Extended Data Fig. 2. Benchmarking NYUTron against a traditional NLP model and other language models on a different clinical prediction task (clinical concept extraction).
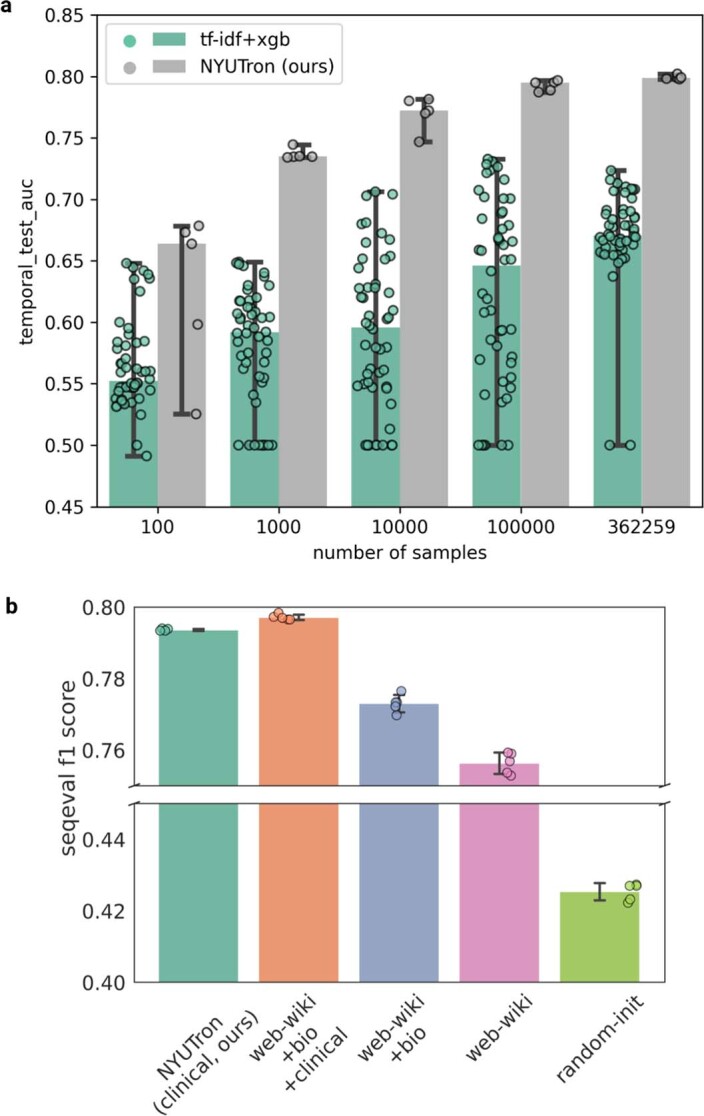
We observe a similar trend as readmission prediction: (a) shows that NYUTron has better performance than tf-idf under different data availability settings and (b) shows that clinically pretrained language models have better performance than non-clinically pretrained language models. This corroborates our findings that health-system scale language models are general purpose clinical prediction engines and that a domain match between pretraining and finetuning corpus contributes to task performance. a, Comparison of temporal test AUCs between NYUTron and a traditional NLP model (tf-idf+xgb). NYUTron has a higher median AUC than tf-idf+xgb for all tested number of finetuning examples. The black vertical line indicates standard deviation over 5 trials of different random seeds (0, 13, 24, 36, 42). b, Comparison of LLMs’ finetuning performances on the NER task. On the i2b2-2012 clinical concept extraction task, the LLMs that are pretrained with clinical corpora (NYUTron, web-wiki+bio+clinical) have a higher average f1 score than LLMs that are not pretrained with clinical corpora (web-wiki+bio, web-wiki, random-init). Specifically, NYUTron and web-wiki+bio+clinical perform better than the randomly initialized model (36.64% higher median seqeval f1 score) and non-clinically pretrained models (2.01%–3.48% higher median seqeval f1 score). Note that the height of each bar is the average f1 score and the half length of each black vertical line indicates the standard deviation over 5 trials of different random seeds (0, 13, 24, 36, 42).
