).
| Gene | Locus | Fold Change | P-Value | FDR | Study |
|---|---|---|---|---|---|
| GP1BA | 17p13.2 | 1.21 | 0.0036 | 0.03 | NHW+AA |
| SLC25A11 | 17p13.2 | 1.08 | 0.0063 | 0.05 | NHW+AA |
| KIF1C | 17p13.2 | 1.07 | 0.0068 | 0.05 | NHW+AA |
| BCL2 | 18q21.33 | −1.17 | 0.0006 | 0.04 | NHW |
AMP-AD data:
We queried the 14 genes from chromosome 17p13.2 locus and 11 genes from chromosome 18q21.33 using web-based database Agora to further investigate the differential gene expression in 7 types of post-mortem brain tissues, which were dorsolateral prefrontal cortex (DLPFC) from ROSMAP, temporal cortex (TCX) and cerebellum (CBE) from Mayo, and inferior frontal gyrus (IFG), superior temporal gyrus (STG), frontal pole (FP), and parahippocampal gyrus (PHG) from MSBB. Out of 14 genes from the chromosome 17p13.2 locus, eight genes showed significant DE in various brain tissues, where four of them (SLC25A11, MINK1, KIF1C, and CAMTA2) showed significant DE in more than one brain tissue type. Seven genes from the chromosome 18q21.33 locus showed significant DE, where five genes showed DE in two type of tissues. Supplemental Figures 6A and 6B shows the forest plots of eight genes from the 17p13.2 locus and five genes from the 18q21.33 locus, respectively.
4. DISCUSSION
We identified two genome-wide significant admixture mapping signals: on chromosomes 17p13.2 and 18q21.33. Our results showed that 18q21.33 locus contributes to increased risk for AD in AAs with the increased frequency of African local ancestry, whereas European local ancestry increases the risk effect of the 17p13.2 locus. These genetic regions are novel in AA populations. Our study with the increased sample size and new ancestry-aware statistical approaches was able to identify the regions that were not previously found in AA genome-wide association studies or the previous AA admixture mapping study7,12.
The finding at the 17p13.2 locus is intriguing because this genetic region was identified as a genome-wide significant AD risk locus in three NHW AD meta-analysis of GWAS’s with sample sizes over 100,0004,6,56. Our follow-up ancestry-aware analysis narrowed the 17p13.2 locus to an interval of 200 kilobase, that is likely to harbor risk variants explaining the admixture peak at the 17p13.2 locus. We compared the variants prioritized through the ancestry-aware fine mapping with previous studies in NHW and AA AD GWAS studies3,5. The lead marker showing significant interaction with European local ancestry (rs72835045; p-value=7.93×10−5) was significantly associated with AD risk in NHW GWAS study3 (p-value = 3.28×10−4), whereas it was not found as a risk marker in AA GWAS study5 (p-value=0.61). Supplemental Table 6 summarize and compare the prioritized markers through this study with NHW and AA GWAS studies3,5. Our findings at the 17p13.2 locus suggest that AA samples inherit their risk allele from European ancestors and have a lower risk of AD if they inherited a segment of African ancestry at the 17p13.2 locus. This finding supports the hypothesis that the variation of both genetic risk loci and their effect across the populations is most likely related to genetic ancestry that is surrounding the risk locus (local ancestry) and not related the overall ancestry (global ancestry)9,10.
Our fine-mapping and multi-omics approach homed in on three genes at the 17p13.2 locus: the Kinesin Family Member 1C (KIF1C), Profilin 1 (PFN1) and Misshapen Like Kinase 1 (MINK1). These genes show strong evidence of regulatory potential and deleterious effect on protein function. All genes are highly expressed in several GTEx brain tissue types. KIF1C and MINK1 genes were differentially expressed between AD and non-AD brain tissues, and additionally KIF1C showed significant differential expression in blood. The KIF1C variant rs17707385 has strong evidence of regulatory potential and deleterious effect on protein function and was in strong LD (r2 = 0.62) with the lead variant from the ancestry-aware regression analysis. KIF1C encodes a motor protein involved in retrograde transport from the Golgi to the endoplasmic reticulum, suggesting a role of intracellular trafficking processes57. Studies have also shown a link between the deficits in the active transport of gene products and neurodegenerative disorders58. The PFN1 gene encodes a member of the profilin family of small actin-binding proteins involved in actin polymerization, and its dynamics are essential for the basic cellular activities59. Rare, missense mutations within the PFN1 gene were associated with the neurodegenerative disorder Amyotrophic lateral sclerosis60. A recent AD study found a variant within the MINK1 gene with significant pleiotropic genetic effect in AD and cardiovascular disease61. Intriguingly, the pleiotropic MINK1 gene variant identified by Broce et al and colleagues is in strong LD (r2=0.97) with the lead variant identified in ancestry-aware regression analysis.
Our follow up using available AD gene expression data revealed an interesting candidate gene at the 18q21.33 locus: BCL2. This gene is highly expressed in several brain tissue types and was differentially expressed between AD and non-AD brain tissues and blood. BCL2 gene play an important role in regulating the autophagy62. Studies have shown that overexpression of Bcl2 gene improves the place recognition memory in mice, protects neuronal cells in vitro from the Aβ-related death, influences the tau processing, and reducing the number of NFTs63,64.
Our study, with the substantially increased sample size, found nominal significance for four regions out of seven regions that were identified in the previous AA admixture mapping study12. All nominally replicated regions showed a similar direction of effect. Notably, the ABCA7 region showed the strongest association compared to other regions (). The sample size of this study was much larger (~65% more) than the previous AA AD admixture mapping study. Additionally, the approaches were different. In our study, the association between AD and local ancestry was assessed within each dataset separately by conducting logistic regression controlled for age, sex, global ancestry (PC1:3), and known and cryptic relatedness, followed by meta-analysis. The study by Hohman et al. did not account for global ancestry in their models, and they performed joint analysis of a combined dataset using Wilcoxon rank-sum test.This study has several limitations. First, our dataset is underpowered to identify association between the local ancestry and AD when the risk-allele frequencies and their effect sizes are similar across the ancestral groups (e.g., African and European). Larger samples will be required to detect these more subtle AD-associated admixture blocks. Second, we combined the AA samples in one discovery dataset to maximize the power of the study and relied on NHW and AA GWAS and gene expression data from peripheral blood and AMP-AD data for validation3,5,55,. An additional independent dataset is needed for replication analysis. Third, ancestry-aware fine-mapping analysis is not well powered in our study for the large genomic region on chromosome 18q21.33 prioritized through admixture mapping. Increasing the sample size in the AA dataset can help to translate the findings from associated genetic regions to potential causal gene(s).
We provided robust evidence for two genomic regions to be associated with the AD risk and prioritized several candidate genes and markers through the fine-mapping approaches. Further investigation of the prioritized loci through long-read sequencing may allow for more comprehensive and accurate detection of variation, especially structural variation. As a next step, integrating the prioritized variants with the cell-type specific data can help to translate the findings into clinically actionable gene sets.
In summary, we identified two novel genome-wide significant loci (17p13.2 and 18q21.33) associated with the increased risk of AD in AAs. Our study generalized known European genetic AD risk locus (17p13.2) to the AA population. The lack of identification of the 18q21.33 region previously in large NHW and incomparably smaller AA GWAS studies, and excess of African ancestry in AD AA individuals at this locus suggest that genetic risk factor(s) is(are) likely to be African-ancestry specific. We found that the overall proportion of African ancestry does not differ between the cases and controls. The similar proportions of parental ancestries in AA cases and controls suggest that African genetic ancestry alone is not likely to explain the AD prevalence difference between AA and NHW populations. Thus, it is likely that social and environmental determinants in concert with the ancestral and genetic factors influence the susceptibility to AD65.
Supplementary Material
Systematic review: The authors reviewed the literature using PubMed. The African American (AA) population is substantially underrepresented in Alzheimer disease (AD) genetic studies, yet its admixed genetic ancestry provides a unique opportunity to identify novel genetic factors associated with AD and related to genetic ancestry.
Interpretation: This study identified two genome-wide significant loci on chromosomes 17p13.2 and 18q21.33. These genetic regions are novel in AAs. Ancestry-aware approach showed that AA individuals have a lower risk of AD if they inherited African ancestry admixture block at the 17p13.2 locus.
Future directions: Further investigation of the prioritized loci through long-read sequencing may allow for more comprehensive and accurate detection of variation, especially structural variation. As a next step, integrating the prioritized variants with the cell-type specific data can help to translate the findings into clinically actionable gene sets.
Highlights.
We identified two genome-wide significant admixture mapping signals: on chromosomes 17p13.2 and 18q21.33, which are novel in African American populations.
Our ancestry-aware regression approach showed that AA individuals have a lower risk of AD if they inherited African ancestry admixture block at the 17p13.2 locus.
We found that the overall proportion of African ancestry does not differ between the cases and controls that suggest African genetic ancestry alone is not likely to explain the Alzheimer disease prevalence difference between African American and non-Hispanic White populations.
5. AKNOWLEDGEMENT
This investigation was supported by grant AG16002, AG032984, AG052410, AG072547, AG057659, AG058654 and AG070864 from the National Institutes on Aging of NIH and the A2018556F grant from the BrightFocus Foundation.
Study specific funding:
This research was supported by the National Institute on Aging (AG16002, AG032984, AG052410, AG072547, AG057659, AG058654 and AG070864) as well as BrightFocus (A2018556F).
Footnotes
CONFLICTS OF INTEREST
Drs. E.R. Martin, W.S. Bush, L.A. Farrer, A. Naj and J.L. Haines received support from NIH/NIA grant AG058654, which supported the work of this manuscript. Dr. F. Rajabli, received support from NIH/NIA grant AG070864, as well as BrightFocus Foundation A2018556F, which supported his effort on this manuscript. Schellenberg has received support from NIH/NIA grants AG16002 and AG032984 that supported the work of this manuscript. Dr. G.W. Beecham has received support from NIH/NIA grants AG052410, AG070935, AG058654, and AG070864, which supported his effort on this manuscript. Dr. G.S Byrd received support from NIA/NIH grant AG052410. Dr C. Reitz received support from NIH/NIA grants AG052410, AG072547, AG056270 and AG064614, that supported her work on this manuscript. Dr. R.P. Mayeux also received support from NIH/NIA grants AG052410 and AG057659. Dr. B.N. Vardarajan received support from NIH/NIA grants AG057659. Dr. B.W. Kunkle received support from grant AG057659. Dr. M.A. Pericak-Vance has received support from NIH/NIA grants AG052410, AG072547, AG057659, AG058654 and AG070864. No other authors have any interests to disclose.
7. REFERENCES
- 1.Folstein MF, Bassett SS, Anthony JC, Romanoski AJ, Nestadt GR. Dementia: case ascertainment in a community survey. J Gerontol. 1991. Jul;46(4):M132–8. doi: 10.1093/geronj/46.4.m132. [DOI] [PubMed] [Google Scholar]
- 2.Gurland BJ, Wilder DE, Lantigua R, Stern Y, Chen J, Killeffer EH, Mayeux R. Rates of dementia in three ethnoracial groups. Int J Geriatr Psychiatry. 1999. Jun;14(6):481–93. [PubMed] [Google Scholar]
- 3.Kunkle BW, Grenier-Boley B, Sims R, Bis JC, Damotte V, Naj AC, Boland A, Vronskaya M, van der Lee SJ, Amlie-Wolf A, Bellenguez C, Frizatti A, Chouraki V, Martin ER, Sleegers K, Badarinarayan N, Jakobsdottir J, Hamilton-Nelson KL, Moreno-Grau S, Olaso R, Raybould R, Chen Y, Kuzma AB, Hiltunen M, Morgan T, Ahmad S, Vardarajan BN, Epelbaum J, Hoffmann P, Boada M, Beecham GW, Garnier JG, Harold D, Fitzpatrick AL, Valladares O, Moutet ML, Gerrish A, Smith AV, Qu L, Bacq D, Denning N, Jian X, Zhao Y, Del Zompo M, Fox NC, Choi SH, Mateo I, Hughes JT, Adams HH, Malamon J, Sanchez-Garcia F, Patel Y, Brody JA, Dombroski BA, Naranjo MCD, Daniilidou M, Eiriksdottir G, Mukherjee S, Wallon D, Uphill J, Aspelund T, Cantwell LB, Garzia F, Galimberti D, Hofer E, Butkiewicz M, Fin B, Scarpini E, Sarnowski C, Bush WS, Meslage S, Kornhuber J, White CC, Song Y, Barber RC, Engelborghs S, Sordon S, Voijnovic D, Adams PM, Vandenberghe R, Mayhaus M, Cupples LA, Albert MS, De Deyn PP, Gu W, Himali JJ, Beekly D, Squassina A, Hartmann AM, Orellana A, Blacker D, Rodriguez-Rodriguez E, Lovestone S, Garcia ME, Doody RS, Munoz-Fernadez C, Sussams R, Lin H, Fairchild TJ, Benito YA, Holmes C, Karamujić-Čomić H, Frosch MP, Thonberg H, Maier W, Roshchupkin G, Ghetti B, Giedraitis V, Kawalia A, Li S, Huebinger RM, Kilander L, Moebus S, Hernández I, Kamboh MI, Brundin R, Turton J, Yang Q, Katz MJ, Concari L, Lord J, Beiser AS, Keene CD, Helisalmi S, Kloszewska I, Kukull WA, Koivisto AM, Lynch A, Tarraga L, Larson EB, Haapasalo A, Lawlor B, Mosley TH, Lipton RB, Solfrizzi V, Gill M, Longstreth WT Jr, Montine TJ, Frisardi V, Diez-Fairen M, Rivadeneira F, Petersen RC, Deramecourt V, Alvarez I, Salani F, Ciaramella A, Boerwinkle E, Reiman EM, Fievet N, Rotter JI, Reisch JS, Hanon O, Cupidi C, Uitterlinden AGA, Royall DR, Dufouil C, Maletta RG, de Rojas I, Sano M, Brice A, Cecchetti R, George-Hyslop PS, Ritchie K, Tsolaki M, Tsuang DW, Dubois B, Craig D, Wu CK, Soininen H, Avramidou D, Albin RL, Fratiglioni L, Germanou A, Apostolova LG, Keller L, Koutroumani M, Arnold SE, Panza F, Gkatzima O, Asthana S, Hannequin D, Whitehead P, Atwood CS, Caffarra P, Hampel H, Quintela I, Carracedo Á, Lannfelt L, Rubinsztein DC, Barnes LL, Pasquier F, Frölich L, Barral S, McGuinness B, Beach TG, Johnston JA, Becker JT, Passmore P, Bigio EH, Schott JM, Bird TD, Warren JD, Boeve BF, Lupton MK, Bowen JD, Proitsi P, Boxer A, Powell JF, Burke JR, Kauwe JSK, Burns JM, Mancuso M, Buxbaum JD, Bonuccelli U, Cairns NJ, McQuillin A, Cao C, Livingston G, Carlson CS, Bass NJ, Carlsson CM, Hardy J, Carney RM, Bras J, Carrasquillo MM, Guerreiro R, Allen M, Chui HC, Fisher E, Masullo C, Crocco EA, DeCarli C, Bisceglio G, Dick M, Ma L, Duara R, Graff-Radford NR, Evans DA, Hodges A, Faber KM, Scherer M, Fallon KB, Riemenschneider M, Fardo DW, Heun R, Farlow MR, Kölsch H, Ferris S, Leber M, Foroud TM, Heuser I, Galasko DR, Giegling I, Gearing M, Hüll M, Geschwind DH, Gilbert JR, Morris J, Green RC, Mayo K, Growdon JH, Feulner T, Hamilton RL, Harrell LE, Drichel D, Honig LS, Cushion TD, Huentelman MJ, Hollingworth P, Hulette CM, Hyman BT, Marshall R, Jarvik GP, Meggy A, Abner E, Menzies GE, Jin LW, Leonenko G, Real LM, Jun GR, Baldwin CT, Grozeva D, Karydas A, Russo G, Kaye JA, Kim R, Jessen F, Kowall NW, Vellas B, Kramer JH, Vardy E, LaFerla FM, Jöckel KH, Lah JJ, Dichgans M, Leverenz JB, Mann D, Levey AI, Pickering-Brown S, Lieberman AP, Klopp N, Lunetta KL, Wichmann HE, Lyketsos CG, Morgan K, Marson DC, Brown K, Martiniuk F, Medway C, Mash DC, Nöthen MM, Masliah E, Hooper NM, McCormick WC, Daniele A, McCurry SM, Bayer A, McDavid AN, Gallacher J, McKee AC, van den Bussche H, Mesulam M, Brayne C, Miller BL, Riedel-Heller S, Miller CA, Miller JW, Al-Chalabi A, Morris JC, Shaw CE, Myers AJ, Wiltfang J, O’Bryant S, Olichney JM, Alvarez V, Parisi JE, Singleton AB, Paulson HL, Collinge J, Perry WR, Mead S, Peskind E, Cribbs DH, Rossor M, Pierce A, Ryan NS, Poon WW, Nacmias B, Potter H, Sorbi S, Quinn JF, Sacchinelli E, Raj A, Spalletta G, Raskind M, Caltagirone C, Bossù P, Orfei MD, Reisberg B, Clarke R, Reitz C, Smith AD, Ringman JM, Warden D, Roberson ED, Wilcock G, Rogaeva E, Bruni AC, Rosen HJ, Gallo M, Rosenberg RN, Ben-Shlomo Y, Sager MA, Mecocci P, Saykin AJ, Pastor P, Cuccaro ML, Vance JM, Schneider JA, Schneider LS, Slifer S, Seeley WW, Smith AG, Sonnen JA, Spina S, Stern RA, Swerdlow RH, Tang M, Tanzi RE, Trojanowski JQ, Troncoso JC, Van Deerlin VM, Van Eldik LJ, Vinters HV, Vonsattel JP, Weintraub S, Welsh-Bohmer KA, Wilhelmsen KC, Williamson J, Wingo TS, Woltjer RL, Wright CB, Yu CE, Yu L, Saba Y; Alzheimer Disease Genetics Consortium (ADGC); European Alzheimer’s Disease Initiative (EADI); Cohorts for Heart and Aging Research in Genomic Epidemiology Consortium (CHARGE); Genetic and Environmental Risk in AD/Defining Genetic, Polygenic and Environmental Risk for Alzheimer’s Disease Consortium (GERAD/PERADES), Pilotto A, Bullido MJ, Peters O, Crane PK, Bennett D, Bosco P, Coto E, Boccardi V, De Jager PL, Lleo A, Warner N, Lopez OL, Ingelsson M, Deloukas P, Cruchaga C, Graff C, Gwilliam R, Fornage M, Goate AM, Sanchez-Juan P, Kehoe PG, Amin N, Ertekin-Taner N, Berr C, Debette S, Love S, Launer LJ, Younkin SG, Dartigues JF, Corcoran C, Ikram MA, Dickson DW, Nicolas G, Campion D, Tschanz J, Schmidt H, Hakonarson H, Clarimon J, Munger R, Schmidt R, Farrer LA, Van Broeckhoven C, O’Donovan MC, DeStefano AL, Jones L, Haines JL, Deleuze JF, Owen MJ, Gudnason V, Mayeux R, Escott-Price V, Psaty BM, Ramirez A, Wang LS, Ruiz A, van Duijn CM, Holmans PA, Seshadri S, Williams J, Amouyel P, Schellenberg GD, Lambert JC, Pericak-Vance MA. Author Correction: Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat Genet. 2019. Sep;51(9):1423–1424. doi: 10.1038/s41588-019-0495-7. Erratum for: Nat Genet. 2019 Mar;51(3):414–430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Jansen IE, Savage JE, Watanabe K, Bryois J, Williams DM, Steinberg S, Sealock J, Karlsson IK, Hägg S, Athanasiu L, Voyle N, Proitsi P, Witoelar A, Stringer S, Aarsland D, Almdahl IS, Andersen F, Bergh S, Bettella F, Bjornsson S, Brækhus A, Bråthen G, de Leeuw C, Desikan RS, Djurovic S, Dumitrescu L, Fladby T, Hohman TJ, Jonsson PV, Kiddle SJ, Rongve A, Saltvedt I, Sando SB, Selbæk G, Shoai M, Skene NG, Snaedal J, Stordal E, Ulstein ID, Wang Y, White LR, Hardy J, Hjerling-Leffler J, Sullivan PF, van der Flier WM, Dobson R, Davis LK, Stefansson H, Stefansson K, Pedersen NL, Ripke S, Andreassen OA, Posthuma D. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat Genet. 2019. Mar;51(3):404–413. doi: 10.1038/s41588-018-0311-9. Epub 2019 Jan 7. Erratum in: Nat Genet. 2020 Mar;52(3):354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kunkle BW, Schmidt M, Klein HU, Naj AC, Hamilton-Nelson KL, Larson EB, Evans DA, De Jager PL, Crane PK, Buxbaum JD, Ertekin-Taner N, Barnes LL, Fallin MD, Manly JJ, Go RCP, Obisesan TO, Kamboh MI, Bennett DA, Hall KS, Goate AM, Foroud TM, Martin ER, Wang LS, Byrd GS, Farrer LA, Haines JL, Schellenberg GD, Mayeux R, Pericak-Vance MA, Reitz C; Writing Group for the Alzheimer’s Disease Genetics Consortium (ADGC), Graff-Radford NR, Martinez I, Ayodele T, Logue MW, Cantwell LB, Jean-Francois M, Kuzma AB, Adams LD, Vance JM, Cuccaro ML, Chung J, Mez J, Lunetta KL, Jun GR, Lopez OL, Hendrie HC, Reiman EM, Kowall NW, Leverenz JB, Small SA, Levey AI, Golde TE, Saykin AJ, Starks TD, Albert MS, Hyman BT, Petersen RC, Sano M, Wisniewski T, Vassar R, Kaye JA, Henderson VW, DeCarli C, LaFerla FM, Brewer JB, Miller BL, Swerdlow RH, Van Eldik LJ, Paulson HL, Trojanowski JQ, Chui HC, Rosenberg RN, Craft S, Grabowski TJ, Asthana S, Morris JC, Strittmatter SM, Kukull WA. Novel Alzheimer Disease Risk Loci and Pathways in African American Individuals Using the African Genome Resources Panel: A Meta-analysis. JAMA Neurol. 2021. Jan 1;78(1):102–113. doi: 10.1001/jamaneurol.2020.3536. Erratum in: JAMA Neurol. 2021 May 1;78(5):620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schwartzentruber J, Cooper S, Liu JZ, Barrio-Hernandez I, Bello E, Kumasaka N, Young AMH, Franklin RJM, Johnson T, Estrada K, Gaffney DJ, Beltrao P, Bassett A. Genome-wide meta-analysis, fine-mapping and integrative prioritization implicate new Alzheimer’s disease risk genes. Nat Genet. 2021. Mar;53(3):392–402. doi: 10.1038/s41588-020-00776-w. Epub 2021 Feb 15. Erratum in: Nat Genet. 2021 Feb 26;: [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Reitz C, Jun G, Naj A, Rajbhandary R, Vardarajan BN, Wang LS, Valladares O, Lin CF, Larson EB, Graff-Radford NR, Evans D, De Jager PL, Crane PK, Buxbaum JD, Murrell JR, Raj T, Ertekin-Taner N, Logue M, Baldwin CT, Green RC, Barnes LL, Cantwell LB, Fallin MD, Go RC, Griffith P, Obisesan TO, Manly JJ, Lunetta KL, Kamboh MI, Lopez OL, Bennett DA, Hendrie H, Hall KS, Goate AM, Byrd GS, Kukull WA, Foroud TM, Haines JL, Farrer LA, Pericak-Vance MA, Schellenberg GD, Mayeux R; Alzheimer Disease Genetics Consortium. Variants in the ATP-binding cassette transporter (ABCA7), apolipoprotein E ϵ4,and the risk of late-onset Alzheimer disease in African Americans. JAMA. 2013. Apr 10;309(14):1483–92. doi: 10.1001/jama.2013.2973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Farrer LA, Cupples LA, Haines JL, et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. APOE and Alzheimer Disease Meta Analysis Consortium. JAMA. 1997. Oct 22–29;278(16):1349–56. [PubMed] [Google Scholar]
- 9.Rajabli F, Feliciano BE, Celis K, Hamilton-Nelson KL, Whitehead PL, Adams LD, Bussies PL, Manrique CP, Rodriguez A, Rodriguez V, Starks T, Byfield GE, Sierra Lopez CB, McCauley JL, Acosta H, Chinea A, Kunkle BW, Reitz C, Farrer LA, Schellenberg GD, Vardarajan BN, Vance JM, Cuccaro ML, Martin ER, Haines JL, Byrd GS, Beecham GW, Pericak-Vance MA. Ancestral origin of ApoE ε4 Alzheimer disease risk in Puerto Rican and African American populations. PLoS Genet. 2018. Dec 5;14(12):e1007791. doi: 10.1371/journal.pgen.1007791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Blue EE, Horimoto ARVR, Mukherjee S, et al. Local ancestry at APOE modifies Alzheimer’s disease risk in Caribbean Hispanics. Alzheimers Dement. 2019. Dec;15(12):1524–1532. doi: 10.1016/j.jalz.2019.07.016. Epub 2019 Oct 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cukier HN, Kunkle BW, Vardarajan BN, Rolati S, Hamilton-Nelson KL, Kohli MA, Whitehead PL, Dombroski BA, Van Booven D, Lang R, Dykxhoorn DM, Farrer LA, Cuccaro ML, Vance JM, Gilbert JR, Beecham GW, Martin ER, Carney RM, Mayeux R, Schellenberg GD, Byrd GS, Haines JL, Pericak-Vance MA; Alzheimer’s Disease Genetics Consortium. ABCA7 frameshift deletion associated with Alzheimer disease in African Americans. Neurol Genet. 2016. May 17;2(3):e79. doi: 10.1212/NXG.0000000000000079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hohman TJ, Cooke-Bailey JN, Reitz C, Jun G, Naj A, Beecham GW, Liu Z, Carney RM, Vance JM, Cuccaro ML, Rajbhandary R, Vardarajan BN, Wang LS, Valladares O, Lin CF, Larson EB, Graff-Radford NR, Evans D, De Jager PL, Crane PK, Buxbaum JD, Murrell JR, Raj T, Ertekin-Taner N, Logue MW, Baldwin CT, Green RC, Barnes LL, Cantwell LB, Fallin MD, Go RC, Griffith P, Obisesan TO, Manly JJ, Lunetta KL, Kamboh MI, Lopez OL, Bennett DA, Hardy J, Hendrie HC, Hall KS, Goate AM, Lang R, Byrd GS, Kukull WA, Foroud TM, Farrer LA, Martin ER, Pericak-Vance MA, Schellenberg GD, Mayeux R, Haines JL, Thornton-Wells TA; Alzheimer Disease Genetics Consortium. Global and local ancestry in African-Americans: Implications for Alzheimer’s disease risk. Alzheimers Dement. 2016. Mar;12(3):233–43. doi: 10.1016/j.jalz.2015.02.012. Epub 2015 Jun 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kukull WA, Higdon R, Bowen JD, McCormick WC, Teri L, Schellenberg GD, van Belle G, Jolley L, Larson EB. Dementia and Alzheimer disease incidence: a prospective cohort study. Arch Neurol. 2002. Nov;59(11):1737–46. doi: 10.1001/archneur.59.11.1737. [DOI] [PubMed] [Google Scholar]
- 14.Beekly DL, Ramos EM, Lee WW, Deitrich WD, Jacka ME, Wu J, Hubbard JL, Koepsell TD, Morris JC, Kukull WA; NIA Alzheimer’s Disease Centers. The National Alzheimer’s Coordinating Center (NACC) database: the Uniform Data Set. Alzheimer Dis Assoc Disord. 2007. Jul-Sep;21(3):249–58. doi: 10.1097/WAD.0b013e318142774e. [DOI] [PubMed] [Google Scholar]
- 15.Beecham GW, Martin ER, Li YJ, Slifer MA, Gilbert JR, Haines JL, Pericak-Vance MA. Genome-wide association study implicates a chromosome 12 risk locus for late-onset Alzheimer disease. Am J Hum Genet. 2009. Jan;84(1):35–43. doi: 10.1016/j.ajhg.2008.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Edwards TL, Scott WK, Almonte C, Burt A, Powell EH, Beecham GW, Wang L, Züchner S, Konidari I, Wang G, Singer C, Nahab F, Scott B, Stajich JM, Pericak-Vance M, Haines J, Vance JM, Martin ER. Genome-wide association study confirms SNPs in SNCA and the MAPT region as common risk factors for Parkinson disease. Ann Hum Genet. 2010. Mar;74(2):97–109. doi: 10.1111/j.1469-1809.2009.00560.x. Epub 2010 Jan 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Haroutunian V, Perl DP, Purohit DP, Marin D, Khan K, Lantz M, Davis KL, Mohs RC. Regional distribution of neuritic plaques in the nondemented elderly and subjects with very mild Alzheimer disease. Arch Neurol. 1998. Sep;55(9):1185–91. doi: 10.1001/archneur.55.9.1185. [DOI] [PubMed] [Google Scholar]
- 18.Tang MX, Stern Y, Marder K, Bell K, Gurland B, Lantigua R, Andrews H, Feng L, Tycko B, Mayeux R. The APOE-epsilon4 allele and the risk of Alzheimer disease among African Americans, whites, and Hispanics. JAMA. 1998. Mar 11;279(10):751–5. doi: 10.1001/jama.279.10.751. [DOI] [PubMed] [Google Scholar]
- 19.Meier IB, Manly JJ, Provenzano FA, Louie KS, Wasserman BT, Griffith EY, Hector JT, Allocco E, Brickman AM. White matter predictors of cognitive functioning in older adults. J Int Neuropsychol Soc. 2012. May;18(3):414–27. doi: 10.1017/S1355617712000227. Epub 2012 Mar 6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Green RC, Cupples LA, Go R, Benke KS, Edeki T, Griffith PA, Williams M, Hipps Y, Graff-Radford N, Bachman D, Farrer LA; MIRAGE Study Group. Risk of dementia among white and African American relatives of patients with Alzheimer disease. JAMA. 2002. Jan 16;287(3):329–36. doi: 10.1001/jama.287.3.329. [DOI] [PubMed] [Google Scholar]
- 21.Lee JH, Cheng R, Graff-Radford N, Foroud T, Mayeux R; National Institute on Aging Late-Onset Alzheimer’s Disease Family Study Group. Analyses of the National Institute on Aging Late-Onset Alzheimer’s Disease Family Study: implication of additional loci. Arch Neurol. 2008. Nov;65(11):1518–26. doi: 10.1001/archneur.65.11.1518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Carrasquillo MM, Zou F, Pankratz VS, Wilcox SL, Ma L, Walker LP, Younkin SG, Younkin CS, Younkin LH, Bisceglio GD, Ertekin-Taner N, Crook JE, Dickson DW, Petersen RC, Graff-Radford NR, Younkin SG. Genetic variation in PCDH11X is associated with susceptibility to late-onset Alzheimer’s disease. Nat Genet. 2009. Feb;41(2):192–8. doi: 10.1038/ng.305. Epub 2009 Jan 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Barnes LL, Shah RC, Aggarwal NT, Bennett DA, Schneider JA. The Minority Aging Research Study: ongoing efforts to obtain brain donation in African Americans without dementia. Curr Alzheimer Res. 2012. Jul;9(6):734–45. doi: 10.2174/156720512801322627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bennett DA, Schneider JA, Bienias JL, Evans DA, Wilson RS. Mild cognitive impairment is related to Alzheimer disease pathology and cerebral infarctions. Neurology. 2005. Mar 8;64(5):834–41. doi: 10.1212/01.WNL.0000152982.47274.9E. [DOI] [PubMed] [Google Scholar]
- 25.Bennett DA, Schneider JA, Buchman AS, Mendes de Leon C, Bienias JL, Wilson RS. The Rush Memory and Aging Project: study design and baseline characteristics of the study cohort. Neuroepidemiology. 2005;25(4):163–75. doi: 10.1159/000087446. Epub 2005 Aug 15. [DOI] [PubMed] [Google Scholar]
- 26.Bennett DA, Wilson RS, Schneider JA, Evans DA, Beckett LA, Aggarwal NT, Barnes LL, Fox JH, Bach J. Natural history of mild cognitive impairment in older persons. Neurology. 2002. Jul 23;59(2):198–205. doi: 10.1212/wnl.59.2.198. [DOI] [PubMed] [Google Scholar]
- 27.Bienias JL, Beckett LA, Bennett DA, Wilson RS, Evans DA. Design of the Chicago Health and Aging Project (CHAP). J Alzheimers Dis. 2003. Oct;5(5):349–55. doi: 10.3233/jad-2003-5501. [DOI] [PubMed] [Google Scholar]
- 28.Murrell JR, Price B, Lane KA, Baiyewu O, Gureje O, Ogunniyi A, Unverzagt FW, Smith-Gamble V, Gao S, Hendrie HC, Hall KS. Association of apolipoprotein E genotype and Alzheimer disease in African Americans. Arch Neurol. 2006. Mar;63(3):431–4. doi: 10.1001/archneur.63.3.431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Logue MW, Schu M, Vardarajan BN, Buros J, Green RC, Go RC, Griffith P, Obisesan TO, Shatz R, Borenstein A, Cupples LA, Lunetta KL, Fallin MD, Baldwin CT, Farrer LA; Multi-Institutional Research on Alzheimer Genetic Epidemiology (MIRAGE) Study Group. A comprehensive genetic association study of Alzheimer disease in African Americans. Arch Neurol. 2011. Dec;68(12):1569–79. doi: 10.1001/archneurol.2011.646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kamboh MI, Minster RL, Demirci FY, Ganguli M, Dekosky ST, Lopez OL, Barmada MM. Association of CLU and PICALM variants with Alzheimer’s disease. Neurobiol Aging. 2012. Mar;33(3):518–21. doi: 10.1016/j.neurobiolaging.2010.04.015. Epub 2010 Jun 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Berg L, McKeel DW Jr, Miller JP, Storandt M, Rubin EH, Morris JC, Baty J, Coats M, Norton J, Goate AM, Price JL, Gearing M, Mirra SS, Saunders AM. Clinicopathologic studies in cognitively healthy aging and Alzheimer’s disease: relation of histologic markers to dementia severity, age, sex, and apolipoprotein E genotype. Arch Neurol. 1998. Mar;55(3):326–35. doi: 10.1001/archneur.55.3.326. [DOI] [PubMed] [Google Scholar]
- 32.Morris JC, Roe CM, Xiong C, Fagan AM, Goate AM, Holtzman DM, Mintun MA. APOE predicts amyloid-beta but not tau Alzheimer pathology in cognitively normal aging. Ann Neurol. 2010. Jan;67(1):122–31. doi: 10.1002/ana.21843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Morris JC, Weintraub S, Chui HC, Cummings J, Decarli C, Ferris S, Foster NL, Galasko D, Graff-Radford N, Peskind ER, Beekly D, Ramos EM, Kukull WA. The Uniform Data Set (UDS): clinical and cognitive variables and descriptive data from Alzheimer Disease Centers. Alzheimer Dis Assoc Disord. 2006. Oct-Dec;20(4):210–6. doi: 10.1097/01.wad.0000213865.09806.92. [DOI] [PubMed] [Google Scholar]
- 34.Storandt M, Grant EA, Miller JP, Morris JC. Longitudinal course and neuropathologic outcomes in original vs revised MCI and in pre-MCI. Neurology. 2006. Aug 8;67(3):467–73. doi: 10.1212/01.wnl.0000228231.26111.6e. [DOI] [PubMed] [Google Scholar]
- 35.Kunkle BW, Carney RM, Kohli MA, et al. Targeted sequencing of ABCA7 identifies splicing, stop-gain and intronic risk variants for Alzheimer disease. Neurosci Lett. 2017. May 10;649:124–129. doi: 10.1016/j.neulet.2017.04.014. Epub 2017 Apr 8. [DOI] [PubMed] [Google Scholar]
- 36.McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology. 1984. Jul;34(7):939–44. doi: 10.1212/wnl.34.7.939. [DOI] [PubMed] [Google Scholar]
- 37.Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007. Sep;81(3):559–75. doi: 10.1086/519795. Epub 2007 Jul 25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Das S, Forer L, Schönherr S, Sidore C, Locke AE, Kwong A, Vrieze SI, Chew EY, Levy S, McGue M, Schlessinger D, Stambolian D, Loh PR, Iacono WG, Swaroop A, Scott LJ, Cucca F, Kronenberg F, Boehnke M, Abecasis GR, Fuchsberger C. Next-generation genotype imputation service and methods. Nat Genet. 2016. Oct;48(10):1284–1287. doi: 10.1038/ng.3656. Epub 2016 Aug 29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Conomos MP, Miller MB, Thornton TA. Robust inference of population structure for ancestry prediction and correction of stratification in the presence of relatedness. Genet Epidemiol. 2015. May;39(4):276–93. doi: 10.1002/gepi.21896. Epub 2015 Mar 23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS Genet. 2006. Dec;2(12):e190. doi: 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cavalli-Sforza LL. Human evolution and its relevance for genetic epidemiology. Annu Rev Genomics Hum Genet. 2007;8:1–15. doi: 10.1146/annurev.genom.8.080706.092403. [DOI] [PubMed] [Google Scholar]
- 42.Delaneau O, Marchini J; 1000 Genomes Project Consortium; 1000 Genomes Project Consortium. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat Commun. 2014. Jun 13;5:3934. doi: 10.1038/ncomms4934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.1000 Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini JL, McCarthy S, McVean GA, Abecasis GR. A global reference for human genetic variation. Nature. 2015. Oct 1;526(7571):68–74. doi: 10.1038/nature15393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Maples BK, Gravel S, Kenny EE, Bustamante CD. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am J Hum Genet. 2013. Aug 8;93(2):278–88. doi: 10.1016/j.ajhg.2013.06.020. Epub 2013 Aug 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Han B, Eskin E. Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. Am J Hum Genet. 2011. May 13;88(5):586–98. doi: 10.1016/j.ajhg.2011.04.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sariya S, Felsky D, Reyes-Dumeyer D, Lali R, Lantigua RA, Vardarajan B, Jiménez-Velázquez IZ, Haines JL, Shellenberg GD, Pericak-Vance MA, Paré G, Mayeux R, Tosto G. Polygenic Risk Score for Alzheimer’s Disease in Caribbean Hispanics. Ann Neurol. 2021. Sep;90(3):366–376. doi: 10.1002/ana.26131. Epub 2021 Jun 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grinde KE, Brown LA, Reiner AP, Thornton TA, Browning SR. Genome-wide Significance Thresholds for Admixture Mapping Studies. Am J Hum Genet. 2019. Mar 7;104(3):454–465. doi: 10.1016/j.ajhg.2019.01.008. Epub 2019 Feb 14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gautam Y, Ghandikota S, Chen S, Mersha TB. PAMAM: Power analysis in multiancestry admixture mapping. Genet Epidemiol. 2019. Oct;43(7):831–843. doi: 10.1002/gepi.22216. Epub 2019 Jun 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Martin ER, Tunc I, Liu Z, Slifer SH, Beecham AH, Beecham GW. Properties of global- and local-ancestry adjustments in genetic association tests in admixed populations. Genet Epidemiol. 2018. Mar;42(2):214–229. doi: 10.1002/gepi.22103. Epub 2017 Dec 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Buzkova P, Lumley T, Rice K. Permutation and parametric bootstrap tests for gene-gene and gene-environment interactions. Ann Hum Genet. 2011. Jan;75(1):36–45. doi: 10.1111/j.1469-1809.2010.00572.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017. Nov 28;8(1):1826. doi: 10.1038/s41467-017-01261-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wang K, Li M, Hakonarson H. (2010) ANNOVAR: Functional annotation of genetic variants from next-generation sequencing data Nucleic Acids Research, 38:e164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019. Jan 8;47(D1):D886–D894. doi: 10.1093/nar/gky1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, Karczewski KJ, Park J, Hitz BC, Weng S, Cherry JM, Snyder M. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012. Sep;22(9):1790–7. doi: 10.1101/gr.137323.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Griswold AJ, Sivasankaran SK, Van Booven D, Gardner OK, Rajabli F, Whitehead PL, Hamilton-Nelson KL, Adams LD, Scott AM, Hofmann NK, Vance JM, Cuccaro ML, Bush WS, Martin ER, Byrd GS, Haines JL, Pericak-Vance MA, Beecham GW. Immune and Inflammatory Pathways Implicated by Whole Blood Transcriptomic Analysis in a Diverse Ancestry Alzheimer’s Disease Cohort. J Alzheimers Dis. 2020;76(3):1047–1060. doi: 10.3233/JAD-190855. [DOI] [PubMed] [Google Scholar]
- 56.Moreno-Grau S, de Rojas I, Hernández I, Quintela I, Montrreal L, Alegret M, Hernández-Olasagarre B, Madrid L, González-Perez A, Maroñas O, Rosende-Roca M, Mauleón A, Vargas L, Lafuente A, Abdelnour C, Rodríguez-Gómez O, Gil S, Santos-Santos MÁ, Espinosa A, Ortega G, Sanabria Á, Pérez-Cordón A, Cañabate P, Moreno M, Preckler S, Ruiz S, Aguilera N, Pineda JA, Macías J, Alarcón-Martín E, Sotolongo-Grau O; GR@ACE consortium; DEGESCO consortium; Alzheimer’s Disease Neuroimaging Initiative, Marquié M, Monté-Rubio G, Valero S, Benaque A, Clarimón J, Bullido MJ, García-Ribas G, Pástor P, Sánchez-Juan P, Álvarez V, Piñol-Ripoll G, García-Alberca JM, Royo JL, Franco E, Mir P, Calero M, Medina M, Rábano A, Ávila J, Antúnez C, Real LM, Orellana A, Carracedo Á, Sáez ME, Tárraga L, Boada M, Ruiz A. Genome-wide association analysis of dementia and its clinical endophenotypes reveal novel loci associated with Alzheimer’s disease and three causality networks: The GR@ACE project. Alzheimers Dement. 2019. Oct;15(10):1333–1347. doi: 10.1016/j.jalz.2019.06.4950. Epub 2019 Aug 28. [DOI] [PubMed] [Google Scholar]
- 57.Aizawa H, Sekine Y, Takemura R, Zhang Z, Nangaku M, Hirokawa N. Kinesin family in murine central nervous system. J Cell Biol. 1992. Dec;119(5):1287–96. doi: 10.1083/jcb.119.5.1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chevalier-Larsen E, Holzbaur EL. Axonal transport and neurodegenerative disease. Biochim Biophys Acta. 2006. Nov-Dec;1762(11-12):1094–108. doi: 10.1016/j.bbadis.2006.04.002. Epub 2006 Apr 19. [DOI] [PubMed] [Google Scholar]
- 59.Witke W. The role of profilin complexes in cell motility and other cellular processes. Trends Cell Biol. 2004. Aug;14(8):461–9. doi: 10.1016/j.tcb.2004.07.003. [DOI] [PubMed] [Google Scholar]
- 60.Wu CH, Fallini C, Ticozzi N, Keagle PJ, Sapp PC, Piotrowska K, Lowe P, Koppers M, McKenna-Yasek D, Baron DM, Kost JE, Gonzalez-Perez P, Fox AD, Adams J, Taroni F, Tiloca C, Leclerc AL, Chafe SC, Mangroo D, Moore MJ, Zitzewitz JA, Xu ZS, van den Berg LH, Glass JD, Siciliano G, Cirulli ET, Goldstein DB, Salachas F, Meininger V, Rossoll W, Ratti A, Gellera C, Bosco DA, Bassell GJ, Silani V, Drory VE, Brown RH Jr, Landers JE. Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis. Nature. 2012. Aug 23;488(7412):499–503. doi: 10.1038/nature11280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Broce IJ, Tan CH, Fan CC, Jansen I, Savage JE, Witoelar A, Wen N, Hess CP, Dillon WP, Glastonbury CM, Glymour M, Yokoyama JS, Elahi FM, Rabinovici GD, Miller BL, Mormino EC, Sperling RA, Bennett DA, McEvoy LK, Brewer JB, Feldman HH, Hyman BT, Pericak-Vance M, Haines JL, Farrer LA, Mayeux R, Schellenberg GD, Yaffe K, Sugrue LP, Dale AM, Posthuma D, Andreassen OA, Karch CM, Desikan RS. Dissecting the genetic relationship between cardiovascular risk factors and Alzheimer’s disease. Acta Neuropathol. 2019. Feb;137(2):209–226. doi: 10.1007/s00401-018-1928-6. Epub 2018 Nov 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Decuypere JP, Parys JB, Bultynck G. Regulation of the autophagic bcl-2/beclin 1 interaction. Cells. 2012. Jul 6;1(3):284–312. doi: 10.3390/cells1030284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Ferreiro E, Eufrásio A, Pereira C, Oliveira CR, Rego AC. Bcl-2 overexpression protects against amyloid-beta and prion toxicity in GT1–7 neural cells. J Alzheimers Dis. 2007. Nov;12(3):223–8. doi: 10.3233/jad-2007-12303. [DOI] [PubMed] [Google Scholar]
- 64.Rohn TT, Vyas V, Hernandez-Estrada T, Nichol KE, Christie LA, Head E. Lack of pathology in a triple transgenic mouse model of Alzheimer’s disease after overexpression of the anti-apoptotic protein Bcl-2. J Neurosci. 2008. Mar 19;28(12):3051–9. doi: 10.1523/JNEUROSCI.5620-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Farrer LA. Intercontinental epidemiology of Alzheimer disease: a global approach to bad gene hunting. JAMA. 2001. Feb 14;285(6):796–8. doi: 10.1001/jama.285.6.796. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.