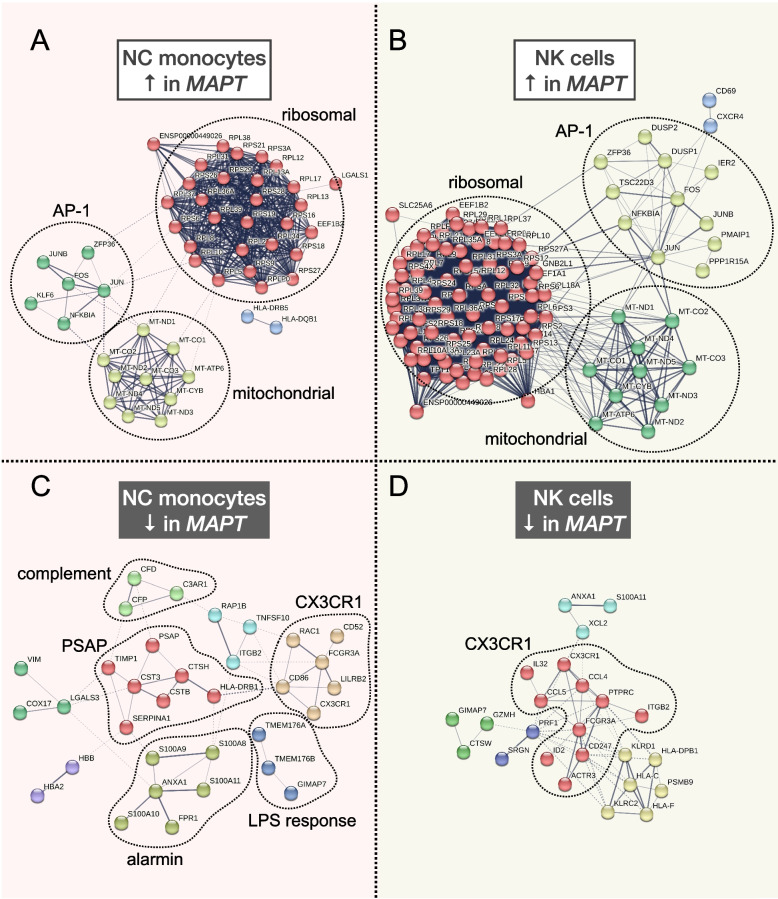
Fig. 3.
STRING interaction networks reveal relationships among nonclassical monocyte and natural killer cell differentially expressed genes. A, B Upregulated DEGs in NC monocytes and NK cells had similar overall network architecture, with large ribosomal and mitochondrial modules, and a third module containing members of the AP-1 transcription factor, among other genes. C The downregulated DEGs in NC monocytes contained a module featuring CX3CR1 and FCGR3A as members, in addition to modules harboring genes involved in LPS response, the alternative and classical complement cascades, the S100 alarmin molecules, as well as PSAP. D Downregulated DEGs in NK cells also featured a large module featuring both CX3CR1 and FCGR3A. All DEGs with pFDR < 0.05 and absolute LFC > 0.1 from clusters 3 and 11 were input into the STRING database as described in the “Methods” section. Modules are colored according to the results of MCL clustering