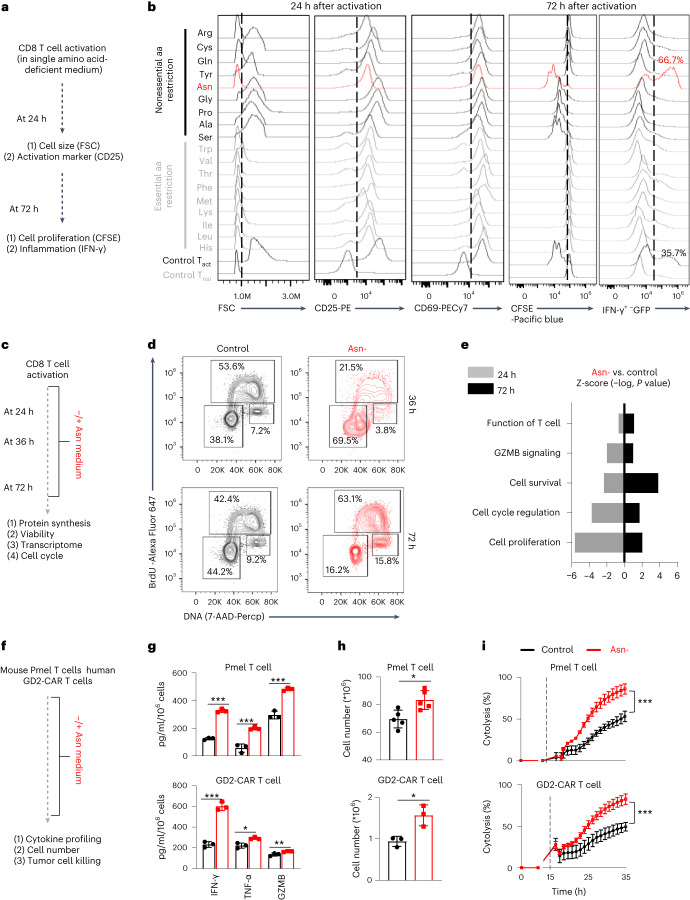
Fig. 1. Asn restriction enhances CD8+ T cell proliferation and effector functions in vitro.
a, Schematic diagram of comprehensively profiled T cell activation, proliferation and IFN-γ production under single amino acid-deficient conditions. FSC, forward scatter. b, CD8+ T cells were activated in a complete medium or indicated single amino acid-deficient medium. Cell size (FSC) and cell surface expression of CD25 and CD69 were determined by flow cytometry in the early phase (24 h). Proliferation (CFSE) and intercellular expression of IFN-γ were determined by flow cytometry in the late phase (72 h). Flow plots are representative of two independent experiments. aa, amino acid; Tnai, Naive CD8+ T cells; Tact, Active CD8+ T cells. c, Schematic diagram of sample collection and assays. d,e, CD8+ T cells were activated in a medium with or without Asn and were collected in the early phase (24–36 h) and late phase (72 h). The cell cycle profile (d) was determined by BrdU incorporation and 7-AAD staining by flow cytometry. The numbers indicate the percentage of cells in the cell cycle stage (n = 3 experimental replicates). Data are representative of three independent experiments. Comparative differently expressed gene signatures (e) were determined by the Ingenuity pathway analysis (IPA) of RNA-seq and listed according to their z-score. Data are representative of one experiment. f, Schematic diagram of in vitro assays. g–i, Indicated proteins in culture supernatants were quantified by ELISA and LEGENDplex (g) (n = 3 experimental replicates; upper panel: P < 0.0001, P = 0.015 and P = 0.0004 for IFN-γ, TNF-α and GZMB, respectively; lower panel: P = 0.002, P = 0.0111 and P = 0.0274 for IFN-γ, TNF-α and GZMB, respectively). The cell number was determined by a cell counter (h) (n = 3 experimental replicates for GD2-CAR T cells and n = 5 experimental replicates for Pmel T cells; P = 0.0108 and P = 0.0176 for the upper and lower panels, respectively). The cytolysis was determined by eSight (i) (n = 3 experimental replicates; P = < 0.0001). Data in g–i are representative of three independent experiments. Error bars are mean ± s.d. *P < 0.05; **P < 0.01; ***P < 0.001. Statistical differences were determined by paired two-tailed Student’s t-test (e), unpaired two-tailed Student’s t-test (g, h) and two-way ANOVA (i).