Abstract
The cell and gene therapy industry has employed the same plasmid technology for decades in vaccination, cell and gene therapy, and as a raw material in viral vector and RNA production. While canonical plasmids contain antibiotic resistance markers in bacterial backbones greater than 2,000 base pairs, smaller backbones increase expression level and durability and reduce the cell-transfection-associated toxicity and transgene silencing that can occur with canonical plasmids. Therefore, the small backbone and antibiotic-free selection method of Nanoplasmid vectors have proven to be a transformative replacement in a wide variety of applications, offering a greater safety profile and efficiency than traditional plasmids. This review provides an overview of the Nanoplasmid technology and highlights its specific benefits for various applications with examples from recent publications.
Keywords: MT: Delivery Strategies, plasmid, minicircle, Nanoplasmid, cell therapy, gene therapy, CRISPR-cas9, homology-directed repair, adeno-associated virus, AAV
Graphical abstract
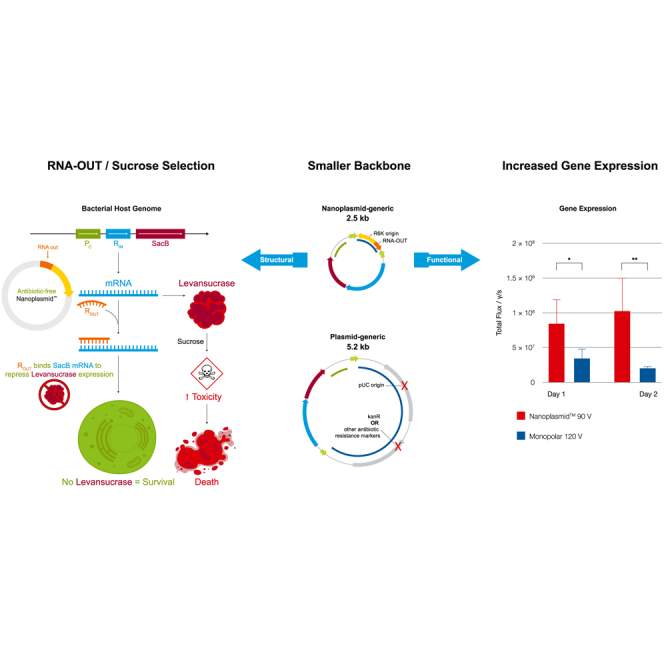
Paez and Williams discuss the Nanoplasmid vector structural differences, including its small backbone and antibiotic-free selection system, resulting in significant performance and safety benefits compared with traditional plasmids. Nanoplasmid’s design reflects the progress of the cell and gene therapy field, not considered during traditional plasmid development in the 1980s.
Introduction
Plasmid DNA is an integral component of cell and gene therapy products. Plasmid technology is utilized as transposon or CRISPR-Cas9 homology-directed repair (HDR) template vectors for cell therapy, non-viral gene therapy vectors, and as a raw material to produce adeno-associated virus (AAV) or lentiviral viral vectors or to produce mRNA vaccines or therapies. For example, plasmid DNA is used to produce AAV virus in two or three plasmid cotransfection systems utilizing adenovirus E1A- and E1B-expressing cell lines such as HEK293. The more commonly utilized three plasmid AAV system comprises: (1) a payload encoding cis plasmid that will be packaged into the AAV capsid, (2) a rep cap-expressing plasmid encoding replicative (rep) and the capsid serotype (cap) genes, and (3) a helper plasmid containing adenovirus genes.1
The cell and gene therapy field has evolved significantly over the past three decades. The ability to therapeutically alter genetic composition via insertion, correction, or removal of a target gene in a disease has created a new paradigm in medicine. Advancements in sequencing technology and gene therapy modalities such as AAV and gene editing modalities such as CRISPR-Cas9 have spearheaded the field. The field of AAV, in particular, is continuing to grow from its first clinical trial for cystic fibrosis patients in 19962 to its first Food and Drug Administration (FDA)-approved AAV-based therapy in 2017.3 The total estimated number of AAV clinical trials ranges from 250 to 300 worldwide,4 and current estimates of active clinical trials using AAV range between 25 and 30.5 However, this figure is expected to grow on the heels of the success of the first European Medical Agency (EMA) approval in 2012, and FDA approval in 2017.5,6
However, in contrast, specific tools used in molecular cloning remain antiquated and limit the progress and safety of the expanding treatment modalities that the cell and gene therapy field has to offer. For example, most cell and gene therapy vectors use canonical plasmid backbones developed in the 1980s. These backbones are typically 2,000–3,000 bp in size and encode antibiotic resistance markers which are discouraged by regulatory agencies. Recently it has been demonstrated that smaller backbone vectors have lower toxicity, reduced payload inactivation, and dramatically fewer metabolic perturbations than canonical plasmids. Antibiotic-free small backbone plasmids such as Nanoplasmid vectors have shown great utility in modifying the genomes and epigenomes of cells for therapeutic purposes.
Plasmid vector: Structural issues
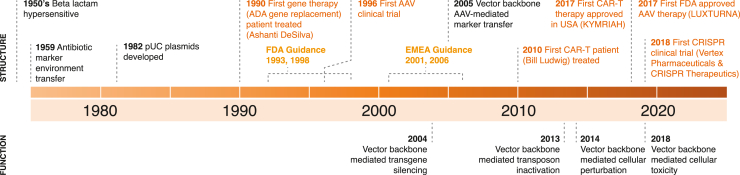
Canonical cloning vector plasmids were developed in the 1980s and combined the promiscuous pUC replication origin with β-lactam ampicillin antibiotic7 or kanamycin antibiotic8 resistance marker genes in a 2.7-kb or larger bacterial backbone (i.e., the region encoding the bacterial replication origin and selectable marker) vector. These vectors were designed for cloning and were not intended for use in gene therapy: promiscuous replication origin-mediated antibiotic marker transfer to the environment (horizontal gene transfer [HGT]) was well known at the time9,10,11,12 as was β-lactam (e.g., penicillin, ampicillin, cephalosporins, etc.) antibiotic-mediated serious hypersensitivity reactions in patients.13,14 However, due to ubiquitous use in cloning laboratories, these pUC-derived antibiotic marker backbones have been incorporated into the plasmids utilized to produce most non-viral and viral gene therapy products that were developed in the 1990s onward (Figure 1).
Figure 1.
A historical perspective on plasmids and their role in cell and gene therapy
Numbered timeline landmarks in the field with given citations (black text indicates vector backbone composition-relevant scientific findings; orange denotes vector regulatory agency guidances; pink denotes vector clinical trial milestones) are as follows: 1950s, β-lactam hypersensitivity in patients13,14; 1959, antibiotic marker environment transfer9,10,11,12; 1982, pUC plasmids developed7,8; 1996, first AAV clinical trial2; 2004, vector backbone-mediated transgene silencing15; 2005, vector backbone AAV-mediated marker transfer16; 2013, vector backbone-mediated transposon inactivation17; 2014, vector backbone-mediated cellular perturbation18; 2017, first FDA approval of an AAV therapy3; 2018, vector backbone-mediated cellular toxicity.19
These structural issues with pUC-derived antibiotic marker backbones were highlighted to the gene and cell therapy communities by regulatory guidance from the FDA20,21 and EMA22,23 (Figure 1) and continue to be highlighted as concerns in current FDA24 and EMA guidelines.25 Consistent with these concerns, HGT to freshwater environmental organisms has been documented due to the unintended release of recombinant pUC-origin plasmid vectors from laboratory or manufacturing facilities.26
AAV-mediated antibiotic marker transfer: an additional concerning form of gene transfer, AAV virus-mediated plasmid backbone transfer to patients, was discovered in 200516 and more recently characterized as ITR-mediated high-frequency reverse packaging of the bacterial backbone during production. This results in significant bacterial backbone-encoded antibiotic resistance marker packaging into AAV viruses, and subsequent marker gene transfer to the patient during therapy1,27,28 (Table 1). Schnödt et al.27 and Tai et al.28 reported AAV contamination with bacterial backbone using AAV genome population sequencing (AAV-GPseq). Gray29 also reports that AAV helper plasmid antibiotic resistance markers are packaged into viral particles, demonstrating the need to remove antibiotic markers from AAV helper plasmids as well as the AAV vector.
Table 1.
AAV virus-mediated marker transfer to patients
| Vector platform | pUC plasmid (>2 kb pUC origin-antibiotic marker spacer region) | Minicircle (MC) retrofit (<100 bp spacer region) |
Result-performance | Result-antibiotic marker gene transfer |
|---|---|---|---|---|
| Self-complementary AAV | pAAV-scGFP | MC.AAVscGFP | MC up to 30-fold improved transducing units27,a | plasmid backbone antibiotic resistance marker packaged in up to 26.1% viral particles27 potential for genome integration in transduced cells |
| Single-stranded AAV | AAV2-ssGFP | MC.AAVssGFP | MC up to 3-fold improved transducing units27,a | plasmid backbone antibiotic resistance marker packaged in up to 2.9%27 or 3%29 of viral particles potential for genome integration in transduced cells |
No improvement in viral titer compared with plasmid was observed with linear Doggybone DNA single-stranded AAV vector retrofits.30 This suggests that short backbone minicircle vector improvement compared with >2 kb pUC-antibiotic marker plasmid backbone vectors require a short backbone circular vector rather than a linear vector such as Doggybone vectors.
Gray29 describes adding a large spacer region to move the bacterial backbone-encoded selection marker away from ITRs. The resultant vector, pAAV CMV GFP is 11 kb, compared with 5–6 kb for a standard plasmid AAV vector. Consequently, it has a reduced transfection efficiency.31
Alternatives to antibiotic-based selection markers
The majority of antibiotic-free selection markers for plasmid maintenance in bacterial cells are protein-based systems, such as those designed for (1) plasmid-borne complementation of an essential gene that has been deleted from the chromosome (auxotrophy complementation) and (2) toxin-antitoxin-based plasmid maintenance systems using post-segregational killing in which a plasmid-borne antitoxin is used to prevent a host cell line-encoded toxin from killing the cell.32 However, when expressed, vector backbone-encoded proteins in gene therapy products may interact with cellular components to alter cell functions or viability and will likely induce an immune response that may be detrimental to the patient.33 The later issue is shared with antibiotic selection markers and will be a concern with AAV vectors due to high-frequency reverse packaging of the bacterial backbone during production as described above (Table 1) and with other gene therapy vectors due to the introduction of the marker into cells during transfection (Table 2).
Table 2.
Minicircle applications with various viral and non-viral vector platforms
| Vector platform | pUC plasmid (>2 kb pUC origin-antibiotic marker spacer region) | Minicircle (MC) retrofit (<100 bp spacer region) | Result-performance | Result-antibiotic marker gene transfer |
|---|---|---|---|---|
| Sleeping Beauty transposon gene integration vector | SB puroR reporter plasmid | SB puroR reporter MC | MC 2-fold increased transposition rate into established cell lines17 | plasmid backbone antibiotic resistance potential expression or genome integration in T cells |
| SB CD19-CAR | MC: SB CD19 CAR | MC 4-fold increased yield of CD19-CAR T cells; reduced transfection associated toxicity with nucleofection delivery34 | ||
| SB | MC: SB (218 bp MC Backbone) |
MC 6- to 7-fold increased transposition into human hematopoietic stem cells; reduced transfection associated toxicity with nucleofection delivery19 | ||
| CRISPR-Cas9 gene editing vectora35 | 3.1 kb 2A-puro/gRNA-1 | 0.85 kb 2A-puro/gRNA-1 | MC 6-fold increased targeted integration into HeLa cells after lipofection of Cas9-2A-puro/gRNA-1 | plasmid backbone antibiotic resistance potential expression or genome integration |
| Direct cell therapy vector36 | pANGPT1 | MC-ANGPT1 | MC 3.7-fold greater increase in ANGPT1 protein expression after electroporation of mesenchymal stem cells | plasmid backbone antibiotic resistance potential expression or genome integration in stem cells |
| Direct gene therapy vector37 | pRSV-hAAT-bpA pEF1α-hFIX-hGHpA |
MC-RSV-hAAT-bpA MC-EF1α-hFIX-hGHpA |
MC > 1 log increased extended duration expression (>20 days post hydrodynamic delivery to mouse liver) (reduced gene silencing)b | plasmid backbone antibiotic resistance marker potential expression or genome integration in liver cells |
Homology-independent “replace editing” using non-homologous end-joining pathway.
Lu et al.37 also report that gene silencing occurs with ≥1 kb random DNA sequences, while shorter spacers ≤500 bp similarly extended duration expression as with minicircle vectors.
Eliminating protein and antibiotic markers using non-coding RNA markers
Ideally, bacterial selection would be mediated by a non-coding selection marker. These have been developed as: (1) plasmid-borne non-coding mRNA that functions by RNA/RNA interaction to inhibit translation of a target chromosomally expressed gene. (2) Plasmid-borne tRNA gene that functions to suppress a nonsense codon mutated chromosomally expressed essential gene38; or (3) by operator repressor titration, wherein a plasmid contains several operator sequences that titrate the repressor, allowing expression of a repressor-regulated selectable marker gene.32
One of these RNA/RNA interaction non-coding selection marker systems is the RNA-OUT marker system39 (Figure 2). This selection process eliminates antibiotic marker transfer when used for cell and gene therapy, and it benefits from a high yielding (up to 2 g plasmid per liter fermentation culture) and robust manufacturing process40,41 that has been scaled to the 200-g lot scale in cGMP manufacture (J.W., personal communication). RNA-OUT vectors have also been tested without safety issues in multiple clinical trials; for example, the EYS606 eye gene therapy for noninfectious uveitis,42 INVAC-1 telomerase DNA vaccine for advanced solid tumors,43 CryJ2-LAMP allergy DNA vaccine for Japanese red cedar pollinosis,44 and COR-1 herpes simplex virus DNA vaccine.45,46
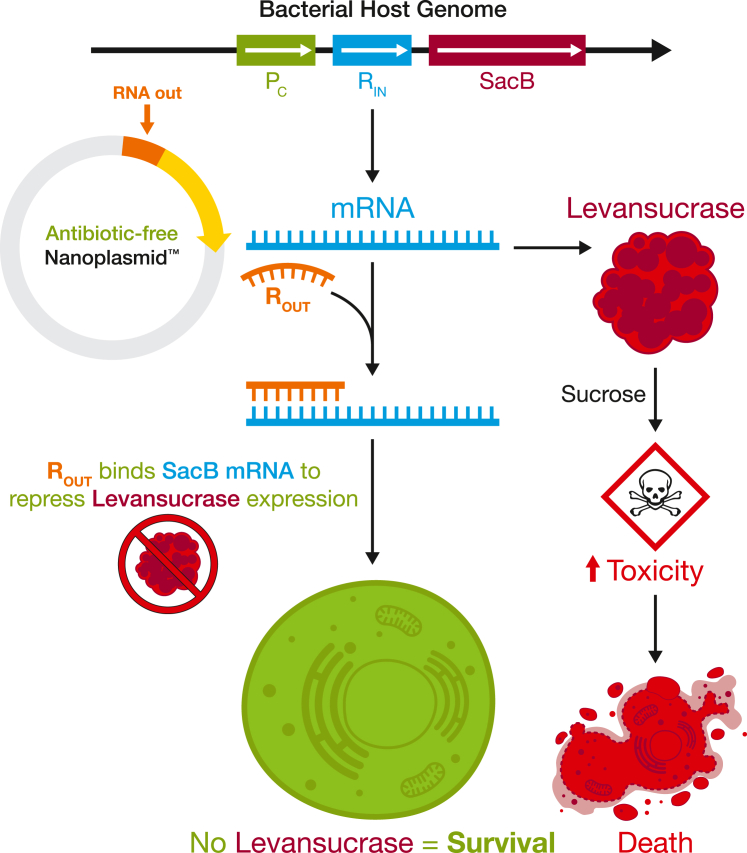
Figure 2.
The Nanoplasmid RNA-OUT sucrose-based selection system eliminates the use of the antiquated antibiotic selection systems seen in traditional plasmid vectors
The proprietary bacterial host strain expresses the SacB gene from a constitutive promoter (Pc), which encodes for levansucrase—an enzyme that reacts with sucrose to create fructose polymers (levan) toxic to bacteria but not humans. Bacterial cells successfully transformed with the Nanoplasmid vector will express RNA-OUT (ROUT), which inhibits the expression of levansucrase at the mRNA level by annealing to the complementary 5′ UTR-encoded RIN sequence, allowing survival and propagation.
Plasmid vector: Functional issues
In the last 20 years, functional drawbacks of pUC-derived antibiotic marker backbones have been discovered, including vector backbone-mediated transgene silencing, insert inactivation, cellular metabolic perturbation, and toxicity after vector transfection into cells (Table 2).
The first functional limitation of canonical plasmid vectors discovered was that transgene expression duration from plasmid vectors is reduced due to promoter inactivation mediated by the bacterial backbone of the vector.15,47 This results in short-duration transgene expression, also known as transgene silencing. A strategy to improve transgene expression duration involves removing the plasmid’s bacterial backbone. One example of this approach in the field was the invention of minicircle vectors. They were first developed in 1997, and their design is devoid of a bacterial backbone, resulting in improved transgene expression and duration when used as a cell and gene therapy vector.15,48,49
Various investigators have subsequently identified that minicircle vectors are also superior to plasmid vectors when used in the production of AAV vectors (resulting in improved transducing unit titers, Table 1) or used as transposon vectors (resulting in increased transposition, Table 2), and CRISPR-Cas9 vectors (increased gene editing, Table 2).50
A drawback of the minicircle strategy lies in how the small bacterial backbone vectors are manufactured. They are produced in a specialized E. coli system that allows the bacterial and eukaryotic backbones to be separated and re-circularized by recombinases on specific recognition sequences on the plasmid. In some methods, a restriction enzyme is then utilized to digest the bacterial backbone circle at a unique site to eliminate this difficult-to-remove contaminant. In one example, the optimal manufacture of minicircle vectors yielded only 5 mg of minicircle per liter culture.51 This laborious manufacturing process makes minicircle vectors expensive and impractical for most commercial applications.
The creation of the Nanoplasmid platform: Next-generation plasmid technology
In minicircle vectors, the eukaryotic region 3′ end is covalently linked to the eukaryotic region 5′ end through a short spacer, typically less than 200 bp comprised of the recombined attachment sites. This linkage or spacer region joining the eukaryotic 5′ and 3′ ends were found to tolerate a much longer spacer sequence in a landmark study published by Lu et al.37 They elegantly discovered that long spacers of ≥1 kb in length resulted in transgene expression silencing in vivo while shorter spacers of ≤500 bp exhibited similar improved transgene expression patterns seen in conventional minicircle DNA vectors. These findings indicated that it was not a specific DNA sequence or structure found in the backbone of a plasmid but the length of the sequence in the spacer that would determine transgene expression levels. This phenomenon was accounted for in the design of Nanoplasmid as its backbone size met the studied 500-bp limit, thereby reducing transgene silencing and gaining the performance benefits observed with minicircles while retaining manufacturing yields seen with conventional plasmid systems.
Nanoplasmid vectors use the RNA-OUT antibiotic-free marker and have additional intrinsic advantages (Table 3). Most notably, the vectors also contain a specialized bacterial R6K replication origin in place of the traditional pUC replication origin, making these vectors replication-incompatible with native organisms. This is an additional safety factor since Nanoplasmids can only replicate within the specialized heat-inducible R6K replication protein encoding E. coli host strain used for manufacturing,52 dramatically reducing the risk of HGT.
Table 3.
DNA technology progression—the birth of next-generation plasmid technology
| Plasmid | Minicircle | Nanoplasmid | |
|---|---|---|---|
| Bacterial region size | >2,000 bp | 100 bpa | 500 bpa |
| Toxicity | increased transfection toxicity | reduced transfection toxicitya | reduced transfection toxicitya |
| Silencing | increased transgene silencing | reduced transgene silencinga | reduced transgene silencinga |
| Transgene expression | low transgene expression | low transgene expression | high transgene expressiona |
| Selection | antibiotic | antibiotic | sucrosea |
| Type of selection marker | protein | protein | RNAa |
| Manufacturing yield | high yielda | low yield | high yielda |
| Scalability of manufacturing | scalablea | complex | scalablea |
| Safety | pUC promiscuous | pUC promiscuous | R6K strain restricteda |
Nanoplasmid encompasses all the performance benefits of minicircles while improving manufacturing and safety profiles for cell and gene therapy applications.
The advantage over multiple technologies comparatively.
In addition, the Nanoplasmid backbone has no coding capacity, eliminating the risk of AAV virus or cell or gene therapy vector-mediated bacterial backbone-encoded protein marker transfer to patients.
Small backbone benefit: Reduced vector inactivation and improved transfection efficiency with Nanoplasmid vectors
As expected from Lu et al.,37 Nanoplasmid vectors exhibit reduced transgene silencing equivalent to minicircle vectors and mini-intronic plasmid (MIP) vectors (short spacer region vectors in which the bacterial backbone is encoded within an intron)53 after hydrodynamic delivery to the murine liver.54 Gene silencing is hypothesized to be the result of vector inactivation through heterochromatin formation on large >1-kb unexpressed plasmid backbones.47,55 Interestingly, both Nanoplasmid and MIP vectors exhibit up to 10-fold higher overall expression levels than conventional plasmids or minicircle vectors.53,54 With plasmids, this is hypothesized to be due to reduced heterochromatinization leading to increased nuclear localization of “non-inactivated” vectors (i.e., reduced vector inactivation). Minicircle vectors may have much lower expression due to the technical difficulties associated with the manufacturing platform,51 which may result in lower purity due to reduced product titer per cell,56 or reduced supercoiling due to DNA repair enzyme depletion. For example, while supercoiling is preserved during recombinase-mediated intramolecular excision of the minicircle and plasmid backbone,57,58 the recombination product is a catenated DNA circle that requires processing by topoisomerase IV into separate minicircle and plasmid backbones.59 This requires carefully controlled growth to ensure cell viability throughout production since depletion of a DNA repair enzyme can prevent decatenation or physiological supercoiling of the recombined vector.60
Robust transgene expression up to 10-fold higher than plasmid vectors has been reported with Nanoplasmid vectors after in vitro delivery to rat tendon or mouse melanoma cells, or in vivo electroporation delivery to rat skin or heart (Figure 3).61,62 Nanoplasmid has also shown robust expression after nanoparticle tail vein delivery,63 hydrodynamic retrograde intrabiliary injection in pigs,64 and after nanoparticle delivery to tumor cells in vitro and in vivo,65 or electroporation delivery of DNA-encoded antibody therapeutics to muscle.66 An immune-oncology gene therapy named EG-70 (also known as detalimogen voraplasmid) is currently in a phase I/II trial for BCG refractory non-muscle invasive bladder cancer (NCT04752722). EG-70 consists of a Nanoplasmid encoding both retinoic acid-inducible gene I (RIG-I) agonist67 and IL-1268 packaged in a proprietary muco-penetrating nanoparticle formulation; and it was administered intravesically to transfect and stimulate the immune system locally in the bladder. No dose-related adverse events were reported, while 67% of patients achieved complete response after one cycle of EG-70.69
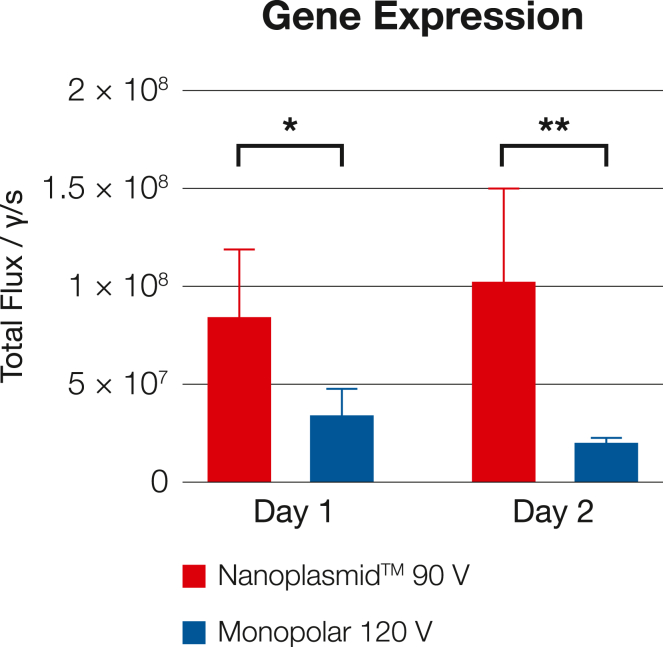
Figure 3.
Nanoplasmid improves gene electrotransfer in vivo to rat myocardial cells at lower voltages
Rat myocardial cells were electroporated in vivo using electrotransfer with either Nanoplasmid (red) at 90 V or traditional plasmids (blue) at 120 V. y axis: total flux (photons/s) measures gene expression by bioluminescence. (∗=p<0.01, ∗∗= p<0.001) This figure was adapted from Boye et al.61 “Reduction of plasmid vector backbone length enhances reporter gene expression,” 2022, Bioelectrochemistry, 144, 107981, https://doi.org/10.1016/j.bioelechem.2021.107981 CC BY-NC -ND 4.0.
Improved Nanoplasmid-encoded transgene expression via transfection in the context of DNA vaccination will translate to increased antigen production, helping prime antigen-presenting cells with the cognate antigen targeted for vaccination.70 This potentiates robust immune responses mediated by using the Nanoplasmid vector as a DNA vaccine. Examples of this have been reported in several publications and a clinical trial. A SARS-CoV-2 DNA vaccine encoding both RIG-I agonist67 and full-length SARS-CoV-2 spike protein encapsulated in proteolipid vehicles formulated with a fusion-associated small transmembrane protein elicited potent neutralizing antibody and significant T cell responses in non-human primates.71 This vaccine has produced positive phase 1 safety, tolerance, and immunological data (Covigenix VAX-001; NCT04591184). Preclinical studies of Nanoplasmid DNA vaccine vectors for Influenza,72 Venezuelan equine encephalitis virus (VEEV), and Ebola virus,73 and a DNA-based vaccine protecting against parasitic antigens expressed by T. cruzi74 all demonstrated improved immune responses compared with plasmids. For example, VEEV Nanoplasmid vectors increased antibody response and significantly increased protection against VEEV challenge compared with a previously optimized pWRG7077-based plasmid comparator.73 In addition, Nanoplasmid delivered T. cruzi Chagas disease TcG2 and TcG4 antigens more efficiently protected infected mice than a pCDNA3.1 plasmid comparator.74
Reported improvements of CAR-T/CAR-NK cell production with Nanoplasmid vectors
Minicircle vectors have recently been shown to have reduced cellular toxicity compared with canonical plasmid vectors.19,75 Toxicity is potentially due to heterochromatin-mediated “danger signal” activation by inactivated transfected plasmid DNA. Consistent with this hypothesis, reduced cellular perturbation at the proteome level has been reported after in vivo transfection of minicircle vectors compared with plasmid vectors,18 and reduced cellular perturbation at the transcriptome level has been reported after in vitro transfection of minicircle or Nanoplasmid vectors compared with plasmid76,77 or lentiviral78 vectors.
Compared with plasmid comparators, superior performance has been reported for non-viral chimeric antigen receptor (CAR) T cell production using Nanoplasmid in the PiggyBac transposon vector system,79,80 and as a DNA template for CRISPR-Cas9 HDR-mediated gene editing (Figure 4).81 Robust non-viral CAR-T and CAR-NK cells were also obtained using a Nanoplasmid TcBuster transposon system.82,83,84 Improved performance may be due to reduced post-transfection cellular toxicity, combined with reduced vector inactivation resulting in higher gene integration into healthier cells. Consistent with this phenotype, PiggyBac-modified CAR-T cells using Nanoplasmid as a template displayed improved transposition efficiency and reduced toxicity in manufacturing compared with a traditional plasmid system. These results showed a shorter manufacturing timeline and a significant increase in stem cell memory T cells, a highly desired and effective phenotype in the production of CAR-T cells in preclinical and clinical studies. Furthermore, patients in a small study who received CAR-T cells manufactured with Nanoplasmid in the P-BCMA-101 clinical trial (NCT03288493) experienced a robust T cell expansion and expression of the CAR in vivo for over 18 months. This resulted in an increased overall response rate (ORR) (67% standard plasmid, 100% Nanoplasmid; ORR) and a 100% complete response rate (CR) (0% standard plasmid, 100% Nanoplasmid; CR) in patients receiving CAR-T cells manufactured with Nanoplasmid.79,80 Oh et al.81 published a comparison of linear double-stranded DNA (dsDNA), a traditional pUC plasmid, and Nanoplasmid employed as a CRISPR-Cas9 HDR donor template for the production of CAR-T cells. In this report, they found that Nanoplasmid yielded twice the number of edited cells (CAR-T cells) compared with the traditional pUC plasmid and 3-fold compared with dsDNA when using 4 μg of the template.
Figure 4.
Nanoplasmid serves as a template for CRISPR-Cas9 HDR knockin—improving 2- and 3-fold of edited CD8+ CAR-T cells yield over conventional plasmids and dsDNA, respectively
Plasmid-based donor templates enable efficient nonviral gene editing of the TRAC locus in primary T cells. Titration of linear dsDNA donor template. (A) Diagram of linear dsDNA knockin construct TRAC-mNG. (B) Bar graphs depicting knockin efficiency, cell viability, total cell recovery, and edited cell recovery (mNG-positive cells) 3 days after electroporation with 1, 2, 4, 6, or 8 μg of linear dsDNA donor template together with Cas9-RNP targeting the TRAC locus. Circles represent individual donors; bars represent median values with range (n = 4). Titration of pUC57 traditional plasmid template. (C) Diagram of pUC57 knockin construct TRAC-mNG. (D) Bar graphs showing the frequency of CD8+ T cells expressing mNG, cell viability, total cell recovery, and edited cell recovery (mNG-positive cells) 3 days after electroporation with 1, 2, 4, 6, or 8 μg of pUC57 plasmid donor template together with Cas9-RNP targeting the TRAC locus. Circles represent individual donors; bars represent median values with range (n = 4). Titration of Nanoplasmid vector template. (E) Diagram of Nanoplasmid knockin construct TRAC mNG. (F) Bar graphs showing the frequency of CD8+ T cells expressing mNG, total cell recovery, and edited cell recovery (mNG-positive cells) 3 days after electroporation with 1, 2, 4, 6, or 8 μg of Nanoplasmid donor template together with Cas9-RNP targeting the TRAC locus. Circles represent individual donors; bars represent median values with range (n = 4). This experiment was performed twice. ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, ∗∗∗∗p < 0.0001 in RM one-way ANOVA with Geisser-Greenhouse correction. This figure was adapted from Oh et al.81 “High-efficiency nonviral CRISPR/cas9-mediated gene editing of human T cells using plasmid donor DNA.” Journal of Experimental Medicine. 2022; 219(5), https://doi.org/10.1084/jem.20211530 CC BY 4.0.
Improving performance, manufacturing, and patient safety
The Nanoplasmid vector system was born out of the progression of technology in the molecular biology field to meet the current demands for cell and gene therapy clinical applications. It encompasses the performance benefits of minicircles not seen in traditional plasmid vectors, including reduced transgene silencing and diminished post-transfection toxicity from its intelligently designed 500-bp bacterial backbone—a discovery utilized from landmark studies in the field indicating that backbone sizes of 1 kb or greater were impacting transgene expression. Nanoplasmid also improves upon the shortcomings that accompany the manufacturing of minicircles, delivering yields up to 2.4 g/L (J.W., personal communication) in optimized systems compared with the milligram quantities per liter seen in documented cases of minicircle production.
Nanoplasmid’s performance benefits are not the singular issue addressed with the design of this next-generation plasmid technology. This plasmid is R6K strain-dependent and has far less promiscuity than pUC vector systems mitigating HGT into the environment or the commensal bacteria of a patient’s skin, mucosal, or intestinal flora. Its RNA-OUT system utilizes a sucrose selection system that eliminates the possibility of antibiotic resistance marker transfer to patients when used in the context of gene or cell therapy, DNA vaccines, lentivirus, or AAV vector applications. The qualities of the Nanoplasmid vector meet or exceed the standards or recommendations of the governing medical agencies such as EMA and FDA while improving performance and increasing manufacturing capabilities in several applications, including AAV, lentivirus, transposon therapy, DNA vaccines, and as a CRISPR-Cas9 HDR donor template. Adoptions of this next-generation plasmid technology in translational research and development programs in the cell and gene therapy field will aid in creating safer therapeutic interventions for patients receiving tomorrow’s therapies.
Acknowledgments
We would like to thank Jessica Lewis, J.D., Ph.D., for reviewing the manuscript and Jim Bolluyt for preparing the graphics.
Author contributions
J.W. and P.P. wrote the manuscript.
Declaration of interests
J.W. and P.P. are employees of Aldevron. J.W. is an inventor on several patent applications and patents to protect Nanoplasmid manufacturing, compositions, and applications.
References
- 1.Pupo A., Fernández A., Low S.H., François A., Suárez-Amarán L., Samulski R.J. AAV vectors: The Rubik’s cube of human gene therapy. Mol. Ther. 2022;30:3515–3541. doi: 10.1016/J.YMTHE.2022.09.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Flotte T., Carter B., Conrad C., Guggino W., Reynolds T., Rosenstein B., Taylor G., Walden s., Wetzel R. A phase I study of an adeno-associated virus-CFTR gene vector in adult CF patients with mild lung disease. Hum. Gene Ther. 1996;7:1145–1159. doi: 10.1089/HUM.1996.7.9-1145. [DOI] [PubMed] [Google Scholar]
- 3.Russell S., Bennett J., Wellman J.A., Chung D.C., Yu Z.F., Tillman A., Wittes J., Pappas J., Elci O., McCague S., et al. Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet. 2017;390:849–860. doi: 10.1016/S0140-6736(17)31868-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.The platform vector gene therapies project: increasing the efficiency of adeno-associated virus gene therapy clinical trial startup | human gene therapy. https://www.liebertpub.com/doi/10.1089/hum.2020.259 [DOI] [PMC free article] [PubMed]
- 5.Kuzmin D.A., Shutova M.V., Johnston N.R., Smith O.P., Fedorin V.v., Kukushkin Y.S., van der Loo J.C.M., Johnstone E.C. The clinical landscape for AAV gene therapies. Nat. Rev. Drug Discov. 2021;20:173–174. doi: 10.1038/D41573-021-00017-7. [DOI] [PubMed] [Google Scholar]
- 6.Wang D., Tai P.W.L., Gao G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 2019;18:358–378. doi: 10.1038/S41573-019-0012-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982;19:259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- 8.Spratt B.G., Hedge P.J., te Heesen S., Edelman A., Broome-Smith J.K. Kanamycin-resistant vectors that are analogues of plasmids pUC8, pUC9, pEMBL8 and pEMBL9. Gene. 1986;41:337–342. doi: 10.1016/0378-1119(86)90117-4. [DOI] [PubMed] [Google Scholar]
- 9.Ochiai K., Yamanaka T., Kimura K., Sawada O. Studies on inheritance of drug resistance between Shigella strains and Escherichia coli strains. Nippon Iji Shimpo. 1959;1861:34–46. [Google Scholar]
- 10.Davies J., Smith D.I. Plasmid-determined resistance to antimicrobial agents. Annu. Rev. Microbiol. 1978;32:469–518. doi: 10.1146/ANNUREV.MI.32.100178.002345. [DOI] [PubMed] [Google Scholar]
- 11.Helinski D.R. R Factors: Infectious Multiple Drug Resistance. S. Falkow. Pion, London, 1975 (distributor, Academic Press, New York). xiv, 300 pp. + plates. $19.95. Pion Advanced Biochemistry Series, 4. Science. 1976;192:778. doi: 10.1126/SCIENCE.192.4241.778.A. [DOI] [PubMed] [Google Scholar]
- 12.O’Brien T.F., Ross D.G., Guzman M.A., Medeiros A.A., Hedges R.W., Botstein D. Dissemination of an antibiotic resistance plasmid in hospital patient flora. Antimicrob. Agents Chemother. 1980;17:537–543. doi: 10.1128/AAC.17.4.537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Idsoe O., Guthe T., Willcox R.R., de Weck A.L. Nature and extent of penicillin side-reactions, with particular reference to fatalities from anaphylactic shock. Bull. World Health Organ. 1968;38:159–188. [PMC free article] [PubMed] [Google Scholar]
- 14.Guthe T., Idsoe O., Willcox R.R. Untoward penicillin reactions. Bull. World Health Organ. 1958;19:427–501. [PMC free article] [PubMed] [Google Scholar]
- 15.Chen Z.Y., He C.Y., Meuse L., Kay M.A. Silencing of episomal transgene expression by plasmid bacterial DNA elements in vivo. Gene Ther. 2004;11:856–864. doi: 10.1038/SJ.GT.3302231. [DOI] [PubMed] [Google Scholar]
- 16.Chadeuf G., Ciron C., Moullier P., Salvetti A. Evidence for encapsidation of prokaryotic sequences during recombinant adeno-associated virus production and their in vivo persistence after vector delivery. Mol. Ther. 2005;12:744–753. doi: 10.1016/J.YMTHE.2005.06.003. [DOI] [PubMed] [Google Scholar]
- 17.Sharma N., Cai Y., Bak R.O., Jakobsen M.R., Schrøder L.D., Mikkelsen J.G. Efficient sleeping beauty DNA transposition from DNA minicircles. Mol. Ther. Nucleic Acids. 2013;2:e74. doi: 10.1038/MTNA.2013.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Geguchadze R., Wang Z., Zourelias L., Perez-Riveros P., Edwards P.C., Machen L., Passineau M.J. Proteomic profiling of salivary gland after nonviral gene transfer mediated by conventional plasmids and minicircles. Mol. Ther. Methods Clin. Dev. 2014;1:14007. doi: 10.1038/MTM.2014.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Holstein M., Mesa-Nuñez C., Miskey C., Almarza E., Poletti V., Schmeer M., Grueso E., Ordóñez Flores J.C., Kobelt D., Walther W., et al. Efficient non-viral gene delivery into human hematopoietic stem sells by minicircle sleeping beauty transposon vectors. Mol. Ther. 2018;26:1137–1153. doi: 10.1016/J.YMTHE.2018.01.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Food and Drug Administration (USA) 1993. Points to Consider in the Characterization of Cell Lines Use to Produce Biologics. [Google Scholar]
- 21.Food and Drug Administration (USA) 1998. Guidance for Human Somatic Cell Therapy and Gene Therapy. [Google Scholar]
- 22.EMEA . 2001. Note for Guidance on the Quality, Preclinical and Clinical Aspects of Gene Transfer Medicinal Products. [Google Scholar]
- 23.EMEA . 2006. Guideline on the Non-Clinical Studies Required Before First Clinical Use of Gene Therapy Medicinal Products. [Google Scholar]
- 24.Food and Drug Administration (USA) FDA; 2016. Recommendations for Microbial Vectors Used for Gene Therapy.https://www.fda.gov/regulatory-information/search-fda-guidance-documents/recommendations-microbial-vectors-used-gene-therapy [Google Scholar]
- 25.EMEA . 2018. Guideline on the Quality, Non-Clinical and Clinical Aspects of Gene Therapy Medicinal Products. [Google Scholar]
- 26.Chen J., Jin M., Qiu Z.G., Guo C., Chen Z.L., Shen Z.Q., Wang X.W., Li J.W. A survey of drug resistance bla genes originating from synthetic plasmid vectors in six Chinese rivers. Environ. Sci. Technol. 2012;46:13448–13454. doi: 10.1021/ES302760S. [DOI] [PubMed] [Google Scholar]
- 27.Schnödt M., Schmeer M., Kracher B., Krüsemann C., Espinosa L.E., Grünert A., Fuchsluger T., Rischmüller A., Schleef M., Büning H. DNA minicircle technology improves purity of Adeno-associated viral vector preparations. Mol. Ther. Nucleic Acids. 2016;5:e355. doi: 10.1038/MTNA.2016.60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tai P.W.L., Xie J., Fong K., Seetin M., Heiner C., Su Q., Weiand M., Wilmot D., Zapp M.L., Gao G. Adeno-associated virus genome population sequencing achieves full vector genome resolution and reveals human-vector chimeras. Mol. Ther. Methods Clin. Dev. 2018;9:130–141. doi: 10.1016/j.omtm.2018.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gray J.T. 2017. Nucleic Acid Molecules Containing Spacers and Methods of Use Thereof. [Google Scholar]
- 30.Karbowniczek K., Rothwell P., Extance J., Milsom S., Lukashchuk V., Bowes K., Smith D., Caproni L. DoggyboneTM DNA: an advanced platform for AAV production. Cell Gene Ther. Insights. 2017;3:731–738. doi: 10.18609/CGTI.2017.074. [DOI] [Google Scholar]
- 31.Kreiss P., Cameron B., Rangara R., Mailhe P., Aguerre-Charriol O., Airiau M., Scherman D., Crouzet J., Pitard B. Plasmid DNA size does not affect the physicochemical properties of lipoplexes but modulates gene transfer efficiency. Nucleic Acids Res. 1999;27:3792–3798. doi: 10.1093/nar/27.19.3792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mignon C., Sodoyer R., Werle B. Antibiotic-Free selection in biotherapeutics: now and forever. Pathogens. 2015;4:157–181. doi: 10.3390/PATHOGENS4020157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Li C., Goudy K., Hirsch M., Asokan A., Fan Y., Alexander J., Sun J., Monahan P., Seiber D., Sidney J., et al. Cellular immune response to cryptic epitopes during therapeutic gene transfer. Proc. Natl. Acad. Sci. USA. 2009;106:10770–10774. doi: 10.1073/PNAS.0902269106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Monjezi R., Miskey C., Gogishvili T., Schleef M., Schmeer M., Einsele H., Ivics Z., Hudecek M. Enhanced CAR T-cell engineering using non-viral Sleeping Beauty transposition from minicircle vectors. Leukemia. 2017;31:186–194. doi: 10.1038/LEU.2016.180. [DOI] [PubMed] [Google Scholar]
- 35.Danner E., Lebedin M., de La Rosa K., Kühn R. A homology independent sequence replacement strategy in human cells using a CRISPR nuclease. Open Biol. 2021;11:200283. doi: 10.1098/RSOB.200283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Florian M., Wang J.P., Deng Y., Souza-Moreira L., Stewart D.J., Mei S.H.J. Gene engineered mesenchymal stem cells: greater transgene expression and efficacy with minicircle vs. plasmid DNA vectors in a mouse model of acute lung injury. Stem Cell Res. Ther. 2021;12:184. doi: 10.1186/S13287-021-02245-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lu J., Zhang F., Xu S., Fire A.Z., Kay M.A. The extragenic spacer length between the 5’ and 3’ ends of the transgene expression cassette affects transgene silencing from plasmid-based vectors. Mol. Ther. 2012;20:2111–2119. doi: 10.1038/MT.2012.65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Marie C., Vandermeulen G., Quiviger M., Richard M., Préat V., Scherman D. pFARs, plasmids free of antibiotic resistance markers, display high-level transgene expression in muscle, skin and tumour cells. J. Gene Med. 2010;12:323–332. doi: 10.1002/jgm.1441. [DOI] [PubMed] [Google Scholar]
- 39.Luke J., Carnes A.E., Hodgson C.P., Williams J.A. Improved antibiotic-free DNA vaccine vectors utilizing a novel RNA based plasmid selection system. Vaccine. 2009;27:6454–6459. doi: 10.1016/J.VACCINE.2009.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Williams J.A. Improving DNA vaccine performance through vector design. Curr. Gene Ther. 2014;14:170–189. doi: 10.2174/156652321403140819122538. [DOI] [PubMed] [Google Scholar]
- 41.Carnes A.E., Luke J.M., Vincent J.M., Anderson S., Schukar A., Hodgson C.P., Williams J.A. Critical design criteria for minimal antibiotic-free plasmid vectors necessary to combine robust RNA Pol II and Pol III-mediated eukaryotic expression with high bacterial production yields. J. Gene Med. 2010;12:818–831. doi: 10.1002/jgm.1499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hoogewoud F., Buggage R., Behar-Cohen F. EYS606 for the Treatment of Non-Infectious Uveitis. Acta Ophthalmol. 2019;97 doi: 10.1111/j.1755-3768.2019.5326. [DOI] [Google Scholar]
- 43.Teixeira L., Medioni J., Garibal J., Adotevi O., Doucet L., Durey M.A.D., Ghrieb Z., Kiladjian J.J., Brizard M., Laheurte C., et al. A first-in-human phase I study of INVAC-1, an optimized human telomerase DNA vaccine in patients with advanced solid tumors. Clin. Cancer Res. 2020;26:588–597. doi: 10.1158/1078-0432.CCR-19-1614. [DOI] [PubMed] [Google Scholar]
- 44.Su Y., Romeu-Bonilla E., Anagnostou A., Fitz-Patrick D., Hearl W., Heiland T. Safety and long-term immunological effects of CryJ2-LAMP plasmid vaccine in Japanese red cedar atopic subjects: A phase I study. Hum. Vaccin. Immunother. 2017;13:2804–2813. doi: 10.1080/21645515.2017.1329070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Dutton J.L., Woo W.P., Chandra J., Xu Y., Li B., Finlayson N., Griffin P., Frazer I.H. An escalating dose study to assess the safety, tolerability and immunogenicity of a Herpes Simplex Virus DNA vaccine, COR-1. Hum. Vaccin. Immunother. 2016;12:3079–3088. doi: 10.1080/21645515.2016.1221872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Chandra J., Woo W.P., Dutton J.L., Xu Y., Li B., Kinrade S., Druce J., Finlayson N., Griffin P., Laing K.J., et al. Immune responses to a HSV-2 polynucleotide immunotherapy COR-1 in HSV-2 positive subjects: A randomized double blinded phase I/IIa trial. PLoS One. 2019;14:e0226320. doi: 10.1371/JOURNAL.PONE.0226320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Suzuki M., Kasai K., Saeki Y. Plasmid DNA sequences present in conventional herpes simplex virus amplicon vectors cause rapid transgene silencing by forming inactive chromatin. J. Virol. 2006;80:3293–3300. doi: 10.1128/JVI.80.7.3293-3300.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Darquet A.M., Cameron B., Wils P., Scherman D., Crouzet J. A new DNA vehicle for nonviral gene delivery: supercoiled minicircle. Gene Ther. 1997;4:1341–1349. doi: 10.1038/SJ.GT.3300540. [DOI] [PubMed] [Google Scholar]
- 49.Chen Z.Y., He C.Y., Ehrhardt A., Kay M.A. Minicircle DNA vectors devoid of bacterial DNA result in persistent and high-level transgene expression in vivo. Mol. Ther. 2003;8:495–500. doi: 10.1016/S1525-0016(03)00168-0. [DOI] [PubMed] [Google Scholar]
- 50.Arévalo-Soliz L.M., Hardee C.L., Fogg J.M., Corman N.R., Noorbakhsh C., Zechiedrich L. Improving therapeutic potential of non-viral minimized DNA vectors. Cell Gene Ther. Insights. 2020;6:1489–1505. doi: 10.18609/CGTI.2020.163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kay M.A., He C.Y., Chen Z.Y. A robust system for production of minicircle DNA vectors. Nat. Biotechnol. 2010;28:1287–1289. doi: 10.1038/NBT.1708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Williams J. 2019. Viral and Non-Viral Nanoplasmid Vectors with Improved Production. [Google Scholar]
- 53.Lu J., Zhang F., Kay M.A. A mini-intronic plasmid (MIP): a novel robust transgene expression vector in vivo and in vitro. Mol. Ther. 2013;21:954–963. doi: 10.1038/MT.2013.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Lu J., Williams J.A., Luke J., Zhang F., Chu K., Kay M.A. A 5’ noncoding exon containing engineered intron enhances transgene expression from recombinant AAV vectors in vivo. Hum. Gene Ther. 2017;28:125–134. doi: 10.1089/HUM.2016.140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Riu E., Chen Z.Y., Xu H., He C.Y., Kay M.A. Histone modifications are associated with the persistence or silencing of vector-mediated transgene expression in vivo. Mol. Ther. 2007;15:1348–1355. doi: 10.1038/SJ.MT.6300177. [DOI] [PubMed] [Google Scholar]
- 56.Carnes A.E., Hodgson C.P., Williams J.A. Inducible Escherichia coli fermentation for increased plasmid DNA production. Biotechnol. Appl. Biochem. 2006;45:155–166. doi: 10.1042/BA20050223. [DOI] [PubMed] [Google Scholar]
- 57.Abremski K., Frommer B., Hoess R.H. Linking-number changes in the DNA substrate during Cre-mediated loxP site-specific recombination. J. Mol. Biol. 1986;192:17–26. doi: 10.1016/0022-2836(86)90460-2. [DOI] [PubMed] [Google Scholar]
- 58.Nash H.A., Pollock T.J. Site-specific recombination of bacteriophage lambda. The change in topological linking number associated with exchange of DNA strands. J. Mol. Biol. 1983;170:19–38. doi: 10.1016/S0022-2836(83)80225-3. [DOI] [PubMed] [Google Scholar]
- 59.Zechiedrich E.L., Khodursky A.B., Cozzarelli N.R. Topoisomerase IV, not gyrase, decatenates products of site-specific recombination in Escherichia coli. Genes Dev. 1997;11:2580–2592. doi: 10.1101/GAD.11.19.2580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Weigl D., Molloy M.J., Clayton T.M., Griffith J., Smith C.R., Steward T., Merrill B.M., DePrince R.B., Jone C.S., Persmark M. Characterization of a topologically aberrant plasmid population from pilot-scale production of clinical-grade DNA. J. Biotechnol. 2006;121:1–12. doi: 10.1016/J.JBIOTEC.2005.06.019. [DOI] [PubMed] [Google Scholar]
- 61.Boye C., Arpag S., Francis M., DeClemente S., West A., Heller R., Bulysheva A. Reduction of plasmid vector backbone length enhances reporter gene expression. Bioelectrochemistry. 2022;144:107981. doi: 10.1016/J.BIOELECHEM.2021.107981. [DOI] [PubMed] [Google Scholar]
- 62.Boye C., Arpag S., Burcus N., Lundberg C., DeClemente S., Heller R., Francis M., Bulysheva A. Cardioporation enhances myocardial gene expression in rat heart. Bioelectrochemistry. 2021;142:107892. doi: 10.1016/J.BIOELECHEM.2021.107892. [DOI] [PubMed] [Google Scholar]
- 63.Mitdank H., Tröger M., Sonntag A., Shirazi N.A., Woith E., Fuchs H., Kobelt D., Walther W., Weng A. Suicide nanoplasmids coding for ribosome-inactivating proteins. Eur. J. Pharm. Sci. 2022;170:106107. doi: 10.1016/J.EJPS.2021.106107. [DOI] [PubMed] [Google Scholar]
- 64.Chan T., Grisch-Chan H.M., Schmierer P., Subotic U., Rimann N., Scherer T., Hetzel U., Bozza M., Harbottle R., Williams J.A., et al. Delivery of non-viral naked DNA vectors to liver in small weaned pigs by hydrodynamic retrograde intrabiliary injection. Mol. Ther. Methods Clin. Dev. 2022;24:268–279. doi: 10.1016/j.omtm.2022.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Merting A.D., Poschel D.B., Lu C., Klement J.D., Yang D., Li H., Shi H., Chapdelaine E., Montgomery M., Redman M.T., et al. Restoring FAS expression via lipid-encapsulated FAS DNA nanoparticle delivery is sufficient to suppress colon tumor growth in vivo. Cancers. 2022;14:361. doi: 10.3390/CANCERS14020361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Vermeire G., de Smidt E., Geukens N., Williams J.A., Declerck P., Hollevoet K. Improved potency and safety of DNA-encoded antibody therapeutics through plasmid backbone and expression cassette engineering. Hum. Gene Ther. 2021;32:1200–1209. doi: 10.1089/HUM.2021.105. [DOI] [PubMed] [Google Scholar]
- 67.Luke J.M., Simon G.G., Söderholm J., Errett J.S., August J.T., Gale M., Hodgson C.P., Williams J.A. Coexpressed RIG-I agonist enhances humoral immune response to influenza virus DNA vaccine. J. Virol. 2011;85:1370–1383. doi: 10.1128/JVI.01250-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Dauphinee S., Goulet M.-L., Veilleux D., Louis K., Lazure D., Stevenson S., Bilimoria D., Dupaul-Chicoine J., Sublemontier S., Amirkhani S., et al. EG-70, a novel non-viral gene therapy for local expression of innate and adaptive immune modulators for treatment of non-muscle invasive bladder cancer. Mol. Ther. 2022;30:155. [Google Scholar]
- 69.Steinberg G.D., Kalota S.J., Lotan Y., Warner L., Dauphinee S., Mazanet R. Clinical results of a phase 1 study of intravesical EG-70 in patients with BCG-unresponsive NMIBC. J. Clin. Oncol. 2023;41:512. [Google Scholar]
- 70.Suschak J.J., Williams J.A., Schmaljohn C.S. Advancements in DNA vaccine vectors, non-mechanical delivery methods, and molecular adjuvants to increase immunogenicity. Hum. Vaccin. Immunother. 2017;13:2837–2848. doi: 10.1080/21645515.2017.1330236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Raturi A., Ablack J., Wee P., Bhandari P., Brown D.W., Hejazi M., McMullen N., Grin L., Vega H., Garcia H., et al. Immunogenicity of a SARS-CoV-2 DNA vaccine formulated with the fusion-associated small transmembrane protein proteolipid vehicle delivery system. SSRN J. 2022 doi: 10.2139/SSRN.4241174. [DOI] [Google Scholar]
- 72.Borggren M., Nielsen J., Bragstad K., Karlsson I., Krog J.S., Williams J.A., Fomsgaard A. Vector optimization and needle-free intradermal application of a broadly protective polyvalent influenza A DNA vaccine for pigs and humans. Hum. Vaccin. Immunother. 2015;11:1983–1990. doi: 10.1080/21645515.2015.1011987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Suschak J.J., Dupuy L.C., Shoemaker C.J., Six C., Kwilas S.A., Spik K.W., Williams J.A., Schmaljohn C.S. Nanoplasmid vectors Co-expressing innate immune agonists enhance DNA vaccines for Venezuelan Equine Encephalitis virus and Ebola virus. Mol. Ther. Methods Clin. Dev. 2020;17:810–821. doi: 10.1016/J.OMTM.2020.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Chowdhury I.H., Lokugamage N., Garg N.J. Experimental nanovaccine offers protection against repeat exposures to Trypanosoma cruzi through activation of Polyfunctional T cell response. Front. Immunol. 2020;11:3333. doi: 10.3389/FIMMU.2020.595039/BIBTEX. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Clauss J., Obenaus M., Miskey C., Ivics Z., Izsvák Z., Uckert W., Bunse M. Efficient non-viral T-cell engineering by sleeping beauty minicircles diminishing DNA toxicity and miRNAs silencing the endogenous T-cell receptors. Hum. Gene Ther. 2018;29:569–584. doi: 10.1089/HUM.2017.136. [DOI] [PubMed] [Google Scholar]
- 76.Bozza M., Green E.W., Espinet E., de Roia A., Klein C., Vogel V., Offringa R., Williams J.A., Sprick M., Harbottle R.P. Novel non-integrating DNA Nano-S/MAR vectors restore gene function in isogenic patient-derived pancreatic tumor models. Mol. Ther. Methods Clin. Dev. 2020;17:957–968. doi: 10.1016/J.OMTM.2020.04.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Roig-Merino A., Urban M., Bozza M., Peterson J.D., Bullen L., Büchler-Schäff M., Stäble S., van der Hoeven F., Müller-Decker K., McKay T.R., et al. An episomal DNA vector platform for the persistent genetic modification of pluripotent stem cells and their differentiated progeny. Stem Cell Rep. 2022;17:143–158. doi: 10.1016/J.STEMCR.2021.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Bozza M., de Roia A., Correia M.P., Berger A., Tuch A., Schmidt A., Zörnig I., Jäger D., Schmidt P., Harbottle R.P. A nonviral, nonintegrating DNA nanovector platform for the safe, rapid, and persistent manufacture of recombinant T cells. Sci. Adv. 2021;7:eabf1333. doi: 10.1126/SCIADV.ABF1333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Moretti A., Ponzo M., Nicolette C.A., Tcherepanova I.Y., Biondi A., Magnani C.F. The past, present, and future of Non-Viral CAR T cells. Front. Immunol. 2022;13:867013. doi: 10.3389/FIMMU.2022.867013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Ostertag E. POSEIDA THERAPEUTICS; 2020. Manufacturing Matters in CAR-T: Small Changes Can Have a Big Impact.https://poseida.com/wp-content/uploads/2021/01/Manufacturing-Matters-in-CAR-T.pdf [Google Scholar]
- 81.Oh S.A., Senger K., Madireddi S., Akhmetzyanova I., Ishizuka I.E., Tarighat S., Lo J.H., Shaw D., Haley B., Rutz S. High-efficiency nonviral CRISPR/Cas9-mediated gene editing of human T cells using plasmid donor DNA. J. Exp. Med. 2022;219:e20211530. doi: 10.1084/JEM.20211530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Pomeroy E.J., Lahr W.S., Chang J.W., Krueger J., Wick B.J., Slipek N.J., Skeate J.G., Webber B.R., Moriarity B.S. Non-Viral Engineering of CAR-NK and CAR-T cells using the Tc Buster Transposon SystemTM. bioRxiv. 2021 doi: 10.1101/2021.08.02.454772. Preprint at. [DOI] [Google Scholar]
- 83.Gurney M., O’Reilly E., Corcoran S., Brophy S., Hardwicke D., Krawczyk J., Hermanson D., Childs R.W., Szegezdi E., O’Dwyer M.E. Tc buster transposon engineered CLL-1 CAR-NK cells efficiently target acute myeloid leukemia. Blood. 2021;138:1725. doi: 10.1182/BLOOD-2021-147244. [DOI] [Google Scholar]
- 84.Gurney M., O’Reilly E., Corcoran S., Brophy S., Krawczyk J., Otto N.M., Hermanson D.L., Childs R.W., Szegezdi E., O’Dwyer M.E. Concurrent transposon engineering and CRISPR/Cas9 genome editing of primary CLL-1 chimeric antigen receptor–natural killer cells. Cytotherapy. 2022;24:1087–1094. doi: 10.1016/J.JCYT.2022.07.008. [DOI] [PubMed] [Google Scholar]