Figure 5.
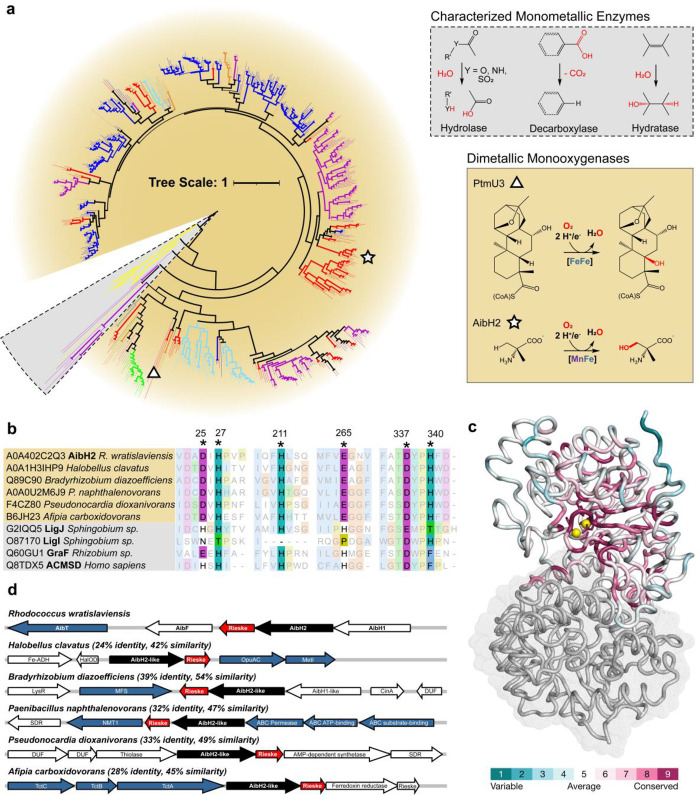
Bioinformatic identification of an uncharacterized family of AibH2-like proteins. (A) Unrooted, neighbor-joining phylogenetic tree of 555 Uniprot Reference Proteome sequences with at least 24% identity to AibH2 and conserved metal-coordinating residues. Biochemically characterized representatives of the PF04909 protein family were included for context. Branch thickness is proportional to the bootstrap values. The branches are colored according to the taxonomic group: red: actinobacteria; magenta: proteobacteria; blue: bacilli; orange: chloroflexi; yellow: eukaryote; teal: haloarchaea; green: cyanobacteria; black: other. The underlying shading reflects the enzymatic reactions (inset) known or expected to be catalyzed by these enzymes. Tree scale, 1.0 amino acid substitution per site. (B) Multiple sequence alignment of representative candidate monooxygenase sequences and characterized monometallic enzymes highlighting the metal binding residues found in AibH2. (C) Ribbon and surface representation of AibH1H2 illustrating the localization of variable (cyan) and conserved (purple) regions of 300 randomly chosen AibH2-like sequences. (D) Genome neighborhood diagrams of AibH2 and AibH2-like proteins. Genes are colored according to an inferred function: black: AibH2-like monooxygenase; red: Rieske-type ferredoxin; blue; small-molecule permease components.