Abstract
Serratia marcescens is now becoming a propensity for its highly antimicrobial-resistant clinical infections. Currently, it provides a novel strategy to prevent and control microbial infection by regulating S. marcescens quorum sensing (QS). Deep-sea-derived fungi are rich in QS bioactive constituents. In this work, the extracts from Cladosporium sphaerospermum SCSGAF0054 showed potent QS-related virulence factors and biofilm-inhibiting activities against S. marcescens NJ01. The swimming motility and multiple virulence factors such as prodigiosin, exopolysaccharide (EPS), lipase, protease and hemolysin were moderately inhibited by the extracts at varied concentrations. The confocal laser scanning microscope (CLSM) and scanning electron microscope (SEM) images revealed that C. sphaerospermum extracts moderately arrested biofilm formation and cell viability. Further, real-time quantitative PCR (RT-qPCR) analysis revealed that expressions of genes associated with virulence factors, flhD, fimA, fimC, bsmA, bsmB, pigA, pigC, and shlA, were significantly down-regulated compared with control. In addition, the extracts combined with imipenem inhibited the QS system of S. marcescens NJ01, disrupted its preformed biofilm, released the intra-biofilm bacteria and killed the bacteria gradually. Therefore, the extracts combined with imipenem can partially restore bacterial drug sensitivity. These results suggest that the extracts from SCSGAF0054 effectively interfere with the QS system to treat S. marcescens infection alone or combining with classical antimicrobial drugs.
Keywords: Cladosporium sphaerospermum, Deep-sea-derived fungus, Biofilm, Serratia marcescens, Quorum sensing
1. Introduction
Serratia marcescens is one clinical pathogen that can cause lung and urinary tract infections and septicemia when the body's immune function decreases [1]. It is also a pathogen of many plants, causing plant diseases such as watermelon by clinging to plant tissue [2]. With the abuse or misuse of antimicrobial drugs, the resistance of S. marcescens to many common antimicrobial drugs is increasing. Carbapenem antimicrobial drugs are commonly used to treat infections of S. marcescens [3]. However, the resistance to carbapenem antimicrobial drugs was more serious, which caused the infection rate of the bacteria to increase gradually. Therefore, new alternative therapies and prevention strategies are urgently needed to update or replace antimicrobial treatments. Quorum sensing (QS) and biofilm inhibition may be one novel alternative strategy to deal with such drug-resistant troubles.
Deep-sea-derived fungi are relatively untapped biological resources for discovering bioactive novel natural products. In the past decades, about 1 000 new natural products have been extracted from deep-sea-derived fungi, including terpenes, alkaloids, mangosteen-derived metabolites, lipids, polyketones and nonribosomal peptides [4]. Some of these compounds have antimicrobial and anticancer activities. Since the first quorum sensing inhibitor (QSI) was isolated from one red algae, some QSIs were developed in marine organisms [5]. QSIs have been found in the deep-sea-derived fungi Sarocladium, Khuskia, Fusarium and so on [6]. Cladosporium from the deep sea is of great interest for its ability to produce many metabolites, including macrolides, alkaloids, tetramic acid derivatives and so on [7]. These metabolites have various biological activities such as antimicrobial, antiviral, insecticidal, etc [8]. Till now, cladodionen isolated from Cladosporium sphaerospermum Z148 has an anti-quorum sensing activity against Pseudomonas aeruginosa [9]. However, no reports of C. sphaerospermum against S. marcescens NJ01 have been found. Hence, the present investigation aims to evaluate the inhibitory effects of C. sphaerospermum SCSGAF0054 extracts against the QS-associated virulence factors and biofilm formation of S. marcescens NJ01.
2. Results
2.1. LC-MS profiling of C. sphaerospermum SCSGAF0054 extracts
The LC-MS analysis of C. sphaerospermum SCSGAF0054 extracts identified 1 057 compounds in the mzCloud database. Among these, 498 compounds exhibited high degree of similarity to the mzCloud database, with a score greater than 60.
Further, cyclo(L-Phe-L-Pro), 3-Phenyllactic acid, pyridoxine, anthranilic acid, thymine and benzoic acid were identified at retention time (Rt) of 12.1 min, Rt 11.5 min, Rt 1.5 min, Rt 10.0 min, Rt 4.7 min and Rt 10.1 min, respectively (Fig. 1).
Fig. 1.
Total ion chromatograms of the extracts by LC-MS.
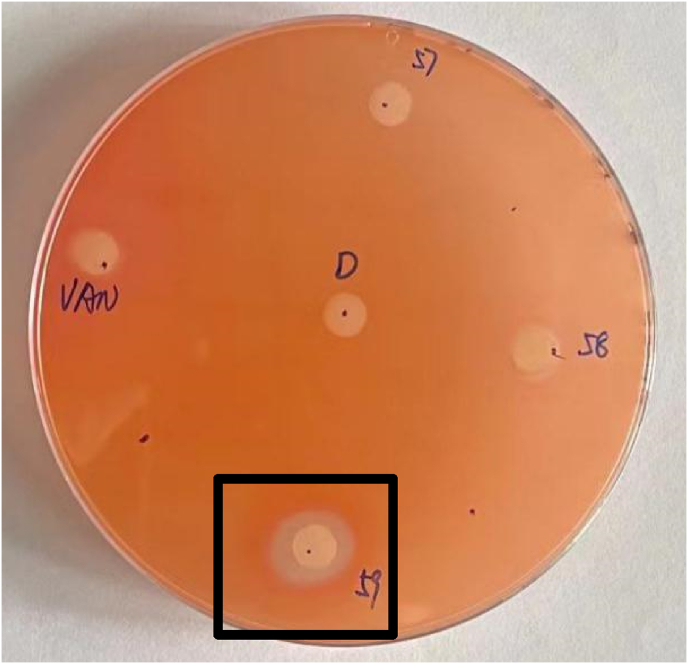
The C. sphaerospermum SCSGAF0054 extract blocks the formation of prodigiosin, which is regulated by quorum sensing (QS) (Fig. 2).
Fig. 2.
Anti-quorum sensing activity of C. sphaerospermum extract against S. marcescens NJ01.
2.2. Determination of minimum inhibitory concentrations (MICs) and growth profiles
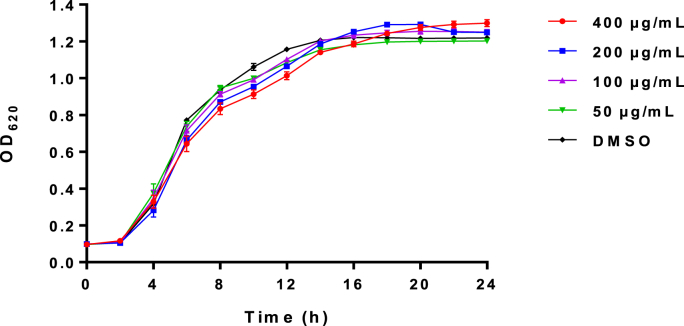
The MIC of the extracts against S. marcescens NJ01 was 800 μg/mL, and the chosen concentration must be lower than the MIC to confirm that it did not inhibit bacterial growth. The result of the bacterial growth profile is represented in Fig. 3. The extracts had no effect on bacterial growth in the range of 50–400 μg/mL, so the concentrations were chosen for further studies. The FICI was 1.5, between 0.5 and 4, indicating that the extracts had no interaction with imipenem.
Fig. 3.
Effects of the extracts against the S. marcescens NJ01 growth at sub-MIC levels. DMSO served as the negative control.
2.3. Inhibition of virulence factors
Fig. 4A,-E demonstrates that 400 μg/mL of the extracts inhibited the synthesis of prodigiosin (57%), protease (46%), lipase (37%), hemolysin (37%) and exopolysaccharides (50%), respectively. Simulatneously, the inhibition rates of positive control vanillic acid (VNA) against prodigiosin, protease, lipase, hemolysin, and exopolysaccharides were 25%, 31%, 30%, 34%, and 14%, respectively, demonstrating that the extracts were more effective than the positive control.
Fig. 4.
Inhibition rates of the extracts on virulence factors of S. marcescens NJ01. (A) Prodigiosin, (B) Protease, (C) Lipase, (D) Hemolysin and (E) Exopolysaccharides. Data shown are mean ± standard deviation (n = 3). One-way ANOVA was used to assess statistical significance. *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
2.4. Inhibition of swimming motility
The swimming ability of S. marcescens NJ01 was reduced after being treated with C. sphaerospermum extracts. The diameter of swimming motility was decreased (1.105 cm) when treated with 400 μg/mL of extracts. The swimming motility inhibition was observed at 53% (Fig. 5).
Fig. 5.
Assessment of swimming motility inhibition of S. marcescens NJ01 treated with C. sphaerospermum extracts. (A) Swimming motility test (B) The measurement of swimming motility diameter (a) DMSO (b) 50 μg/mL extracts (c) 100 μg/mL extracts (d) 200 μg/mL extracts (e) 400 μg/mL extracts. Data shown are mean ± standard deviation (n = 3). One-way ANOVA was used to assess statistical significance. *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
2.5. Inhibition of biofilm formation
The biofilm inhibition ratio for extracts at 400 μg/mL concentration was recorded as 50% (Fig. 6A). Similarly, the cell survival rate decreased by 44% when treated with 400 μg/mL of C. sphaerospermum extracts.
Fig. 6.
Antibiofilm activity of C. sphaerospermum extracts against S. marcescens NJ01. (A) Crystal violet, (B) MTT. Data shown are mean ± standard deviation (n = 3). One-way ANOVA was used to assess statistical significance. *p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001.
Additionally, CLSM and SEM observations have been performed to analyze the changes in the architecture of biofilms treated with the extracts. The CLSM images revealed that C. sphaerospermum extracts reduced and loosened biofilms, but the control image was uniformly dispersed (Fig. 7). Similarly, SEM images revealed that DMSO-exposed biofilms were dense and evenly distributed, whereas biofilms treated with variable concentrations of extracts loosened and disintegrated (Fig. 8).
Fig. 7.
Confocal laser scanning microscopic observation of S. marcescens NJ01 biofilms treated with C. sphaerospermum extracts (a) and (a') DMSO (b) and (b') 400 μg/mL extracts.
Fig. 8.
Scanning electron microscopic observation of S. marcescens NJ01 biofilms treated with C. sphaerospermum extracts. (a) DMSO (b) 50 μg/mL extracts (c) 100 μg/mL extracts (d) 200 μg/mL extracts and (e) 400 μg/mL extracts.
2.6. Damage of preformed biofilm in combination with imipenem
As illustrated in Fig. 9, treatment with imipenem alone had no significant eradication effect on the preformed biofilm, whereas the combination of 0.2 μg/mL of imipenem and 400 μg/mL of the extracts substantially eradicated the preformed biofilm by 45%. In addition, CLSM was used to observe changes in biofilms treated with the combination of extracts and 0.2 μg/mL of imipenem. CLSM analysis revealed that the biofilm treated with imipenem alone was dense and evenly distributed, whereas the biofilms were reduced, flattened, and loosened after exposure with 400 μg/mL of the extracts and 0.2 μg/mL of imipenem (Fig. 10).
Fig. 9.
Dispersion of preformed biofilm by extracts combined with imipenem by crystal violet method. Data shown are mean ± standard deviation (n = 3). T-test was used to assess statistical significance. *p < 0.05, **p < 0.01 and ***p < 0.001.
Fig. 10.
CLSM observation confirms that the synergistic effect of extracts and imipenem disintegrates preformed biofilm architecture. (a) DMSO (b) imipenem (c) 400 μg/mL extract and (d) imipenem with 400 μg/mL extract.
2.7. Expression profile of virulence genes in S. marcescens NJ01
The alteration of gene expression profile associated with virulence factors in S. marcescens NJ01 was analyzed by RT-qPCR. The expression of QS-related virulence genes flhD, fimA and fimC were significantly downregulated by 31%, 47% and 6%, respectively, when treated with 400 μg/mL of C. sphaerospermum extracts. Further, the virulence genes bsmA (15%) and bsmB (27%) exhibited downregulated gene expression. A downregulation in gene expression was also observed for the virulence genes pigA (30%), pigC (33%) and shlA (10%) (Fig. 11).
Fig. 11.
Effects of the C. sphaerospermum extracts at 400 μg/mL on gene expressions of S. marcescens NJ01. Data shown are mean ± standard deviation (n = 3). T-test was used to assess statistical significance. *p < 0.05, **p < 0.01.
3. Materials and methods
3.1. Strains
The deep-sea-derived fungus SCSGAF0054 was obtained from the Third Institute of Oceanography (TIO) of the Ministry of Natural Resources. S. marcescens NJ01 was acquired from Q.H. Gong (Ocean University of China) which was cultivated in Luria-Bertani (LB) broth (Hopebio, Qingdao, China) and incubated at 28 °C, for 17 h at 180 rpm.
3.2. Fermentation and compound extraction from SCSGAF0054
The fungus SCSGAF0054 was cultivated at 28 °C on minimal glycerol (MG) plate for five days, then grown in FL3 liquid medium (5% mannitol, 0.75% yeast extract, 1.25% monosodium glutamate (MSG), 2.5% glucose, 0.75% maltose, 1.25% soya peptone) and incubated aerobically at 28 °C, 130 rpm for seven days. After fermentation, the solution was subjected to ultrasonic extraction using ethyl acetate. The extracts were obtained and stored for further experiments. The bacterial pathogen, S. marcescens NJ01 was cultivated in LB broth and maintained at 28 °C, 180 rpm for 17 h. Initially, 500 μL culture with OD620 = 0.1 was added to 50 mL of luke warm LB agar and mixed well. The mixture was poured into each petri plate and left for solidification. Then 30 μL extract was added to the solidified medium and incubated at 28 °C for 24 h. The anti-QS activity was assessed by observing the pigment inhibition zone.
3.3. LC-MS analysis of the extracts
The compounds were identified from the extracts of SCSGAF0054 by non-targeted LC-MS (Thermo Scientific™ Q Exactive™) analysis in the ion trap scanning mode. The mobile phase were prepared with sterilized distilled water with 0.1% of formic acid (solvent A) and HPLC grade 100% methanol (solvent B) [10]. The elution gradient program was carried out as follows, initially 98% A and 2% B for 1 min; 80% A and 20% B for 5 min; 50% A and 50% B for 10 min; 20% A and 80% B for 15 min; 5% A and 95% B for 20 min; 5% A and 95% B for 27 min; 98% A and 2% B for 28 min; 98% A and 2% B for 30 min. The sample injection volume was fixed at 2 μL. The obtained data were processed and analyzed using the Thermo Scientific Compound Discoverer 3.3 SP1 software. The detection of metabolites is primarily dependent on comparing MS/MS data to the mzCloud database. The unidentified signals were re-entered into the ChemSpider database using MS data and were confirmed by a FISh score greater than 50.
3.4. MICs and growth measurement
The minimum inhibitory concentrations (MICs) of the extracts and imipenem against S. marcescens NJ01 were determined using CLSI method (Clinical and Labrotory Standards Institute, 2021). The pathogen S. marcescens NJ01 was cultivated and calibrated to 0.5 McFarland standard (1 × 108 CFU/mL). Different doses of active extracts (50–800 μg/mL) and were loaded into the each well contained S. marcescens NJ01 and incubated at 28 °C, 180 rpm for 24 h. The absorbance of the bacterial density was measured using 96-well micro plate reader at 620 nm [11].
The bacterial growth curve was measured by a slightly modified method [12]. The different concentrations of active extracts and the same amount of DMSO were added to the 0.5 McFarland calibrated cultures. The bacterial growth was measured at 620 nm every 2 h using a microplate reader.
3.5. Virulence factors assay
S. marcescens NJ01 was cultured in LB broth containing extracts at the desired concentrations and incubated for 24 h at 28 °C. DMSO and vanillic acid were served as the negative and positive controls, respectively. Further, the cultures were centrifuged at 15 294×g for 5 min. The sterile supernatants and pellets were collected and used for the following studies [13].
For the determination of protease activity, 75 μL of sterile supernatants were mixed with 125 μL of 0.3% azocasein in 0.05 M Tris-HCl. Then the mixture was incubated at 37 °C for 3 h. The reaction was terminated by adding 600 μL of 10% trichloroacetic acid. After that, the reaction mixture was centrifuged at 15 294×g at 4 °C and harvested supernatants. The absorbance of the mixture of 0.5 mL supernatants and equal volume sodium hydroxide solution was measured at 420 nm [14].
Prodigiosin production assay was performed based on previous report with minor modifications [15]. Briefly, 1 mL of acidified ethanol (4%, 1 M HCl) was mixed with the pellets. The solution mixture was centrifuged at 15 294×g for 5 min at 4 °C and collected supernatants. The absorbance of the harvested supernatants was determined at OD534.
Lipase activity assay was performed based on previous report with minor modifications [16]. That is, 100 μL of cell-free supernatant was mixed with 900 μL of buffer solution (1 vol 0.3% (w/v) p-p-nitrobenzyl palmitate dissolved in isopropyl alcohol, 9 vol 0.2% (w/v) sodium deoxycholate and 0.1% (w/v) acacia dissolved in 50 mM disodium hydrogen phosphate) and incubated for 60 min. The solution mixture was centrifuged at 15 294×g for 5 min and collected supernatants. The absorbance of the supernatants were measured at OD410 and the amount of lipase was calculated.
The hemolysin production assay was performed based on previous report with minor modifications [17]. Briefly, 100 μL of the sterile supernatant was added with 900 μL of 2% sheep blood and incubated for 1 h. The mixture was centrifuged at 1 057×g for 10 min. The absorbance of the harvested supernatants was measured at 530 nm and the hemolysin production was calculated.
The exopolysaccharides (EPS) production assay was performed based on previous report with minor modifications [18]. That is, 1 mL of 0.4 M sodium hydroxide solution was added to the pellet and dissolved by shaking for 15 min. The supernatant was collected after 15 min of centrifugation at 1 057×g. The insoluble EPS was obtained by filtering the above supernatant. The EPS was precipitated by adding two volumes of absolute ice-cold ethanol to the supernatant and kept at 4 °C for 24 h. The crude EPS precipitate was then recovered by centrifugation at 12 000×g for 20 min at 4 °C and dissolved in distilled water. The EPS was lyophilized and dissolved in 125 μL of 0.4 M sodium hydroxide solution. Then, 500 μL sulphuric acid was added and mixed thoroughly. Subsequently, 100 μL of 5% phenol was added to the solution mixture and incubated at 37 °C for 15 min. Finally, the EPS content was evaluated at 490 nm using microplate reader.
3.6. Swimming assay
The swimming assay was performed based on previous report with minor modifications [19]. The swimming motility was evaluated on LB agar plates (0.3% Agar, 0.5% NaCl, 1% peptone, pH 7.2) modified with the desired concentration of bioactive extracts. S. marcescens NJ01 was spotted on the center of the plate using a sterile swab and incubated at 28 °C for 24 h in an upright position. DMSO served as the control group. In the plates, the swimming motility of the cells was observed and the percentage of motility was estimated.
3.7. Biofilm inhibition assay
The biofilm inhibition assay was performed using a 96-well polystyrene microtitre plate [20]. Briefly, overnight broth culture of S. marcescens NJ01 was added to the freshly prepared LB broth containing the extracts (50–400 μg/mL) and incubated for 24 h at 28 °C. The biofilm was stabilized by adding 200 μL of methanol to each well for 15 min. Then the plate was desiccated at 60 °C. Biofilms were observed by labeling each well with 0.05% crystal violet dye (100 μL) after the plate was incubated at room temperature for 15 min.The biofilms were rinsed with phosphate-buffered saline (PBS) to remove planktonic cells and residual dye. Crystal violet was extracted from biofilms with 95% ethanol, and the absorbance of the dissolved pigment was monitored at 570 nm by employing a 96-well microtiter plate analyzer. The MTT test was employed to determine the number of colonies in biofilms.
3.8. CLSM and SEM analysis
Based on previous report with minor modifications [21]. Briefly, 1% overnight culture was added to LB broth with or without desired concentrations of extracts and incubated at 28 °C for 24 h. The planktonic bacteria on the coverslips were washed with PBS (pH 7.2).
The biofilms were stained with 0.1 mg/mL acridine orange (AO) and kept for 15 min , After that, the biofilms were rinsed thrice with PBS to remove residual dyes. Finally, the biofilms were fixed in coverslips and observed under confocal laser scanning microscopy (CLSM, Leica Microsystem, Germany).
The biofilms were fixed with 2.5% glutaraldehyde for 12 h of incubation. After drying the samples were dehydrated in a gradient of 50%, 70%, 80%, 90%, and 100% ethanol. The dried biofilm was sputtered with a two-sided metal stub of 10 nm gold (Au) thickness. The biofilm architecture and associated bacterial cell damage were observed by scanning electron microscopy (SEM, Thermo Scientific, Verios G4 UC, USA).
3.9. Dispersion of preformed biofilm by C. sphaerospermum extracts combining with imipenem
The dispersion of preformed biofilm by C. sphaerospermum extracts combining with imipenem were tested by the crystal violet method reported previously [20].
The bacterial biofilms were cultivated in LB broth at 28 °C for 24 h. Then the biofilms were washed with PBS (pH 7.2 to remove planktonic cells. Various concentrations of the extracts and imipenem were added to LB broth in a 96-well microplate containing biofilms, respectively. The plates were incubated at 28 °C for 24 h. After that, the biofilms were rinsed with PBS and subsequently 200 μL of methanol was added to each well to stabilize the biofilms for 15 min. Then the plate was dried at 60 °C. Biofilms were observed by staining each well with 0.05% crystal violet dye (100 μL) after the plate was incubated at room temperature for 15 min. The biofilms were rinsed with phosphate-buffered saline (PBS) to remove planktonic cells and residual dye. Crystal violet was eluted from biofilms with 95% ethanol, and the absorbance of the dissolved dye was evaluated at 570 nm by using a 96-well microtiter plate reader.
The biofilms were stained with AO (0.1 mg/mL) for 15 min. The residual stains were washed with PBS (pH 7.2). The coverslips were attached to the cavity slide and observed under CLSM (Leica Microsystem, Germany).
3.10. Expression profile of virulence genes by RT-qPCR analysis
The gene expression was performed by RT-PCR method with slight modifications [17]. The overnight culture was inoculated in LB broth containing 400 μg/mL of extracts and the same amount of DMSO and incubated for 24 h at 28 °C at 180 rpm. Subsequently, the cultured solution was centrifuged at 10 621×g for 10 min. Further, the total RNA was isolated from the bacterial pellets using Spin Column Bacteria Total RNA Purification kit (Solarbio Technology, Beijing, China). The cDNA synthesis was performed using Reverse Transcription Kit (with dsDNase) (LABGIC Technology, Beijing, China) synthesized cDNA for RT-qPCR detection. Supreme Enzyme Mix (LABGIC Technology, Beijing, China), including Reverse Transcriptase, Ribonuclease Inhibitor and dsDNase, was used to remove genomic DNA and reverse transcription. RT-qPCR was carried out using SYBR Green qPCR Mix Kit (LABGIC Technology, Beijing, China) on a fluorescence quantitative PCR instrument (Analytik-qTOEWER3). The housekeeping gene rpsL was used as a reference gene. The primer details of genes are shown in Table S1.
3.11. Statistical analysis
All experiments were performed in triplicate. Statistical differences were evaluated by one-way analysis of variance (ANOVA) or t-test using GraphPad Prism 6 (GraphPad Software Inc., USA). Statistical significance was expressed using the following terminologies: *p < 0.05, **p < 0.01, ***p < 0.001.
4. Discussion
Serratia marcescens NJ01 has an innate resistance to various antimicrobial agents, such as antibiotics and antimicrobial peptides [3]. Clinical isolates of S. marcescens are frequently identified as multidrug-resistant [22]. The suppression of quorum sensing is an alternative to hindering bacterial growth to mitigate bacterial toxicity and treating infections [23]. Quorum-sensing inhibitors (QSIs) have been demonstrated to reduce virulence and increase antibiotic susceptibility. Microorganisms originating from the ocean's depths are more likely to produce new bioactive molecules with unique mechanisms of action for drug discovery [24]. C. sphaerospermum from the deep sea has produced an abundance of bioactive molecules such as macrolides, alkaloids, tetramic acid derivatives etc [8,25]. The secondary metabolites of the deep-sea fungus C. sphaerospermum EIODSF 008 exhibited considerable antibacterial effects in preliminary studies [26]. Cladodionen was isolated from C. sphaerospermum Z148, showing QSI activity against P. aeruginosa [9]. The marine-derived fungus Cladosporium sp. KFD33 was isolated from a blood cockle in China's Haikou Bay. Altertoxins VIII–XII and cladosporol I were extracted from this fungus fermentation liquid and have been used as a QSI against C. Violaceum CV026 [27]. However, no reports of C. sphaerospermum against S. marcescens NJ01 have been found. Hence, the present investigation aims to evaluate the inhibitory effects of C. sphaerospermum extracts SCSGAF0054 against the QS-associated virulence factors and biofilm formation of S. marcescens NJ01.
The SCSGAF0054 extracts contained 1 057 compounds that were identified in the mzCloud database. Among these, a total of 498 compounds exhibited a strong correlation with the mzCloud database, with a score greater than 60. In LC-MS spectra, the compound with a Rt of 4.7 min was identified as thymine which has been reported to increase the sensitivity of numerous gram-negative pathogens to multiple antibiotics by activating bacterial metabolism and respiration, triggering the generation of reactive oxygen species (ROS) [28]. The compound with a Rt of 1.5 min was pyridoxal, a vitamin B6 that served an essential role in maintaining epidermal function. Further, the compound with a Rt of 12.1 min was cyclo(L-Phe-L-Pro). Previous reports suggested that the Cyclo(L-Phe-L-Pro) (cyclic dipeptide) isolated from the marine fungus C. sphaerospermum F14. It has been reported to have numerous biological activities, including anti-inflammatory, antitumor, and antibacterial properties against L. hongkongensis, M. luteus and Ruegeria sp [29,30]. The compound with a Rt of 11.5 min was 3-phenyllactic acid which exhibited QSI and anti-biofilm activity against P. aeruginosa [21,31]. Anthranilic acid with a Rt of 10.0 min, could inhibit hypha formation in S. scitamineum [32,33]. Benzoic acid with a Rt of 10.1 min, which showed QSI and anti-biofilm activity against E. coli K88 [34]. Some of these compounds have been reported to possesses QSI or anti-biofilm activity. The extract of SCSGAF0054 exhibited significant QSI and anti-biofilm activity against S. marcescens NJ01. However, these compounds are not the main ingredients of the extracts. Therefore, we hypothesized that all the compounds possessed QSI and anti-biofilm activity against S. marcescens NJ01. In addition, C. sphaerospermum contains multiple secondary metabolites and hence it was hypothesized that the extracts could contain novel anti-quorum sensing and anti-biofilm compounds.
Numerous virulence factors of S. marcescens, including biofilm formation, motility, prodigiosin, protease, lipase and hemolysin, are associated with QS [11,35]. Protease in S. marcescens can degrade immunoglobulins A and G in the host, promoting bacterial invasion and inhibiting host defense [36,37]. The cytotoxicity of hemolysin damages host cells through inflammation and hinders neutrophils defensive function [38]. The chosen concentrations of C. sphaerospermum extracts were selected below the MIC value against S. marcescens. S. marcescens NJ01 treated with 400 μg/mL SCSGAF0054 extracts showed reductions in the synthesis of prodigiosin (57%), protease (46%), lipase (37%), hemolysin (37%), and EPS (50%). The motility of S. marcescens NJ01 was reduced when treated with various concentrations of extracts. Swimming mediated by flagella was decreased by 53% at 400 μg/mL. In a previous study, 3-phenylpropan-1-amine (3-PPA) was found to inhibit S. marcescens NJ01 biofilm formation and virulence factors, such as prodigiosin, protease, lipase, hemolysin, and swimming motility [13].
S. marcescens NJ01 forms biofilms through a series of defined stages that culminate in a highly porous, filamentous biofilm composed of cell chains, filaments, and cell clusters dependent on the quorum-sensing systems [13,39,40]. Bacteria susceptible to antimicrobial agents can become resistant after forming a biofilm. According to reports, biofilm is a major contributor to bacterial resistance and chronic infection [16,28]. S. marcescens NJ01 was treated with 400 μg/mL extracts of SCSGAF0054, which resulted in a reduction of biofilm formation (50%) and bacterial cell viability (44%). The results of SEM and CLSM indicated that biofilms were diminished, loosened and weakened after treatment with different concentrations of extracts, compared with control biofilms.
S. marcescens possesses innate resistance to numerous antibiotics, including carbapenems. In healthcare contexts, carbapenem-resistant S. marcescens (CRSM) is an emerging problem, and its enzyme can hydrolyze antibacterial drugs, including imipenem [3,41]. Synergistic effects of QS inhibitors and conventional antimicrobial drugs is a strategy to restore bacterial sensitivity [15,42]. In this study, SCSGAF0054 extracts increased the susceptibility of S. marcescens NJ01 to imipenem and disrupted membrane structures.
Genes associated with virulence factors were selected to investigate the potential molecular mechanism of the extracts inhibiting virulence factors of S. marcescens NJ01. Bacterial fimbriae are crucial for colonization to human epithelial cells and biofilm formation [43]. It was demonstrated that type 1 fimbria of S. marcescens is encoded by fimABCD genes in the early stages of biofilm formation [44,45]. The extracts of SCSGAF0054 inhibited the expression of fimA and fimC by 47% and 6%, respectively. The flhD gene was inhibited by 31%, which is responsible for bacterial motility. The biofilm formation genes including bsmA and bsmB were arrested by 15% and 27%, respectively. The extracts suppressed the prodigiosin-producing pigA and pigC genes expression by 30% and 33%, respectively. Moreover, the extracts inhibited the QS-associated hemolysin-encoding shlA (10%) gene expression.
5. Conclusion
This is the first study using the deep sea fungus, C. sphaerospermum extracts against S. marcescens pathogen. A total of 1 057 bioactive compounds were identified in the C. sphaerospermum extracts by LC-MS analysis. Among them, 498 compounds strongly correlated with the mzCloud database, with a score greater than 60. The biofilm formation and virulence factors of the S. marcescens NJ01, including prodigiosin, protease, lipase, hemolysin, EPS, and swimming motility, were considerably inhibited by various concentrations of C. sphaerospermum extracts. The SEM and CLSM images revealed that C. sphaerospermum extracts moderately arrested biofilm formation and cell viability. The combination of C. sphaerospermum extract with imipenam moderately disrupted preformed biofilm. RT-qPCR analysis revealed that the C. sphaerospermum extract moderately hindered the QS-associated virulence genes. These findings suggest that SCSGAF0054 extracts effectively inhibit the QS system to treat S. marcescens infections alone or in combination with conventional antimicrobial agents. Therefore, developing QSI derived from C. sphaerospermum extract to treat infections caused by S. marcescens and reduce its resistance is strongly encouraged.
CRediT authorship contribution statement
Dan-Rui Liu: Writing – original draft, Visualization, Formal analysis, Methodology, Conceptualization, Data curation, Validation. Qing-Xiang Yan: Resources, Validation, Investigation. Zheng-Biao Zou: Visualization. Chun-Lan Xie: Validation. Xian-Wen Yang: Methodology, Writing – review & editing, Funding acquisition, Resources. Ai-Qun Jia: Methodology, Writing – review & editing, Funding acquisition, Resources.
Declaration of competing interest
The authors declare have no competing financial interests.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (82160664), and the Natural Science Foundation of Hainan Province (221CXTD434, 221QN170).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.bioflm.2023.100146.
Contributor Information
Dan-Rui Liu, Email: liudanrui970503@163.com.
Qing-Xiang Yan, Email: youngqx@126.com.
Zheng-Biao Zou, Email: zhengbiaozou@njust.edu.cn.
Chun-Lan Xie, Email: xiechunlanxx@xmu.edu.cn.
Xian-Wen Yang, Email: yangxianwen@tio.org.cn.
Ai-Qun Jia, Email: ajia@hainanu.edu.cn.
Appendix A. Supplementary data
The following is the Supplementary data to this article.
Data availability
Data will be made available on request.
References
- 1.Bakkiyaraj D., Sivasankar C., Pandian S.K. Inhibition of quorum sensing regulated biofilm formation in Serratia marcescens causing nosocomial infections. Bioorg Med Chem Lett. 2012;22(9):3089–3094. doi: 10.1016/j.bmcl.2012.03.063. [DOI] [PubMed] [Google Scholar]
- 2.Liang Z., Liao W., Yu Y., et al. How does vegetable waste decomposition influence the antibiotic resistome and the human bacterial pathogen structure in leachates? ACS ES&T Water. 2021;2(1):226–236. [Google Scholar]
- 3.Tavares-Carreon F., De Anda-Mora K., Rojas-Barrera I.C., et al. Serratia marcescens antibiotic resistance mechanisms of an opportunistic pathogen: a literature review. PeerJ. 2023;11 doi: 10.7717/peerj.14399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gladfelter A.S., James T.Y., Amend A.S. Marine fungi. Curr Biol. 2019;29(6):R191–R195. doi: 10.1016/j.cub.2019.02.009. [DOI] [PubMed] [Google Scholar]
- 5.Manefield M., de Nys R., Naresh K., et al. Evidence that halogenated furanones from Delisea pulchra inhibit acylated homoserine lactone (AHL)-mediated gene expression by displacing the AHL signal from its receptor protein. Microbiology. 1999;145(2):283–291. doi: 10.1099/13500872-145-2-283. [DOI] [PubMed] [Google Scholar]
- 6.Chen J., Wang B., Lu Y., et al. Quorum sensing inhibitors from marine microorganisms and their synthetic derivatives. Mar Drugs. 2019;17(2):80. doi: 10.3390/md17020080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Choub V., Ajuna H.B., Won S.J., et al. Antifungal activity of Bacillus velezensis CE 100 against anthracnose disease (Colletotrichum gloeosporioides) and growth promotion of walnut (Juglans regia L.) trees. Int J Mol Sci. 2021;22(19) doi: 10.3390/ijms221910438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mohamed G.A., Ibrahim S.R.M. Untapped potential of marine-associated Cladosporium species: an overview on secondary metabolites, biotechnological relevance, and biological activities. Mar Drugs. 2021;19(11):645. doi: 10.3390/md19110645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang M., Zhao L., Wu H., et al. Cladodionen is a potential quorum sensing inhibitor against Pseudomonas aeruginosa. Mar Drugs. 2020;18(4):205. doi: 10.3390/md18040205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nybo S.E., Lamberts J.T. Integrated use of LC/MS/MS and LC/Q-TOF/MS targeted metabolomics with automated label-free microscopy for quantification of purine metabolites in cultured mammalian cells. Purinergic Signal. 2019;15(1):17–25. doi: 10.1007/s11302-018-9643-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sethupathy S., Ananthi S., Selvaraj A., et al. Vanillic acid from Actinidia deliciosa impedes virulence in Serratia marcescens by affecting S-layer, flagellin and fatty acid biosynthesis proteins. Sci Rep. 2017;7(1) doi: 10.1038/s41598-017-16507-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhou J.W., Ruan L.Y., Chen H.J., et al. Inhibition of quorum sensing and virulence in Serratia marcescens by hordenine. J Agric Food Chem. 2019;67(3):784–795. doi: 10.1021/acs.jafc.8b05922. [DOI] [PubMed] [Google Scholar]
- 13.Yin L.J., Zhang P.P., Wang W., et al. 3-Phenylpropan-1-amine enhanced susceptibility of Serratia marcescens to ofloxacin by occluding quorum sensing. Microbiol Spectr. 2022;10(5) doi: 10.1128/spectrum.01829-22. 22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Shanks R.M.Q., Stella N.A., Kalivoda E.J., et al. A Serratia marcescens OxyR homolog mediates surface attachment and biofilm formation. J Bacteriol. 2007;189(20):7262–7272. doi: 10.1128/JB.00859-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Song Y., Sun M., Ma F., et al. Lactiplantibacillus plantarum DLPT4 protects against cyclophosphamide-induced immunosuppression in mice by regulating immune response and intestinal flora. Probiot. Antimicrob. Proteins. 2023:1–13. doi: 10.1007/s12602-022-10015-9. [DOI] [PubMed] [Google Scholar]
- 16.Devi K.R., Srinivasan S., Ravi A.V. Inhibition of quorum sensing-mediated virulence in Serratia marcescens by Bacillus subtilis R-18. Microb Pathog. 2018;120:166–175. doi: 10.1016/j.micpath.2018.04.023. [DOI] [PubMed] [Google Scholar]
- 17.Sun B., Luo H., Jiang H., et al. Inhibition of quorum sensing and biofilm formation of esculetin on Aeromonas hydrophila. Front Microbiol. 2021;12 doi: 10.3389/fmicb.2021.737626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Salini R., Pandian S.K. Interference of quorum sensing in urinary pathogen Serratia marcescens by Anethum graveolens. Pathogens Dis. 2015;73(6) doi: 10.1093/femspd/ftv038. [DOI] [PubMed] [Google Scholar]
- 19.Mattmann M.E., Blackwell H.E. Small molecules that modulate quorum sensing and control virulence in Pseudomonas aeruginosa. J Org Chem. 2010;75(20):6737–6746. doi: 10.1021/jo101237e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Koh K.S., Lam K.W., Alhede M., et al. Phenotypic diversification and adaptation of Serratia marcescens MG1 biofilm-derived morphotypes. J Bacteriol. 2007;189(1):119–130. doi: 10.1128/JB.00930-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chatterjee M., D'morris S., Paul V., et al. Mechanistic understanding of Phenyllactic acid mediated inhibition of quorum sensing and biofilm development in Pseudomonas aeruginosa. Appl Microbiol Biotechnol. 2017;101:8223–8236. doi: 10.1007/s00253-017-8546-4. [DOI] [PubMed] [Google Scholar]
- 22.Sopovski D.S. Purdue University Graduate School; 2019. Antimicrobial resistance in Serratia marcescens. [Google Scholar]
- 23.Ge Z., Kuang Z., Chen J., et al. Comparative genomics analysis of Bacillus velezensis LOH112 isolated from a nonagenarian provides insights into its biocontrol and probiotic traits. Gene. 2022;835 doi: 10.1016/j.gene.2022.146644. [DOI] [PubMed] [Google Scholar]
- 24.Wang Y.N., Meng L.H., Wang B.G. Progress in research on bioactive secondary metabolites from deep-sea derived microorganisms. Mar Drugs. 2020;18(12):614. doi: 10.3390/md18120614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cho H.S., Park S.Y., Ryu C.M., et al. Interference of quorum sensing and virulence of the rice pathogen Burkholderia glumae by an engineered endophytic bacterium. FEMS (Fed Eur Microbiol Soc) Microbiol Ecol. 2007;60(1):14–23. doi: 10.1111/j.1574-6941.2007.00280.x. [DOI] [PubMed] [Google Scholar]
- 26.Liang X., Huang Z.H., Ma X., et al. Unstable tetramic acid derivatives from the deep-sea-derived fungus Cladosporium sphaerospermum EIODSF 008. Mar Drugs. 2018;16(11):448. doi: 10.3390/md16110448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang F., Zhou L., Kong F., et al. Altertoxins with quorum sensing inhibitory activities from the marine-derived fungus Cladosporium sp. KFD33. Mar Drugs. 2020;18(1):67. doi: 10.3390/md18010067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liu Y., Yang K., Jia Y., et al. Thymine sensitizes gram-negative pathogens to antibiotic killing. Front Microbiol. 2021;12 doi: 10.3389/fmicb.2021.622798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Padmavathi A.R., Abinaya B., Pandian S.K. Phenol, 2,4-bis(1,1-dimethylethyl) of marine bacterial origin inhibits quorum sensing mediated biofilm formation in the uropathogen Serratia marcescens. Biofouling. 2014;30(9):1111–1122. doi: 10.1080/08927014.2014.972386. [DOI] [PubMed] [Google Scholar]
- 30.Qi S.H., Xu Y., Xiong H.R., et al. Antifouling and antibacterial compounds from a marine fungus Cladosporium sp. F14. World J Microbiol Biotechnol. 2009;25:399–406. [Google Scholar]
- 31.Barchanska H., Płonka J., Nowak P., et al. Metabolic profiles and fingerprints for the investigation of the influence of nitisinone on the metabolism of the yeast Saccharomyces cerevisiae. Sci Rep. 2023;13(1):1473. doi: 10.1038/s41598-023-28335-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Song S., Sun X., Guo Q., et al. An anthranilic acid-responsive transcriptional regulator controls the physiology and pathogenicity of Ralstonia solanacearum. PLoS Pathog. 2022;18(5) doi: 10.1371/journal.ppat.1010562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Song S., Yin W., Sun X., et al. Anthranilic acid from Ralstonia solanacearum plays dual roles in intraspecies signalling and inter-kingdom communication. ISME J. 2020;14(9):2248–2260. doi: 10.1038/s41396-020-0682-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bonetti A., Tugnoli B., Rossi B., et al. Nature-identical compounds and organic acids reduce E. coli K88 growth and virulence gene expression in vitro. Toxins. 2020;12(8):468. doi: 10.3390/toxins12080468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Smith E.E., Smith Z.H.G., Goldstein I.J. Protein–carbohydrate interaction. A turbidimetric study of the interaction of concanavalin A with amylopectin and glycogen and some of their enzymic and chemical degradation products. Biochem J. 1968;107(5):715–724. doi: 10.1042/bj1070715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Guo Y., Wang L., Hanson A., et al. Identification of protective amino acid metabolism events in nursery pigs fed thermally oxidized corn oil. Metabolites. 2023;13(1):103. doi: 10.3390/metabo13010103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Liu S., Lin M., Hu X., et al. Improved sensitivity of the anti-microcystin-LR ELISA using phage-displayed alpha-type anti-idiotypic nanobody. Anal Biochem. 2023;664 doi: 10.1016/j.ab.2022.115030. [DOI] [PubMed] [Google Scholar]
- 38.Sen S., Chen S., Feng B., et al. Preventive effects of north American ginseng (Panax quinquefolium) on diabetic nephropathy. Phytomedicine. 2012;19(6):494–505. doi: 10.1016/j.phymed.2012.01.001. [DOI] [PubMed] [Google Scholar]
- 39.Mangalampalli B., Dumala N., Grover P. Toxicity assessment of magnesium oxide nano and microparticles on cancer and non-cancer cell lines. Nucleus. 2019;62(3):227–241. [Google Scholar]
- 40.Zhou J.W., Ji P.C., Wang C.Y., et al. Synergistic effect of propyl gallate and antibiotics against biofilms of Serratia marcescens and Erwinia carotovora in vitro. Lebensm Wiss Technol. 2023;173 [Google Scholar]
- 41.Sudharshana T.N., Venkatesh H.N., Nayana B., et al. Anti-microbial and anti-mycotoxigenic activities of endophytic Alternaria alternata isolated from Catharanthus roseus (L.) G. Don.: molecular characterisation and bioactive compound isolation. Mycology. 2019;10(1):40–48. doi: 10.1080/21501203.2018.1541933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Xiang Y., Liang B., Zhang X., et al. Atheroprotective mechanism by which folic acid regulates monocyte subsets and function through DNA methylation. Clin Epigenet. 2022;14(1):1–17. doi: 10.1186/s13148-022-01248-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Liu X., Zou Z., Zhang C., et al. Knockdown of the trehalose-6-phosphate synthase gene using RNA interference inhibits synthesis of trehalose and increases lethality rate in Asian citrus psyllid, Diaphorina citri (Hemiptera: psyllidae) Insects. 2020;11(9):605. doi: 10.3390/insects11090605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Garcez W.S., Martins D., Garcez F.R., et al. Effect of spores of saprophytic fungi on phytoalexin accumulation in seeds of frog-eye leaf spot and stem canker-resistant and-susceptible soybean (Glycine m ax L.) cultivars. J Agric Food Chem. 2000;48(8):3662–3665. doi: 10.1021/jf991146o. [DOI] [PubMed] [Google Scholar]
- 45.Wattana-Amorn P., Charoenwongsa W., Williams C., et al. Antibacterial activity of cyclo(L-Pro-L-Tyr) and cyclo(D-Pro-L-Tyr) from Streptomyces sp. strain 22-4 against phytopathogenic bacteria. Nat Prod Res. 2016;30(17):1980–1983. doi: 10.1080/14786419.2015.1095747. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data will be made available on request.