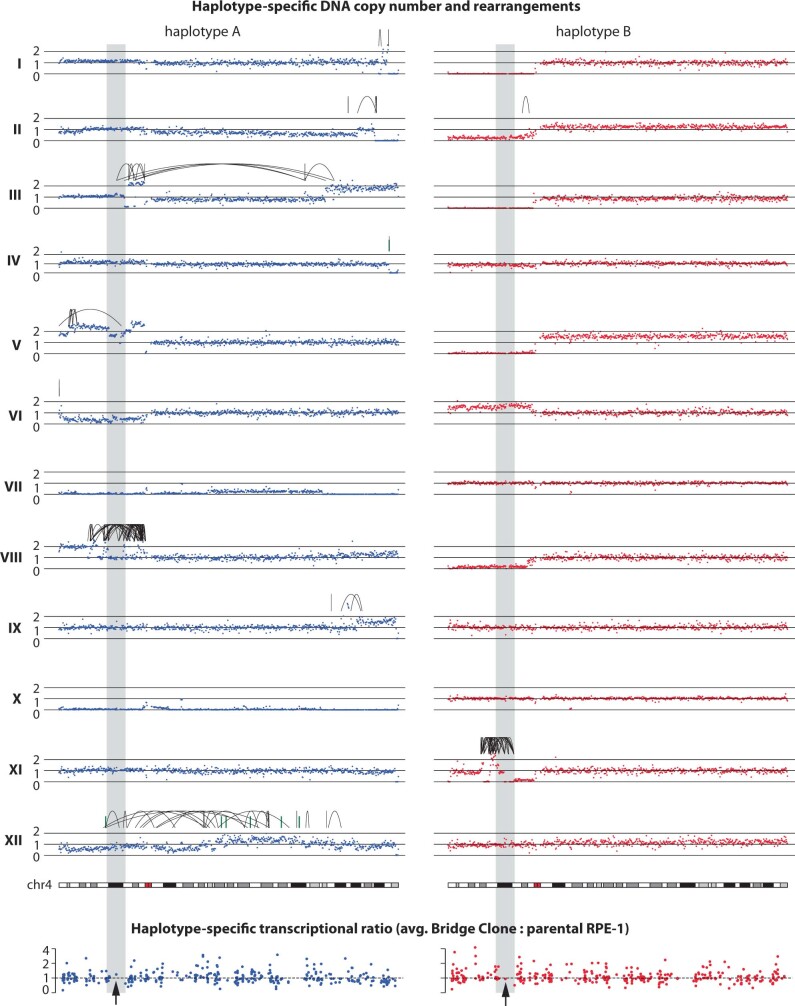
Extended Data Fig. 11. Haplotype-specific DNA copy number, rearrangements, and transcriptional levels of chromosome 4 in 12 bridge clones.
Top: Haplotype-specific Chr.4 DNA copy number (250 kb bins) determined from newly generated DNA-Seq data (~5X mean sequencing depth) of the bridge clones. Black arcs represent intra-chromosomal rearrangements determined from previous whole-genome sequencing of the same bridge clones and subclones (10-60X mean depth)11. Rearrangements are phased to each homologue based on haplotype-specific copy-number changes at rearrangement breakpoints. The shaded box denotes the region of 27-37 Mb on Chr.4p with reduced ATAC-Seq signal as shown in Fig. 5c. We detected no clonal or subclonal breakpoints in this region (based on both DNA copy-number changepoints and rearrangement junctions) in bridge clones I, II, IV, VI, VII, IX, X. Bridge clone VIII and XI contain the most breakpoints within this region but display no significant change of ATAC density after normalization for copy-number variation. In bridge clone III, ATAC reduction is most prominent between 27 and 30.5 Mb and this region is far away from rearrangements that affect two segments between 32.19-32.23 Mb. Bridge clones V and XII both contain multiple copy-number and rearrangement breakpoints in this region and display modest but significant ATAC reduction (based on the permutation test). Bottom: Haplotype-specific gene transcription (TPM) ratio on Chr.4. Each dot represents the average TPM ratio of a single gene calculated from all 12 bridge clones, excluding samples with complete DNA deletion. Arrows point to the PCDH7 gene residing in the region with reduced ATAC signal.