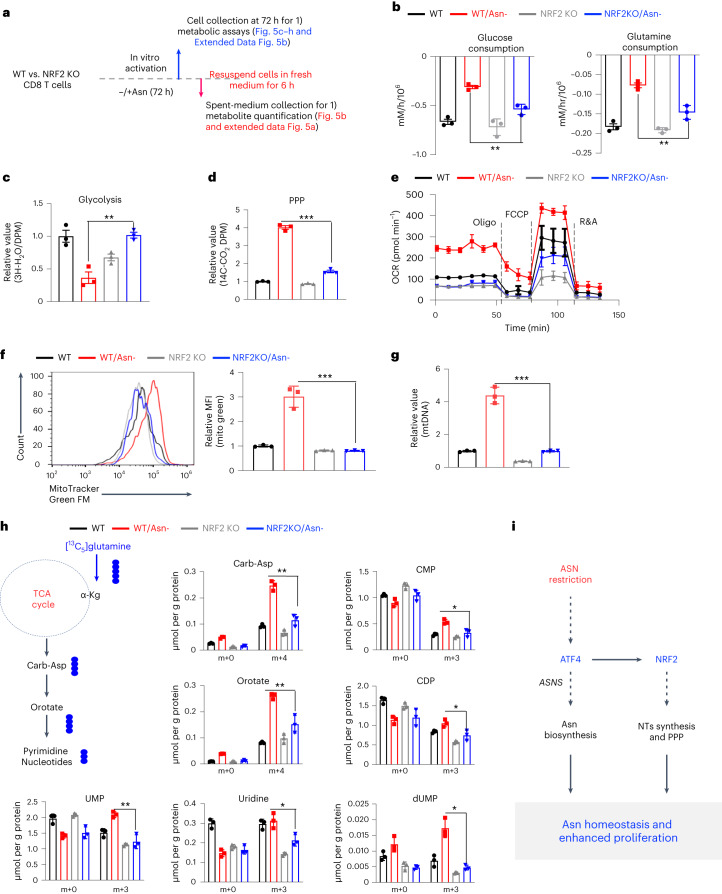
Fig. 5. NRF2 is required to rewire the carbon metabolic program and enhance nucleotide biosynthesis under Asn restriction.
a, Schematic diagram of sample collection and assays. b, The indicated metabolites were quantified by the YSI bioanalyzer. Consumption was determined by calculating the difference between blank and spent medium (6-h incubation of T cells collected in the late phase (72 h)) (n = 3 experimental replicates; P = 0.0027 and P = 0.0043 for glucose and glutamine, respectively). c,d, T cells were collected in the late phase (72 h). Glycolysis activity (c) (n = 3 experimental replicates; P = 0.0027) and PPP activity (d) (n = 3 experimental replicates; P < 0.0001) were determined by measuring 3H2O generated from D-[5-3H(N)]glucose and 14CO2 generated from [1-14C]D-glucose, respectively. e, OCR (in the late phase (72 h)) was determined by Seahorse (n = 3 experimental replicates). f,g, Mitochondrial mass was determined by MitoTracker Green FM staining by flow cytometry (f) (n = 3 experimental replicates; P = 0.0009), and mitochondrial DNA was quantified by qPCR (g) (n = 3 experimental replicates; P = 0.0003). Data in b–g are representative of three independent experiments. MFI, mean fluorescence intensity; mtDNA, mitochondrial DNA. h, Diagram of [13C5]glutamine catabolism through entering the downstream TCA cycle and pyrimidine biosynthesis. Filled circles denote the 13C label of all carbons of indicated metabolites derived from [13C5]glutamine catabolism (left panel). Metabolites in cells (in the late phase (72 h)) were analyzed by IC–UHR-FTMS (right panel); numbers on the x axis represent those of 13C atoms in given metabolites, and numbers on the y axis represent the levels of the metabolites (μmol per g protein). CDP, cytidine diphosphate. n = 3 experimental replicates; P = 0.0013, P = 0.0061, P = 0.0035, P = 0.0228, P = 0.0129, P = 0.028 and P = 0.0021 for Carb-Asp, orotate, UMP, uridine, CMP, CDP and dUMP, respectively. Data are representative of one experiment. Error bars represent mean ± s.d. *P < 0.05; **P < 0.01; ***P < 0.001. Statistical differences were determined by unpaired two-tailed Student’s t-test (b–d and f–h) and paired two-tailed Student’s t-test (e). i, The conceptual model of ATF4-dependent and NRF2-dependent carbon assimilation and robust effector CD8+ T cell proliferation.