Effector ILC2s retain phenotypic flexibility after hematogenous trafficking and can adapt to distant tissue niches, irrespective of their origin. Retinoic acid signaling is an important driver of intestine-specific ILC2 adaptation with functional implications for acute helminth infections.
Abstract
Adaptation of immune cells to tissue-specific microenvironments is a crucial process in homeostasis and inflammation. Here, we show that murine effector type 2 innate lymphoid cells (ILC2s) from various organs are equally effective in repopulating ILC2 niches in other anatomical locations where they adapt tissue-specific phenotypes of target organs. Single-cell transcriptomics of ILC2 populations revealed upregulation of retinoic acid (RA) signaling in ILC2s during adaptation to the small intestinal microenvironment, and RA signaling mediated reprogramming of kidney effector ILC2s toward the small intestinal phenotype in vitro and in vivo. Inhibition of intestinal ILC2 adaptation by blocking RA signaling impaired worm expulsion during Strongyloides ratti infection, indicating functional importance of ILC2 tissue imprinting. In conclusion, this study highlights that effector ILC2s retain the ability to adapt to changing tissue-specific microenvironments, enabling them to exert tissue-specific functions, such as promoting control of intestinal helminth infections.
Introduction
Group 2 innate lymphoid cells (ILC2s) have emerged as important regulators of type 2 immune responses in helminth infection and allergic diseases. In the context of tissue injury, ILC2s facilitate repair processes and restoration of homeostasis by producing mediators that instruct other immune cells, directly stimulate proliferation of epithelial cells, and promote extracellular matrix remodeling (Klose and Artis, 2016). Thus, in recent years, therapeutic targeting of ILC2s has been identified as a potential strategy to enhance tissue regeneration in acute and chronic inflammation of various organs, e.g., the lung, kidney, and intestine (Düster et al., 2018; Monticelli et al., 2011, 2015; Riedel et al., 2017; Turner et al., 2013). Uncontrolled activation of ILC2s, however, can have deleterious side effects and result in organ-specific fibrosis (Hams et al., 2014; McHedlidze et al., 2013). With regard to the suitability of ILC2s as a therapeutic target, this dichotomy requires careful evaluation of ILC2 responsiveness and effector mechanisms in different anatomical locations.
ILC2s are equipped with a wide array of receptors to sense, integrate, and respond to local cues provided by other immune cells, epithelial cells, and stromal cells of the tissue niche they reside in (Karagiannis and Wilhelm 2018; Klose and Artis, 2016). Accordingly, recent studies have demonstrated that ILC2s in different non-lymphoid organs are phenotypically and functionally distinct (Gadani et al., 2017; Ricardo-Gonzalez et al., 2018; Simoni et al., 2017). Mature effector ILC2s in peripheral tissues can differentiate locally from progenitor populations (Bando et al., 2015; Zeis et al., 2020) that either seed non-lymphoid organs during fetal development and early postnatal life (Bando et al., 2015; Schneider et al., 2019) or may derive from ILC progenitors recruited from distant sites, such as the bone marrow (Bar-Ephraim et al., 2019; Lim et al., 2017; Zeis et al., 2020). While local intratissue differentiation seems to be the main mechanism for ILC2 renewal and expansion in homeostasis and acute inflammation (Gasteiger et al., 2015; Zeis et al., 2020), in chronic inflammation, ILC progenitors recruited from the circulation, and potentially, interorgan trafficking of mature ILC2s can contribute to replenishment of the ILC2 pool in affected tissues (Huang et al., 2018; Ricardo-Gonzalez et al., 2020). However, whether mature effector ILC2s in peripheral tissues retain phenotypic flexibility after hematogenous trafficking to other anatomical locations and which microenvironmental factors might shape organ-specific effector ILC2 adaptation and function is still unclear.
Here, we show that upon adoptive transfer into alymphoid recipients, murine effector ILC2s isolated from various non-lymphoid organs are equally effective in repopulating ILC2 niches in other peripheral tissues, arguing against the preferential organ-specific homing of ILC2 populations. Transferred effector ILC2s acquired the tissue-specific phenotype of organs they came to reside in—a process we refer to as “tissue adaptation.” Adaptation to the small intestinal microenvironment required retinoic acid (RA) signaling in ILC2s and inhibition of intestinal ILC2 adaptation by blocking RA signaling impaired worm expulsion after Strongyloides ratti infection.
In summary, we provide evidence for retained flexibility of mature effector ILC2s to adapt to new tissue microenvironments with important implications for organ-specific ILC2 functions, and potentially, for organ-specific therapeutic targeting of these cells.
Results
Organ-specific subset distribution and phenotype of ILC2s
Therapeutic targeting of ILCs has been identified as a potential strategy to enhance tissue repair in various organs, e.g., the kidney, lung, and intestine (Düster et al., 2018; Monticelli et al., 2011, 2015; Riedel et al., 2017; Turner et al., 2013). To evaluate ILC responsiveness and effector mechanisms in different anatomical locations, we first analyzed organ-specific subset distribution of helper-like ILCs under homeostatic conditions in the kidney, lung, and small intestinal lamina propria (SILP) of naïve C57BL/6 and BALB/c mice by flow cytometry (Fig. 1, A–C; and Fig. S1). These analyses identified GATA3+ ILC2s as a major ILC subset in all the three tissues. As previously described (Penny et al., 2018), RORγt+ ILC3s were abundant in SILP; however, they were scarce in the lungs and kidneys of naïve mice. CD127+NKp46+T-bet+Eomes− ILC1s represented a minor helper-like ILC population in all examined tissues (Fig. 1, A and C); however, low CD127 expression on kidney and lung ILC1s might result in an underestimation of ILC1 abundance in these locations with our gating strategy (Fig. S2, A and B).
Figure 1.
Organ-specific ILC subset distribution and phenotype of ILC2s. (A) Flow cytometric characterization and gating strategy of ILC subsets isolated from kidney, lung, and SILP of naïve C57BL/6 mice (blue = ILC2s, red = ILC3s, and yellow = ILC1s). Numbers indicate the percentage of events in the respective gates. (B) Frequencies and absolute numbers of total ILCs in kidney, lung, and SILP. (C) Frequencies of ILC subset distribution in the respective organs. Symbols represent individual data points and bars indicate mean ± SEM. (D) Principal component analysis of bulk RNAseq data from ILC2s purified from the indicated organs by flow cytometry. ILC2s from the kidneys and lungs of C57BL/6 mice were sorted as CD45+Lin−CD127+CD25+Sca-1+, whereas ILC2s from SILP were sorted as CD45+Lin−CD127+CD25+KLRG1+. Symbols represent individual ILC2 samples sorted from independent pools of mice (n = 3–4 samples with five mice per pool). (E) Sample distance plot based on all detected transcripts in the individual samples of the three organs. (F) Venn diagram of differentially expressed transcripts in each tissue compared with the other two tissues (false discovery rate < 0.1 and Log2FC > 1). Numbers indicate the up- (arrow up) or downregulated (arrow down) transcripts. Selected transcripts are specified for each organ with transcripts of interest marked in bold. (G) Flow cytometric analysis of Lin−GATA-3+ ILC2s from PBS- and IL-33–treated (400 ng i.p. injection on four consecutive days) wild type C57BL/6 mice (n = 3–5 mice per group). Analyses were performed at 2–3 wk after the first injection. Heat maps show geometric MFI of various surface markers of ILC2s in the indicated organs normalized to surface marker expression of kidney ILC2s. (H) UMAP clustering of the flow cytometry data from G. Lin−GATA-3+ ILC2s of the lung, kidney, and SILP were concatenated (40,000 events for each organ), and unbiased clustering was performed. Plots show combined and single-organ contribution of kidney (orange), lung (blue), and SILP (green) ILC2s to the UMAP clustering. (I) Absolute numbers of Lin−GATA-3+ ILC2s analyzed in G. (J and K) Representative flow cytometry plots (J) and absolute numbers (K) of ILC2s isolated from the kidney, lung, and SILP of PBS- (n = 8) and IL-33–treated (n = 10) wild type C57BL/6 mice. (L and M) Representative flow cytometry plots (L) and frequencies (M) of different cytokines produced by ILC2s of PBS- (n = 6) and IL-33–treated C57BL/6 mice (n = 5). Numbers in the flow cytometry plots indicate the percentage of events in the respective gates. Data in A, B, and G–I are representative of at least two independent experiments with similar results. Data in J and K are pooled from three and in L and M are pooled from two individually performed experiments. Symbols represent individual data points, and bars indicate mean ± SEM. Statistical analysis was performed using unpaired two-tailed Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001).
Figure S1.
Characterization of ILC subsets and ILC2s in different organs in BALB/c and C57BL/6 mice. (A) Flow cytometric characterization and gating strategy of ILC subsets isolated from kidney, lung, and SILP of naïve BALB/c mice (blue = ILC2s, red = ILC3s, and yellow = ILC1s). Numbers indicate the percentage of events in the respective gates. (B) Frequencies and absolute numbers of total ILCs in kidney, lung, and SILP. (C) Frequencies of ILC subset distribution in the respective organs. Symbols represent individual data points, and bars indicate mean ± SEM. (D) Flow cytometric analysis of Lin−GATA-3+ ILC2s from PBS- or IL-33–treated (i.p. injection on four consecutive days) wild type BALB/c mice at day 12–14 after the first injection. Heat maps show MFI of various surface markers of ILC2s in the indicated organs normalized to surface marker expression of kidney ILC2s. (E) Unbiased UMAP clustering of the flow cytometry data from D. Plots show combined and single-organ contribution of kidney (orange), lung (blue), and SILP (green) ILC2s to the UMAP clustering. Data are representative of at least two independent experiments with similar results with n = 3–5 mice per group. (F) Representative histogram overlays showing surface marker expression on Lin−GATA-3+ ILC2s isolated from kidney (orange), lung (blue), and SILP (green) of PBS- or IL-33–treated (i.p. injection on four consecutive days) wild type C57BL/6 mice. (G) Concatenated flow cytometry data from A shown in an unbiased UMAP clustering along with multigraph color mapping of cell surface markers on UMAP projection. Lowest expression is indicated by blue and highest expression by red. (H) Absolute numbers of different cytokine-producing ILC2s (see Fig. 1, L and M) isolated from the kidney, lung, and SILP of PBS- (n = 6) or IL-33–treated (n = 5) wild type C57BL/6 mice. Data are pooled from two individually performed experiments.
Figure S2.
Gating and sorting strategies for analysis and purification of ILCs from different organs. (A) Backgating of ILC populations on the CD45+ live lymphocyte gate in naïve C57BL/6. (B) MFI for CD127 expression on the different ILC populations in kidney, lung, and SILP. Data are representative of at least two independent experiments with similar results with n = 3–5 mice per group. (C) Sorting strategy for kidney, lung, and SILP ILC2s from naïve C57BL/6 mice without using genetically modified fluorescent reporter mice for RNAseq analyses (see Fig. 1). While the Lin−CD127+CD25+Sca-1+ gate reliably identified GATA-3+ ILC2s in the kidney and lung, SILP ILC2s were better identified by the Lin−CD127+CD25+KLRG1+ gate. (D) Gating on all GATA-3+ ILC2s showed that the gating strategy depicted in C represents >80% of the total ILC2 population in naïve C57BL/6 mice. (E) Sorting strategy for IL-33–expanded ILC2s from CD45.1 BALB/c mice (see Fig. 2, A and B, for experimental setup and sorting purity). (F) Sorting strategy for IL-33–expanded effector ILC2s from C57BL/6 Red5 mice (see Fig. 3 for sorting purity). (G) Sorting strategy for IL-33–expanded ILC2s from BALB/c IL-13fm donor mice and BALB/c Rag2−/−Il2rg−/− mice recipient mice (see Fig. 4 A for experimental setup and Fig. S3, A and B, for sorting purity). Numbers in flow cytometry plots indicate the percentage of events in the repective gates. Symbols in B represent individual data points and bars indicate mean ± SEM. Statistical analysis was performed using one-way ANOVA with Tukey’s post-hoc test (*P < 0.05, **P < 0.01, ***P < 0.001).
In the following, we focused on the ILC2 populations found in all three organs. To explore how tissue-specific factors shape the transcriptome of ILC2s in different anatomical locations at steady state, we next performed bulk RNA sequencing (RNAseq) of ILC2s purified by flow cytometry from kidney, lung, and SILP of naïve mice (Fig. 1, D–F). The sorting strategy was designed based on the ILC2 surface marker expression to exclude potential transcriptional alterations by reporter transgenes (Fig. S2, B–C). To obtain high purity (99%, Fig. S2 C) and coverage (>80%, Fig. S2 D) of the Lin−CD127+GATA3+ ILC2 population, a combination of Sca-1 and CD25 was found to be optimal for ILC2 isolation from kidney and lung, while SILP ILC2s were best identified as CD25+KLRG1+. Unbiased clustering in principal component analyses revealed a tissue-specific distinct transcriptomic signature of ILC2s (Fig. 1 D). However, a sample distance analysis based on all detected transcripts demonstrated that kidney and lung ILC2s were transcriptionally more closely related compared with SILP ILC2s (Fig. 1 E). Several hundred transcripts were differentially expressed between ILC2s isolated from different locations (Fig. 1 F). Among these, numerous genes regulating responsiveness to environmental stimuli (cytokines and growth factors) and cell migration/adhesion were specifically up- or downregulated in each organ.
To characterize the phenotypic differences of ILC2s in the kidney, lung, and SILP with a second approach, we selected a panel of cytokine receptors and other common ILC2 surface markers and analyzed the ILC2 phenotype by flow cytometry in unchallenged C57BL/6 and BALB/c mice and after activation of ILC2s by i.p. IL-33 injection (Fig. 1, G and H; and Fig. S1, F and G). The comparison of geometric mean fluorescence intensity (MFI) of these surface markers among ILC2 populations confirmed our RNAseq results at protein level (see Il17rb, Il1lr1, and Thy1), e.g., with high expression of IL-17RB and KLRG1 but low expression of IL-33R and CD90.2 in small intestinal ILC2s (Fig. 1 G and Fig. S1). To visualize similarities and differences of the tissue-specific ILC2 phenotypes in an unsupervised approach, we next concatenated the flow cytometry data from different organs followed by dimensionality reduction and clustering using the Uniform Manifold Approximation and Projection (UMAP) algorithm (Fig. 1 H and Fig. S1). In line with the transcriptomic analyses, this approach revealed a substantial phenotypic overlap of kidney and lung ILC2s, while ILC2s from SILP formed a clearly separated cluster, which was preserved after IL-33 treatment.
While SILP ILC2s express much lower levels of IL-33R than kidney and lung ILC2s, they are not completely negative (compare Fig. S1 F). In line, they show accumulation and increased cytokine production in response to systemic IL-33 treatment, as used for in vivo ILC2 expansion in our experiments (Fig. 1, I–M; and Fig. S1 H). Consistent with lower IL-33R expression, IL-33–mediated in vivo ILC2 expansion was less prominent in the SILP (approximately factor 5) compared with lung and kidney (approximately factor 10–20; Fig. 1 I). Although the frequency Ki67 positivity in IL-33–expanded ILC2s at the time of analysis (2–3 wk after the first IL-33 injection) was not increased, the absolute number of Ki67+ ILC2s was significantly higher after IL-33 treatment in all organs, indicating recent proliferation (Fig. 1, J and K). The cytokine production profile differed between the three organs as SILP ILC2s, in general, produced more cytokines compared with kidney and lung ILC2s (Fig. 1, L and M), while lung ILC2s particularly showed lower IL-4 production than the other organs. The distinct surface marker phenotype (e.g., IL-33RloIL-17RBhiCD90.2loKLRG1hiICOSlo in SILP, Fig. 1 G) and cytokine production profile (Fig. 1, L and M) were preserved after application of IL-33, indicating that tissue imprinting of ILC2s is maintained during IL-33–induced ILC2 activation and expansion in the tissues.
Taken together, these data suggest that tissue-specific factors in the small intestine drive a unique ILC2/ILC3 subset distribution and a distinct ILC2 phenotype.
ILC2 phenotype is dictated by the tissue microenvironment
Next, we asked the question of whether these tissue-specific ILC2 phenotypes are irreversibly imprinted during differentiation or whether ILC2s from peripheral sites retain the ability to adapt to new tissue microenvironments. To address this, we established an adoptive transfer model in which highly purified congenitally marked ILC2s from the kidney, lung, and SILP of IL-33–treated mice were transferred into alymphoid Rag2−/−Il2rg−/− mice, and tissue repopulation was assessed after 3 wk (Fig. 2, A–D). In these experiments, ILC2s were sorted as CD45+Lin−CD127+KLRG1+, allowing for reliable identification and high-purity sorting of ILC2s from all anatomical locations in IL-33–treated mice (Fig. S2 E and Fig. 2 B). Flow cytometric analyses of the recipient mice revealed that CD45.1+ ILC2s repopulated all tissues examined, regardless of the origin of transferred ILC2 populations. The distribution of ILC2 frequencies and numbers in all kidneys, lungs, and SILP was reminiscent of wild type mice (Fig. 2, C and D; and Fig. 1, A and B). Moreover, transferred ILC2s, again regardless of their tissue of origin, adopted the surface marker expression pattern specific for the respective target organ, as seen in wild type mice without cell transfer (Fig. 2, E and F; and Fig. 1 G).
Figure 2.
ILC2 phenotype is dictated by the tissue microenvironment. (A) Schematic representation of the adoptive transfer model. ILC2s (sorted as CD45+Lin−CD127+KLRG1+) isolated from kidney, lung, and SILP of IL-33–treated (i.p. injection on four consecutive days) C57BL/6 CD45.1 wild type mice were adoptively transferred i.v. into C57BL/6 Rag2−/−Il2rg−/− mice. Tissues from the recipient mice were analyzed after 3 wk of reconstitution. (B) Flow cytometry plots of ILC2 sorting purity from A. The numbers represent frequencies of sorted ILC2s in the total lymphocyte population. (C–F) Flow cytometric analysis of leukocytes isolated from the kidney, lung, and SILP of C57BL/6 Rag2−/−Il2rg−/− mice after 3 wk of ILC2 transfer and without transfer. (C) Representative plots comparing CD45.1+ ILC2s isolated from the three indicated organs of recipient mice (rows) with transferred ILC2 populations derived from different organ origins (columns). Numbers indicate the percentage of cells in each gate. (D) Frequencies and absolute numbers of CD45.1+ ILC2s in kidney, lung, and SILP at 3 wk after transfer. Symbols represent individual data points, and bars indicate mean ± SEM (n = 4 per organ analyzed). (E) Representative histogram overlays showing surface marker expression of kidney, lung, and SILP CD45.1+ ILC2s isolated after 3 wk of reconstitution. (F) Heat maps of the MFI of various surface markers of ILC2s 3 wk after ILC2 transfer normalized to surface marker expression of kidney ILC2s. All data are representative of two independent experiments with at least three animals per group.
These data argue against tissue-specific homing capacity and irreversible imprinting of ILC2s during differentiation and provide evidence for continuous adaptation of mature effector ILC2s to the tissue microenvironment they reside in.
Effector ILC2s can adapt to new tissue microenvironment
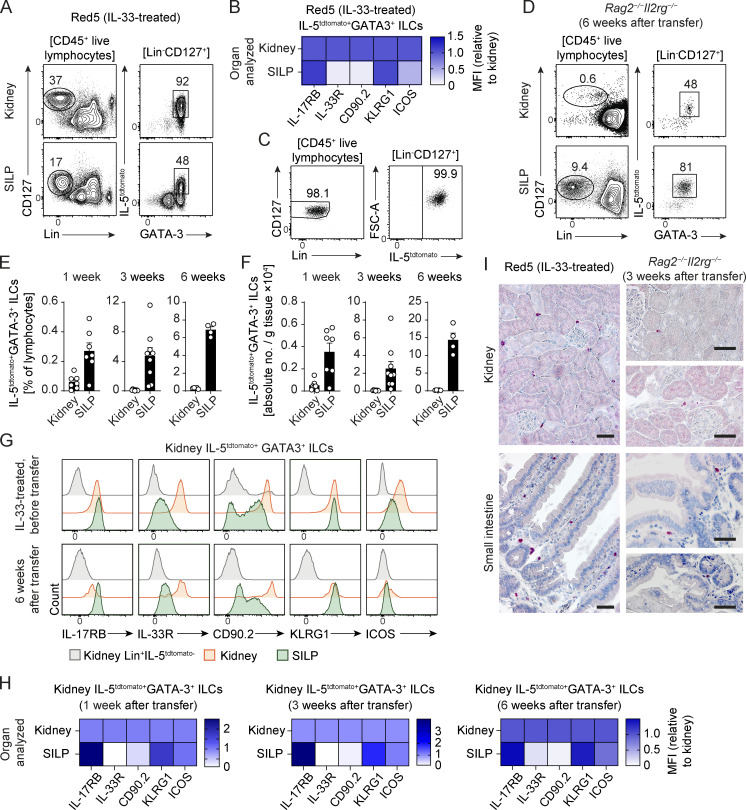
Next, we wanted to investigate whether tissue adaptation of ILC2s was due to true flexibility of mature effector ILC2s rather than a result of tissue ILC progenitors that might contaminate the transferred Lin−CD127+KLRG1+ ILC2 populations, seed the empty niches, and proliferate in target tissues. Since tissue ILC2 progenitors were recently shown to express only intermediate levels of IL-5 (Zeis et al., 2020), we sorted IL-5hi effector ILC2s from the kidney of IL-33–treated IL-5 reporter (Red5) mice to high purity, transferred them into alymphoid Rag2−/−Il2rg−/− mice, and assessed tissue repopulation after 1–6 wk (Fig. 3). In these experiments, ILC2s were sorted as CD45+Lin−CD127+IL-33R+KLRG1+IL-5tdTomato hi to exclude IL-5lo ILC2 tissue precursor populations (Fig. S2 F and Fig. 3 C). The IL-5tdTomato+GATA3+ ILC2s exhibited similar surface marker expression as seen in the total GATA3+ ILC2s population in wild type mice, with high expression of IL-17RB and KLRG1 and low expression of IL-33R, CD90.2, and ICOS in SILP compared with kidney (Fig. 3, A, B, and G, upper row).
Figure 3.
Effector ILC2s can adapt to new tissue microenvironment. (A) Flow cytometric analysis of IL-5tdtomato+GATA3+ ILCs isolated from kidney and SILP of IL-33–treated Red5 mice (i.p. injection on four consecutive days). Numbers indicate the percentage of cells in each gate. (B) Heat map of MFI of different surface markers on IL-5tdtomato+GATA3+ ILCs isolated from kidney and SILP normalized to surface marker expression of kidney ILC2s. (C) Representative sorting purity of transferred ILC2s (sorted as CD45+Lin−CD127+IL-33R+KLRG1+IL-5tdtomato+) originating from the kidney of IL-33–treated Red5 mice. Sorted kidney ILC2s were transferred to C57BL/6 Rag2−/−Il2rg−/−. (D–F) Representative flow cytometric plots (D), frequencies (E), and absolute numbers (F) of IL-5tdtomato+GATA3+ ILC2s obtained from the kidney and SILP (n = 4–7 per organ) of C57BL/6 Rag2−/−Il2rg−/− mice at different time points after transfer. Numbers in D indicate the percentage of events in each gate. Symbols represent individual data points, and bars indicate mean ± SEM. (G and H) Representative histogram overlays (G) and heat map of MFI (H) of different surface markers of IL-5tdtomato+GATA3+ ILCs originating from the kidney of Red5 mice analyzed 1–6 wk after transfer in the kidney and SILP of C57BL/6 Rag2−/−Il2rg−/− mice and normalized to surface marker expression on kidney ILC2s. (I) Immunohistochemical staining of tdTomato-positive ILC2s (deep red) in the kidney and SILP tissue of IL-33–treated Red5 mice and in C57BL/6 Rag2−/−Il2rg−/− mice 3 wk after kidney IL-5tdtomato+ ILC2 transfer. Scale bars, 50 µm. Data in D–H represent five independent experiments.
Already at 1 wk after transfer, high-purity Lin−CD127+IL-33R+KLRG1+IL-5tdTomato hi effector ILC2s started to repopulate not only kidneys but also SILP. ILC2 numbers and frequencies in tissues further increased over time (Fig. 3, D–F). Of note, the ratios between kidney and SILP ILC2s were comparable with that observed in wild type mice at all time points (Fig. 3, D–F). Irrespective of their kidney origin, within 1 wk, transferred ILC2s adopted the phenotype of the target tissues, as assessed by our defined set of surface markers (Fig. 3, G and H). To visualize ILC2s in their respective tissue niches, we performed immunohistochemistry for tdTomato-positive ILC2s in kidney and SILP. We observed that kidney ILC2s resided in the tubulointerstitial compartment and periglomerular space, and SILP ILC2s were found in villi and close to the crypt bases in both IL-33–treated Red5 mice and Rag2−/−Il2rg−/− mice 3 wk after ILC2 transfer (Fig. 3 I).
While formal proof of mature effector ILC2 adaptation would require clonal single-cell readouts, these data further substantiate that ILC2s seem to retain flexibility even after differentiation to the effector state and indicate that migrating effector ILC2s can occupy the same niche as the resident ILC2 populations.
Adaptable, non-adaptable, and hardwired features of organ-specific ILC2 transcriptional programs
Next, we used an unbiased approach to investigate transcriptional programs specific to kidney and SILP ILC2s, and subsequently, to delineate which features of these programs are flexible during adaptation to the small intestinal microenvironment. To this end, we adoptively transferred kidney ILC2s from IL-33–treated Il13Cre × R26reYFP (IL-13 fate mapping mice, IL-13fm) mice to Rag2−/−Il2rg−/− mice and analyzed transferred ILC2s isolated from SILP of recipient mice at two time points after transfer (2 and 8 wk) using single-cell RNAseq (scRNAseq; see Fig. 4 A for experimental setup). In these experiments, ILC2s were sorted as CD45+Lin−CD127+IL-13fm+, which allowed for unequivocal identification of transferred ILC2 populations in recipient mice, facilitated cell sorting, and thereby minimized contamination in scRNAseq analyses (Fig. S2 G; and Fig. S3, A and B). In addition, kidney and SILP ILC2s from IL-13fm donor mice were also subjected to scRNAseq. scRNAseq analysis of IL-13fm+ donor ILC2s isolated from the kidney again confirmed the mature effector identity of the transferred population with high levels of Il5, Il13, Klrg1, and Il1rl1 transcripts and minimal (co-) expression of the recently described ILC2 progenitor markers Tcf7, Il18r1, Pdcd1 (encoding for PD-1), and Zbtb16 (encoding for PLZF; Zeis et al., 2020; Fig. S3, C–F).
Figure 4.
Adaptable and hardwired features of kidney and SILP ILC2s. (A) Schematic representation of the experimental setup for scRNAseq of ILC2 populations. ILC2s from kidney and SILP were isolated from IL-33–treated BALB/c IL-13fm mice (i.p. injection on four consecutive days) and sorted for scRNA sequencing analysis (donors). The cells were sorted as CD45+Lin−CD127+IL-13fm+. Simultaneously, kidney ILC2s from the same isolated cell pool were adoptively transferred i.v. into BALB/c Rag2−/−Il2rg−/− mice. After 2 and 8 wk, IL-13fm+ ILC2s were recovered from SILP and subjected to scRNAseq analysis. (B) Unsupervised UMAP clustering of combined scRNAseq data of kidney and SILP ILC2s from donors and of SILP ILC2s after 2 and 8 wk from recipients. (C) Heat map of differentially expressed ILC2 surface markers in donor and recipient clusters. (D) DE analysis between donor kidney and SILP ILC2s. Genes that are significantly upregulated with ≥twofold change are shown as red dots. (E) Gene set enrichment analysis (GSEA) for HALLMARK pathways of the Mouse Molecular Signatures Database using the genes significantly upregulated in SILP (left, dark blue) and kidney (right, dark green) ILC2s. (F) Violin plots depicting the expression score of SILP program and kidney program, defined as the organ-specific upregulated genes in the DE analyses in D, in the respective conditions. The indicated comparisons represent adaptable SILP features (= comp. 1), non-adaptable SILP features (= comp. 2), and hardwired kidney features (= comp. 3). (G) DE expression analysis of the comparisons indicated in F. Significantly upregulated genes (≥twofold) in the indicated samples are shown as red dots. (H) Venn diagram of SILP-specific and kidney-specific HALLMARK pathways from E and classification of the pathways as indicated. (I) Specific adaptable and non-adaptable HALLMARK pathways in SILP ILC2s identified by filtering as shown in H.
Figure S3.
Sorting purity and scRNAseq analysis of ILC2s. (A) Representative sorting purity of ILC2s (sorted as Lin−CD127+IL-13fm+) isolated from the kidney and SILP of IL-33–treated BALB/c IL-13fm mice (donors) for scRNAseq. (B) Representative sorting purity of ILC2s (sorted as Lin−CD127+IL-13fm+) isolated from the SILP (2 and 8 wk after transfer) of BALB/c Rag2−/−Il2rg−/− mice that received adoptive i.v. transfer of kidney ILC2s. (C) Unsupervised UMAP clustering of kidney ILC2s from donor mice. (D) Heat map of top 15 cluster-defining genes obtained from C. (E) Feature plots depicting expression of key markers on the UMAP of the kidney ILC2 dataset from C. (F) Overlay of Tcf7 and Il18r1 genes known to be co-expressed in ILC precursors.
Unbiased UMAP clustering of the merged scRNAseq data set revealed four distinct clusters, one for each organ analyzed at different time points (Fig. 4 B and Fig. S4 A). As expected, donor kidney and SILP ILC2s clustered separately from each other and from SILP ILC2s of recipient mice, whereas SILP ILC2s obtained 2 and 8 wk after transfer clustered together. Analysis of mRNA expression of the defined surface marker set in the ILC2 clusters confirmed our previous flow cytometry–based findings on mRNA level of the switch of transferred kidney ILC2s to the SILP ILC2 phenotype after transfer (Fig. 4 C). To identify organ-specific ILC2 transcriptional programs, we first performed differential expression (DE) analyses of donor kidneys versus donor SILP ILC2s (Fig. 4 D). The organ-specific upregulated genes in this comparison were defined as the “kidney program” and the “SILP program,” respectively, and subjected to HALLMARK pathway gene set enrichment analyses using the Mouse Molecular Signatures Database (Fig. 4 E). These analyses revealed several pathways upregulated in SILP ILC2s (n = 27), some of which were immune cell related, while kidney ILC2s showed upregulation of a smaller number (n = 6) of metabolic and general signaling pathways (Fig. 4 E). In line with rapid ILC2 tissue adaptation, the kidney program was markedly downregulated in adapting kidney ILC2s already at 2 wk and almost completely lost at 8 wk after transfer (Fig. 4 F, right panel). The (partial) upregulation of the SILP program in ex-kidney ILC2s was also already observed at 2 wk after transfer and increased until 8 wk but remained incomplete even at the late time point (Fig. 4 F, left panel). Next, we defined three DE comparisons between the groups (Fig. 4, F and G) that identify differentially regulated gene sets, representing the “adaptable” SILP features (comparison 1), “non-adaptable” SILP features (comparison 2), and “hardwired” kidney features (comparison 3). These gene sets were then subjected to HALLMARK pathway gene set enrichment analyses (Fig. S4 B). However, several of the upregulated pathways seemed to be unspecific (e.g., upregulated in all three comparisons) or related to the adoptive transfer setting into alymphoid Rag2−/−Il2rg−/− mice (upregulated in ex-kidney SILP ILC2s at 8 wk in comparisons to both non-transfer settings). Thus, we used a Venn diagram of SILP-specific pathways to identify “fully adaptable” SILP features, “partially adaptable” SILP features, and “non-adaptable” SILP features (Fig. 4 H, upper panel). A similar strategy was used for filtering kidney-specific pathways (Fig. 4 H, lower panel). Interestingly, when using these filtering steps, we failed to extract any “hardwired” kidney ILC2 features from the data. In contrast, several immune-related pathways, e.g., TNF/NF-κB signaling, were found among the (partially) adaptable SILP features that were acquired by ex-kidney ILC2s in the small intestine (Fig. 4 I). Non-adaptable SILP features were related to DNA replication, DNA repair, as well as Notch signaling and response to estrogens (Fig. 4 I).
Figure S4.
Cluster-defining genes and pathway analysis of scRNAseq data. (A) Heat map depicting top 10 cluster-defining genes for each cluster obtained from scRNAseq analysis of donor kidney and SILP ILC2s as well as recipient SILP ILC2s (see Fig. 4 A for experimental setup). Genes associated with RA and Notch signaling are marked with an arrow and labeled in red. (B) Gene set enrichment analysis (GSEA) for HALLMARK pathways of the Mouse Molecular Signatures Database using the genes significantly upregulated in the indicated comparisons between ILC2 populations (see Fig. 4 A for experimental setup). Specific pathways identified by further filtering (see Fig. 4, G and H) are indicated by blue boxes and marked in bold.
Adapting ILC2s gradually acquire the transcriptional program of small intestinal ILC2
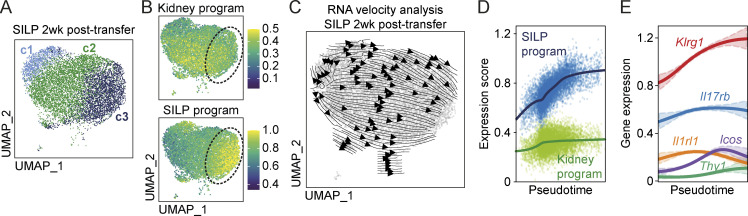
To further investigate the dynamics of intestinal ILC2 adaptation after transfer, we focused on scRNAseq analysis of ex-kidney ILC2s isolated from SILP at 2 wk after transfer. At this time point, adapting kidney ILC2s have acquired a substantial part of the SILP program while still expressing a residual kidney signature (see Fig. 4 F). UMAP clustering of the 2-wk sample revealed three clusters (Fig. 5 A), of which cluster 3 showed substantial upregulation of the SILP program and expressed the kidney program at a low level (Fig. 5, A and B). To identify the dynamics of the adaptation process, we used RNA velocity analysis that, based on the amount of unspliced and spliced mRNAs in each cell, assigns directed, dynamic information to the state of each cell in a transition process (Fig. 5 C; La Manno et al., 2018). The directional information derived from this approach showed a trajectory from cluster 2 with low SILP program expression toward cluster 3 with high SILP program expression (Fig. 5 C). RNA velocity-based pseudotime analyses demonstrated a gradual increase in the SILP program and most of our defined SILP ILC2 surface markers in adapting ex-kidney ILC2s at 2 wk after transfer (Fig. 5, D and E). The residual kidney program (including Il1rl1) remained stable over pseudotime, indicating that most kidney features were already downregulated within the first 2 wk of the adaptation process (Fig. 5, D and E; see Fig. 4 F).
Figure 5.
Gradual acquisition of the SILP ILC2 phenotype by transferred kidney ILC2s. (A) Unsupervised UMAP subclustering of scRNAseq data of ex-kidney ILC2s isolated from the SILP of recipient BALB/c Rag2−/−Il2rg−/− mice 2 wk after transfer (see Fig. 4 A for experimental setup). (B) Color mapping of kidney and SILP program expression on the UMAP subclustering of adapting ex-kidney ILC2s at 2 wk after transfer. Lowest expression is indicated by blue and highest expression by yellow. (C) Dynamic information of cell states in the transition process generated using RNA velocity analysis of adapting ex-kidney ILC2s at 2 wk after transfer. (D and E) RNA velocity-based pseudotime analyses of the SILP and kidney programs (D) and selected surface markers (E) in adapting ex-kidney ILC2s at 2 wk after transfer.
Taken together, these data further support the hypothesis that mature effector ILC2s can gradually adapt the tissue-specific ILC2 transcriptional program of a new microenvironment.
RA signaling activity is increased in ILC2s adapting to the small intestine
RA is an important mediator in the intestinal microenvironment and its production by SILP dendritic cells has been shown to imprint intestine-specific features on T cells (Iwata, 2009). Two RA receptors, Rara and Rxrg, were among the significantly upregulated genes in bulk-like DE analysis of the scRNAseq data from SILP versus kidney ILC2s. In addition, Rxrg was found among the top 10 cluster-defining genes of SILP-adapted ex-kidney ILC2s in the scRNAseq analyses (Fig. S4 A). Expression analyses of all known RA receptors in our single-cell and bulk RNAseq data sets showed that Rara, Rxrb, and Rxrg expression was higher in SILP ILC2s than in kidney ILC2s, while Rarb, Rarg, and Rxra expression was lower in SILP ILC2s (Fig. 6, A and B). In addition, screening of SILP ILC2 program genes (Fig. 4 D) and of the top 30 cluster-defining genes upregulated in the SILP-adapted ex-kidney ILC2 clusters (Fig. S4 A) identified several genes that were previously described to be induced by RA signaling (Nrip1, Tgif1, Ctsh, Dusp1, Irf1; Fig. 6, C and D). To assess RA signaling in adapting ILC2s in a broader sense, we generated an RA signaling score by combining Gene Ontology (GO) terms for RA receptor binding (GO:0046965), RA receptor signaling pathway (GO:0048384), and response to RA (GO:0032526) in one expression module (Fig. 6 E). This expression score was significantly upregulated in all SILP ILC2 clusters from the transfer experiments compared with kidney ILC2s (Fig. 6 E, see Fig. 4, A and B). In addition, analyses focused on ex-kidney ILC2s isolated from the SILP at 2 wk after transfer (compare Fig. 5) showed upregulation of the RA module in SILP phenotype ILC2s (cluster 3; Fig. 6 F). RNA velocity-based pseudotime analyses of adapting ILC2s demonstrated gradual upregulation of four out of the five RA signaling genes during acquisition of the SILP ILC2 program (Fig. 6 G and Fig. 5 D), suggesting a role for RA signaling activity in ILC2 adaption to the intestinal microenvironment.
Figure 6.
RA signaling drives functional adaptation of ILC2s to the intestinal phenotype. (A) Dot plot of RA receptor expression in the scRNAseq data set of kidney and SILP ILC2s from donors and of SILP ILC2s after 2 and 8 wk from recipients of kidney ILC2s (see Fig. 4 A for experimental setup). (B) Heat map of RA receptor expression in bulk RNAseq data set of ILC2s isolated from the kidneys, lungs, and SILP of naïve C57BL/6 mice (see Fig. 1). (C) Dot plot of RA signaling genes selected from the DE analyses (see Fig. 4 D) and top 30 cluster-defining genes (see Fig. S4 A). (D) Heat map of RA signaling genes in the bulk RNAseq data set of ILC2s isolated from naïve C57BL/6 mice (see Fig. 1). (E) Violin plots depicting the expression score of RA signaling associated genes in the respective conditions. (F) Color mapping of RA expression score on the UMAP subclustering of adapting ex-kidney ILC2s at 2 wk after transfer (see Fig. 5). Lowest expression is indicated by blue and highest expression by yellow. (G) RNA velocity-based pseudotime analyses of selected RNA signaling genes in adapting ex-kidney ILC2s at 2 wk after transfer. (H and I) Representative histogram overlays (H) and MFI (I) for surface marker expression of sorted kidney ILC2s after culture with IL-2 (n = 4 for each condition) in the presence or absence of RA (1 µM). (J) Cytokine quantification in the supernatant of the cultured kidney ILC2s stimulated with subthreshold IL-25 or IL-33 (1 ng/ml each; n = 4 for each condition). Data in H–J are representative of three individual experiments with similar results. Symbols represent individual data points and bars indicate mean ± SEM. Statistical analysis was performed using unpaired two-tailed Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001). (K) Hypothetical model of RA signaling network in SILP ILC2s based on genes upregulated in the RNAseq data sets and a literature review of described downstream pathways. CREB, cAMP response element-binding protein; ISG, interferon-stimulated gene.
RA signaling drives functional adaptation of ILC2s to the intestinal phenotype in vitro
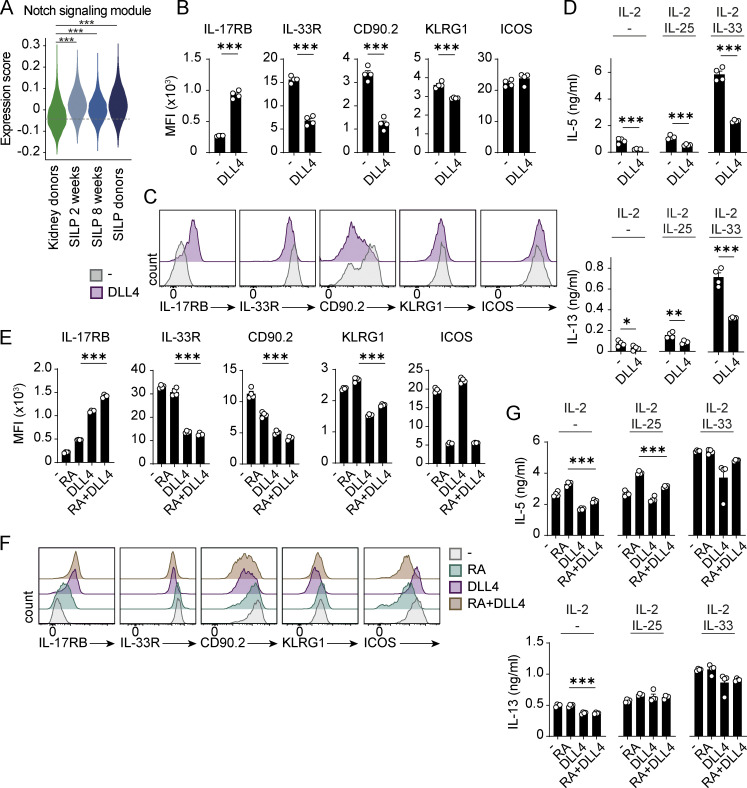
To validate the hypothesis that RA signaling activity drives phenotypic ILC2s adaptation to the intestinal microenvironment, we next cultured kidney ILC2s with or without RA and analyzed selected surface markers using flow cytometry after 4 d of culture (Fig. 6, H and I). Indeed, we observed that cells treated with RA upregulated IL-17RB and KLRG1 expression while downregulating CD90.2 and ICOS, mimicking the “intestinal” ILC2 phenotype, with the exception that we did not observe RA-mediated downregulation of the IL-33R (Fig. 6, H and I). To further validate whether downregulation of IL-17RB was of functional relevance, we studied the responsiveness of cultured kidney ILC2s to IL-25 and IL-33 after RA-induced switch to the intestinal phenotype. These experiments confirmed that RA treatment not only upregulates IL-17RB expression on ILC2s but also increases their responsiveness to subthreshold IL-25 (1 ng/ml), as measured by enhanced production of IL-5 and IL-13 in RA-preconditioned cells (Fig. 6 J). In line with unchanged receptor expression, responsiveness to IL-33 was unaltered by RA (Fig. 6 J). These data identify RA signaling as one important regulator of the phenotypic switch of ILC2s to the SILP microenvironment. Based on the RA-induced genes that were upregulated during intestinal ILC2 adaptation in our RNAseq data sets and a literature review of described downstream pathways (Bartholin et al., 2006; Borowczyk et al., 2021; Jeffrey et al., 2007; Lu et al., 2008; Samarut and Rochette-Egly, 2012; White et al., 2003; Zhuang et al., 2021), we devised a hypothetical model of the broader consequences of RA signaling on intestinal ILC2 biology (Fig. 6 K).
Role of Notch signaling for adaptation of ILC2s to the intestinal phenotype in vitro
While we established RA signaling as one driving factor for ILC2 adaptation to the intestinal microenvironment, other tissue cues most likely act in concert with RA to induce their full phenotypic switch (including, e.g., IL-33R downregulation). The central adaptor of the Notch pathway, Hes1, was the most upregulated gene in the SILP program (see Fig. 4 D) and was also found among the 10 cluster-defining genes of SILP-adapted ex-kidney ILC2s in the scRNAseq analyses (Fig. S4 A). Although in unbiased analyses, the HALLMARK Notch signaling pathway was identified as one of the non-adaptable SILP features (see Fig. 4 I, lower panel), Hes1 was strongly upregulated in adapting ex-kidney ILC2s (see Fig. 4 G, comparison 1). In line, an expression score of genes related to Notch signaling (KEGG PATHWAY: hsa04330) was significantly upregulated in all SILP ILC2 clusters compared with kidney ILC2s (Fig. S5 A), pointing to a role of Notch signaling in ILC2 adaptation to the SILP. To address this, we performed another set of in vitro experiments, culturing kidney ILC2s in the absence and presence of the Notch ligand DLL4 (Fig. S5, B and C). Although some features of the intestinal ILC2 phenotype in kidney ILC2s could be induced by activation of Notch signaling (upregulation of IL-17RB, downregulation of IL-33R and CD90.2), other markers remained unchanged (ICOS) or were regulated inversely (downregulation of KLRG1; Fig. S5, B and C). Importantly, despite upregulation of IL-17RB, DLL4 stimulation significantly decreased IL-25 responsiveness of ILC2s, indicating that activation of the Notch pathway might be a general inhibitor of ILC2 function (Fig. S5 D). Finally, a combination of DLL4 and RA treatment was able to induce ICOS downregulation but failed to restore upregulation of KLRG1 and the increased cytokine responsiveness observed with RA treatment alone (Fig. S5, E–G). In conclusion, in addition to RA and Notch signaling, other unknown factors are required to simulate a full phenotypic and functional switch of kidney ILC2s to the intestinal ILC2 program in vitro.
Figure S5.
Notch signaling in intestine-specific adaptation of kidney ILC2s in vitro. (A) Violin plots depicting Notch signaling score in the scRNAseq data of kidney and SILP donors and recipient mice (see Fig. 4 A for experimental setup). (B and C) MFI (B) and representative histogram overlays (C) of surface marker expression of sorted kidney ILC2s after in vitro culture with IL-2 in the presence or absence of the Notch ligand DLL4 (2.5 µg/ml). Cells were sorted as CD45+Lin−CD127+IL-33R+KLRG1+ from the kidney of IL-33–treated C57BL/6 mice and cultured for 4 d in a 96-well plate (∼10,000 cells/well; n = 4 for each condition). (D) Cytokine quantification in the supernatant of the cultured kidney ILC2s stimulated with subthreshold IL-25 or IL-33 (1 ng/ml each). Cells were stimulated on day 3 for 24 h. (E and F) MFI (E) and representative histogram overlays (F) for surface marker expression of kidney ILC2s after culture with IL-2 in the presence or absence of RA (1 µM) and DLL4 (n = 4 for each condition). (G) Cytokine quantification in the supernatant of the cultivated kidney ILC2s stimulated as in E. Data are representative of three individual experiments with similar results. Symbols represent individual data points and bars indicate mean ± SEM. Statistical analysis was performed using unpaired two-tailed Student’s t test (B and D) or one-way ANOVA with Tukey’s post-hoc test (E and G); *P < 0.05, **P < 0.01, ***P < 0.001.
RA is essential for intestinal adaptation of ILC2s in vivo
As RNAseq and in vitro analyses identified RA signaling as one factor for adaptation of intestine-specific ILC2 phenotype, we decided to validate this effect in vivo by blocking RA signaling using a pan-RA receptor inverse agonist (Spencer et al., 2014; BMS493; hereafter referred to as retinoic acid inhibitor [RAi]). For this, we performed an adoptive transfer of IL-13fm+ ILC2s into Rag2−/−Il2rg−/− mice and treated the recipients with RAi from 1 d prior to cell transfer for 3 wk. Control recipients received DMSO injections (see Fig. 7 A for experimental setup). As observed in previous analyses, transferred kidney ILC2s occupied the niches in kidney, lung, and SILP in ratios comparable to wild type mice in both groups (Fig. 7, B and C) and adopted the intestinal IL-17RBhiIL-33RloCD90.2loKLRG1hiICOSlo phenotype in the SILP of DMSO-treated mice (Fig. 7 D). In line with our hypothesis, RAi prevented the adaptation of kidney ILC2s to the SILP phenotype, reducing upregulation of IL-17RB and KLRG1, while increasing expression of CD90.2 and ICOS on ex-kidney SILP ILC2s (Fig. 7 D). As observed under in vitro conditions, IL-33R expression remained unchanged between the groups.
Figure 7.
RA signaling is essential for intestinal adaptation of ILC2s in vivo. (A) Experimental design of the RAi (BMS493 220 µg) treatment of ILC2-transferred BALB/c Rag2−/−Il2rg−/− mice. Kidney ILC2s isolated from IL-33–treated BALB/c IL-13fm mice (i.p. injections on four consecutive days) were adoptively transferred through i.v. injections into BALB/c Rag2−/−Il2rg−/− recipient mice (n = 11). The recipient mice received either DMSO (n = 5) or RAi (n = 6) treatment at the indicated time points. (B) Representative FACS plots of Lin−IL-13fm+ ILC2s isolated from the kidneys, lungs, and SILP of BALB/c Rag2−/−Il2rg−/− mice after 3 wk of transfer and treatment with either DMSO or RAi. Numbers indicate the percentage of cells in each gate. (C) Frequencies and absolute numbers of IL-13fm+ ILC2s isolated from the kidney, lung, and SILP of BALB/c Rag2−/−Il2rg−/− mice treated with either DMSO or RAi. (D) MFI of different surface marker expressions of ILC2s isolated from the respective organs. (E) Experimental design of the RAi treatment of BALB/c Rag2−/− and BALB/c IL-13fm mice. The mice received either DMSO (n = 5) or RAi (n = 7) treatment for 3 wk, and ILC2s from the kidney, lung, and SILP were analyzed. (F and G) Absolute numbers and ILC2/ILC3 ratio of Lin−GATA-3+ ILC2s, Lin−IL-13fm+ ILC2s, and Lin−RORγt+ ILC3s isolated from the kidney, lung, and SILP of BALB/c Rag2−/− (F) and BALB/c IL-13fm (G) mice. (H and I) MFI of different surface marker expression of ILC2s isolated from the respective organs of BALB/c Rag2−/− (H) and BALB/c IL-13fm mice (I). Data are pooled from two individually performed experiments. Symbols represent individual data points and bars indicate mean ± SEM. Statistical analysis was performed using unpaired two-tailed Student’s t test (*P < 0.05, **P < 0.01, ***P < 0.001).
As cellular adaptation to tissues should occur constitutively, even at steady state, we next asked the question of whether RA signaling inhibition could also reverse intestinal adaptation of endogenous ILC2s in a non-transfer setting. To address this, we treated T cell–deficient Rag2−/− and immunocompetent Il13fm mice with RAi for 3 wk and assessed abundance and phenotype of ILC2s in kidney, lung, and SILP (Fig. 7, E–I). As previously observed (Spencer et al., 2014), RAi treatment led to a significant increase in ILC2 abundance in the SILP with a shift in the ILC2/ILC3 ratio (Fig. 7, F and G). Endogenous SILP ILC2s from RAi-treated mice significantly downregulated IL-17RB and KLRG1 expression, while upregulating CD90.2 and ICOS expression (Fig. 8, H and I), indicating a partial reversal of the intestinal ILC2 phenotype, as defined by our surface marker set. Taken together, these data confirm that RA contributes to continuous phenotypic ILC2 adaptation to the small intestinal microenvironment in vivo.
Figure 8.
RA signaling inhibition reverses intestinal ILC2 adaptation and impairs anti-helminth immunity. (A) Experimental setup of RAi treatment and subsequent S. ratti infection in BALB/c Rag2−/− and BALB/c Rag2−/−Il2rg−/− mice. (B) Representative flow cytometric plots of ILC2 and ILC3 populations isolated from the colon of BALB/c Rag2−/− mice treated with either DMSO or RAi and infected with S. ratti. In S. ratti experiments, colon was analyzed for the RAi effect on ILC2s because the small intestine was needed for worm counting. Gating strategy is indicated in brackets and numbers indicate the percentage of cells in each gate. (C) Frequencies (upper panel) and absolute numbers (lower panel) of GATA-3+ ILC2s and RORγt+ ILC3s populations as well as their proportional shift (right panel) in the colon of BALB/c Rag2−/− mice infected with S. ratti after 3 wk of DMSO or RAi treatment. (D and E) Representative histogram overlays (D) and MFI (E) of the surface marker expression in the two groups. (F) Worm burden in the small intestine of Rag2−/− and Rag2−/−Il2rg−/− mice 6 d after infection with S. ratti. (G) Mast cell activity assessed as mMCPT-1 serum concentration in Rag2−/− and Rag2−/−Il2rg−/− mice infected with S. ratti. Data are pooled from two (Rag2−/−) or three (Rag2−/−Il2rg−/−) individually performed experiments with three to five mice per group. (H and I) Absolute numbers (H) and MFI (I) of IL-17RB and IL-33R of Lin−IL-13fm+ ILC2s isolated from different intestinal segments of naïve BALB/c IL-13fm mice. Data represent at least two individually performed experiments with three mice per group. Symbols represent individual data points and bars indicate mean ± SEM. Statistical analysis was performed using unpaired two-tailed Student’s t test (C–G) or one-way ANOVA with Tukey’s post-hoc test (H and I); *P < 0.05, **P < 0.01, ***P < 0.001.
RA signaling inhibition impairs anti-helminth immunity
To study the importance of RA-mediated adaptation on ILC2 functionality in a model that induces a strong intestinal type 2 immune response, we decided to block RA signaling in a helminth infection model. To this end, we applied RAi in Rag2−/− and Rag2−/−Il2rg−/− mice for 3 wk before infecting them with S. ratti, a model for human gastrointestinal nematode infection (Breloer and Abraham, 2017). Control recipients received DMSO injections (see Fig. 8 A for experimental setup). In these experiments, the small intestine was used for the analysis of worm burden, precluding the analysis of ILC2s from the SILP. Thus, we confirmed the effect of RAi on endogenous ILC2s isolated from the colon of infected Rag2−/− mice (Fig. 8, B–D). As observed in uninfected mice (see Fig. 7, E–I), RAi treatment of worm-infected mice resulted in a shift of the ILC2/ILC3 ratio and a change of colonic ILC2 surface marker expression toward a non-intestinal phenotype (e.g., IL-17RBloCD90hiKLRG1loICOShi; Fig. 8, D and E). In line with a functional relevance of this ILC2 phenotypic switch and despite increased ILC2 frequencies, worm burden in the small intestine was significantly higher in RAi-treated Rag2−/− mice than in DMSO controls (Fig. 8 F). In Rag2−/−Il2rg−/− mice, serving as controls for Rag2−/− mice since they lack endogenous ILC population, the adult worm burden remained unchanged after RAi treatment. In addition, mast cell activation after S. ratti infection, as assessed by serum mouse mast cell protease 1 (mMCPT-1), was significantly reduced in the RAi-treated Rag2−/− mice but not in RAi-treated Rag2−/−Il2rg−/− mice compared with their respective controls (Fig. 8 G).
These results are discrepant with a previous report demonstrating that increased ILC2 abundance in conditions of impaired RA signaling improves anti-helminth response in chronic Trichuris muris infection of the cecum (Spencer et al., 2014). However, the observation that ILC2 frequencies and phenotype substantially differ between intestinal segments might explain the distinct sensitivity of these two helminths species to interventions targeting ILC2 adaptation, according to their target segment in the intestine (S. ratti, small intestine versus T. muris, cecum). To formally address this, we analyzed ILC2 abundance and IL-17RB/IL-33R expression in the different intestinal segments of IL-13fm mice (Fig. 8, H and I). Indeed, the abundance of ILC2s in the cecum was much lower than that in small intestinal segments. In addition, IL-17RB expression was substantially lower on cecal than on small intestinal ILC2s, while IL-33R expression was higher. This supports the hypothesis that the expulsion of T. muris from the cecum is less sensitive to RAi-induced perturbations in ILC2 tissue adaptation, e.g., impaired upregulation of IL-17RB, as IL-17RB expression is already low on cecal ILC2s in steady state. Increases in ILC2 abundance under RAi treatment, on the other hand, are likely to improve worm expulsion in T. muris infection as steady-state ILC2 levels are low in the cecum.
In summary, these data confirm that blocking RA signaling can partially reverse the small intestinal ILC2 phenotype, impairing the immune response against S. ratti.
Discussion
Tissue-specific phenotype adaptation is a process widely studied for various immune cell types. The dogma that organ-specific microenvironments dictate phenotype and functionality of immune cells has been long-established for cells of the myeloid lineage (Lavin et al., 2014; Roquilly et al., 2022). In contrast, lymphocyte residency and adaptation in non-lymphoid tissues is an emerging concept that has recently been brought forward by landmark studies on tissue-resident memory T cells and ILCs (Masopust and Soerens, 2019; Meininger et al., 2020). With respect to the ILC2 subset, several recent reports have demonstrated that these tissue-resident lymphocytes exhibit substantial heterogeneity between tissues, and it is presumed that this heterogeneity is imprinted by local niches with their specific microenvironmental cues (Ricardo-Gonzalez et al., 2018; Spits and Mjösberg, 2022).
Using an adoptive transfer model, we show here that murine effector ILC2s originating from non-lymphoid organs could effectively repopulate their niche in various peripheral tissues of alymphoid mice, arguing against an imprinted homing capacity at developmental stages determining the fate of ILC2 localization, as has been previously suggested (Kim et al., 2015; Yang et al., 2016). ILC2s, regardless of their tissue of origin, adapted the phenotype of the tissue they came to reside in, which is in line with previous studies on macrophages and ILC3s that were also capable of adapting to different tissue niches after transfer (Lavin et al., 2014; Nussbaum et al., 2017). In the present study, we observed flexible phenotypic adaptation of the transferred effector ILC2 populations isolated from the kidney, lung, and SILP. We used several strategies in combination to minimize the probability that the effects observed here are due to expansion and differentiation of transferred ILC2 tissue precursor populations: (1) sorting of IL-33R+KLRG1+IL-5hi ILC2s to a high purity (99.9%, see Fig. 3 C) minimizes ILC2 tissue progenitor contamination; (2) scRNAseq of ILC2 populations before transfer shows absence of TCF-7+IL-18R+ ILC2 tissue progenitors in the transferred ILC2 population (see Fig. S3); and (3) RNA velocity analyses of adapting kidney ILC2s in the small intestine supports continuous adaptation of effector ILC2s from the “kidney program” toward the “small intestinal program” (see Fig. 5). However, formal proof of mature effector ILC2 adaptation would require clonal single-cell readouts, which is extremely challenging in innate cells that cannot be tracked using rearranged antigen receptors for endogenous single-cell barcoding. In the future, somatic mitochondrial DNA mutations might offer a possibility for endogenous barcoding of innate lymphoid cell populations to address these questions on a clonal level (Rückert et al., 2022). Moreover, further investigations will be required to assess the ability of mature ILC2s from other tissues (e.g., skin, meninges, and adipose tissue) to replicate these findings.
ILC2s are believed to be distributed to tissues during the perinatal window. Here, they undergo proliferation and expansion and maintain residency in homeostatic conditions, replenishing through tissue precursor expansion (Gasteiger et al., 2015; Huang et al., 2018; Moro et al., 2016; Schneider et al., 2019). However, under inflammatory conditions, such as allergic lung inflammation and prolonged helminth infection, ILC2s from non-lymphoid organs or their precursors from bone marrow have been reported to undergo hematogenous trafficking to affected organs where they contribute to the tissue ILC2 pool (Gasteiger et al., 2015; Huang et al., 2018; Karta et al., 2018; Ricardo-Gonzalez et al., 2020; Stier et al., 2018; Zeis et al., 2020). Collectively, these data suggest that interorgan trafficking of effector ILC2s, as modeled in our adoptive transfer experiments, and our finding of their retained capacity for tissue-specific adaptation is relevant, especially under inflammatory conditions.
Although distinct ILC2 phenotypes have been well recognized in various tissues, the signaling pathways that define tissue-specific ILC2 programs remain poorly understood, and it remains unclear to which extent these tissue signatures can be adapted by ILC2s originating from distant sites. Here, we applied scRNAseq to ex-kidney ILC2s adapting to the small intestinal microenvironment to extract fully adaptable, partially adaptable, and non-adaptable features of the SILP ILC2 transcriptional program, as well as “hardwired” kidney ILC2 features that would not adapt to the new location. Interestingly, we failed to extract any hardwired kidney features from the data, which might indicate that kidney ILC2s exhibit a minimum of tissue-specific adaptation and express a “core ILC2 signature” that is shared with ILC2s in SILP (and presumably other locations). In contrast, several immune-related pathways, such as TNF/NF-κB, ROS, and complement signaling, were identified in the adaptable SILP-specific ILC2 signature and were (partially) acquired by kidney ILC2s in the small intestine.
Using the scRNAseq data set, we then identified RA signaling as one potential mediator of intestine-specific adaptation of effector ILC2s. A prominent feature of successful ILC2 adaptation to the SILP was upregulation of the IL-25 receptor (IL-17RB) and downregulation of the IL-33 receptor (IL-33R), a phenotype that has been observed in previous studies (Huang et al., 2018; Ricardo-Gonzalez et al., 2018; von Moltke et al., 2016). Our in vitro analysis confirmed that RA was sufficient to drive ILC2 upregulation of IL-17RB, as well as changes in other surface markers indicative of the intestine-specific phenotype (CD90.2loKLRG1hiICOSlo). In line with IL-17RB upregulation, RA increased responsiveness of ILC2s to subthreshold IL-25 stimulation, indicating functional importance of this phenotypic switch. In contrast, IL-33R expression was unaltered, and sensitivity of ILC2s to IL-33 stimulation was maintained in the presence of RA in vitro, which was in line with a previous study examining the effect of RA on lung ILC2s (Seehus et al., 2017). Importantly, blocking of RA signaling in vivo after adoptive ILC2 transfer partially inhibited the intestine-specific adaptation of ILC2s, again with the exception of IL-33R downregulation. Our scRNAseq analyses pointed at Notch signaling as another potential pathway involved, a pathway that has been implicated in ILC2 development (Wong et al., 2012) and may also play a role in ILC2 to ILC3 plasticity (Zhang et al., 2017). However, our in vitro validation of this candidate pathway indicated that Notch activation, despite upregulation of IL-17RB and downregulation of IL-33R, acted as a general inhibitor of IL-25– and IL-33–stimulated cytokine production by ILC2s. Although combined activation of the RA and Notch pathways in vitro recapitulated most surface markers defining the intestinal ILC2 phenotype, RA was not able to functionally rescue the general inhibitory effect of Notch ligands on ILC2 cytokine responsiveness (see Fig. S5). Thus, while we established RA signaling as one driving factor for ILC2 adaptation to the intestinal microenvironment, it is evident that a complex combination of signals through various pathways (e.g., RA, Notch, and others) are integrated by SILP ILC2s to shape their full intestine-specific phenotype and function.
As tissue-specific surface receptor expression of ILC2s impacts their functional response in the lung and skin (Ricardo-Gonzalez et al., 2018; Zhang et al., 2022), we hypothesized that RA-mediated ILC2 adaptation to the intestinal environment is required for an optimal type 2 immune response against helminths in which ILC2s play an important role (Meiners et al., 2020; Neill et al., 2010). In line with this hypothesis, we observed that inhibition of RA signaling in Rag2−/− mice leads to reversal of the intestinal phenotype of endogenous ILC2s, which was accompanied by reduced mast cell activation and impaired worm expulsion in acute S. ratti infection. Importantly, this effect depended on the presence of ILCs since it was not observed in ILC-deficient Rag2−/−Il2rg−/− mice. One potential explanation for this finding could be that failed phenotypic ILC2 adaptation impairs their IL-9 production, resulting in suboptimal mast cell activation that is instrumental in S. ratti elimination from the small intestine (Meiners et al., 2020; Reitz et al., 2018). In such a setting, impaired ILC2 activation could be due to a reduced responsiveness of ILC2s to IL-25, caused by a failure to upregulate IL-17RB in the intestine under conditions of impaired RA signaling. This hypothesis is supported by several landmark studies demonstrating that IL-25 is constitutively produced by tuft cells in the small intestine, and a tuft cell/ILC2 feed-forward loop is crucial to mount an efficient anti-helminth immune response (Gerbe et al., 2016; Howitt et al., 2016; von Moltke et al., 2016). In this context, it would also be important to study the effect of IL-25 stimulation on expansion and phenotypic adaptation of ILC2s from the SILP. However, despite extensive efforts, we were unable to retrieve cells from the SILP of mice for several weeks after IL-25 application (200–400 ng per mouse i.p. for four consecutive days), presumably because of intense intestinal inflammation and mucus production induced by IL-25 treatment. Interestingly, detailed literature research revealed a lack of studies on IL-25–expanded murine SILP ILC2s, indicating that other research groups face similar technical challenges.
As previously described (Spencer et al., 2014), RA inhibition also affected ILC2/ILC3 subset distribution, resulting in a significant increase of the ILC3/ILC2 ratio in the small intestine and colon (see Fig. 7, F and G; and Fig. 8, B and C). However, while increased ILC2 abundance in conditions of impaired RA signaling was shown to improve anti-helminth response in chronic T. muris infection of the cecum (Spencer et al., 2014), we observed the opposite effect in acute S. ratti infection. This discrepancy might be explained by considerable differences in these two helminth infection models, regarding the route of infection (S. ratti, subcutaneous versus T. muris, oral), the infection dynamics (S. ratti, acute, self-limiting infection versus T. muris, chronic infection), and probably the most important, the target segment in the intestine (S. ratti, small intestine versus T. muris, cecum). Our data indeed showed low abundance and low IL-17RB expression of ILC2s in the cecum as compared with the SILP (see Fig. 8, H and I). This supports the hypothesis that an increase in ILC2 abundance under RAi treatment (Spencer et al., 2014) is likely to have a positive impact on worm expulsion in cecal T. muris infection, while IL-17RB upregulation is less relevant (as IL-17RB levels are generally low on cecal ILC2s). In S. ratti infection of the small intestine, in contrast, impaired expression of IL-17RB (and potentially other functional markers) under RA signaling inhibition is functionally relevant (as IL-17RB levels are high at steady state), and hence, impaired ILC2 adaptation results in impaired control of helminth infections.
In conclusion, the results presented here attribute an important role to RA signaling in ILC2 adaptation to the small intestine, a mechanism that had not been demonstrated before. However, the study must be considered in the context of several limitations. First, to definitively prove the cell-intrinsic role of RA signaling in intestinal adaptation of ILC2s, transfer experiments with ILC2s deficient in RA signaling would be required. However, owing to the complexity of RA signaling pathway and potential involvement of several RA receptors (e.g., Rara, Rxrb, and Rxrg were expressed in SILP ILC2s), this would require targeting multiple receptors or their downstream signaling components in mice by complex genetic modifications. Second, although demonstrated by ex vivo ILC2 culture experiments, the direct in vivo link between RA-mediated upregulation of IL-17RB expression on SILP ILC2s, increased IL-25 sensitivity, and consequently, alteration of ILC2 in vivo function (e.g., increased cytokine production) has not been shown here. In addition, the predicted RA-induced modulation of IL-17RB downstream signaling (see Fig. 6 K) awaits further exploration to substantiate its mechanistic basis. Finally, we propose a differential response of ILC2s in different intestinal segments (e.g., small versus large intestine) which might offer an explanation for the opposing effects of RA inhibition on defense against different helminth species (Spencer et al., 2014). While the segment-specific ILC2 abundance and expression pattern of IL-17RB and IL-33R (Fig. 8, H and I) suggests differential in vivo functions, a formal proof of this hypothesis is still lacking and warrants further investigations.
Regardless of these limitations, our study highlights that effector ILC2s, after hematogenous trafficking, retain the capacity to adapt to new tissue-specific microenvironments, which may enable them to exert tissue-specific functions, such as promoting control of intestinal helminth infections.
Materials and methods
Animals
C57BL/6 wild type, BALB/c wild type, C57BL/6 Rag2−/−Il2rg−/−, C57BL/6 CD45.1, BALB/c Rag2−/−Il2rg−/−, BALB/c Rag2−/−, and C57BL/6 IL5tdTomato (Red5) mice were bred in the animal facility of the University Medical Centre Hamburg-Eppendorf under specific pathogen–free conditions. BALB/c Il13YFP/Cre were purchased from the Jackson Laboratory and crossed with BALB/c LSL-R26rtdTomato mice to generate IL13fm mice and bred in the animal facility of the University Medical Centre Hamburg-Eppendorf under specific pathogen–free conditions. Adult male and female mice with the appropriate sex- and age-matched controls were used in all experiments. All animal experiments were performed according to national and institutional animal care and ethical guidelines and were approved by the local committees (approval number: N054/2020; Behörde für Gesundheit und Verbraucherschutz, Freie und Hansestadt Hamburg).
In vivo treatments
For ILC2 activation and expansion, mice received injections of 200 or 400 ng recombinant mouse (rm) IL-33 (BioLegend) in 200 μl PBS i.p. on four consecutive days. Controls received 200 μl PBS. ILC2s were isolated at 2–3 wk after the first IL-33 injection. To block RA signaling, a pan-RAi (BMS493; R&D) was dissolved in DMSO (final conc. 29.3 µg/μl). 220 µg RAi/animal (diluted in PBS) was administered i.p. every second day. The same DMSO concentration was administered to control animals.
S. ratti infection, adult worm count, and mast cell activation
The S. ratti cycle was maintained in Wistar rats, and stage 3 larvae were isolated from their charcoal feces culture 1 d before infection using the Baermann technique. Infection of BALB/c Rag2−/−Il2rg−/− and BALB/c Rag2−/− mice was performed by s.c. injection of 2,000 stage 3 larvae in 30 μl PBS into the hind footpad, as described earlier (Eschbach et al., 2010). To count the adult worms on day 6 after infection, the small intestine was cut open longitudinally, washed to remove feces, and tissue was incubated for 3 h at 37°C in tap water while thoroughly shaking every hour for 10 s by hand. After allowing the worms to settle down (∼30 min at room temperature), supernatant was removed and adult worms were counted under a bright-field microscope in a double-blinded fashion. For mast cell activation analysis, blood was collected on day 6 after infection. Serum was analyzed using the MCPT-1 Uncoated ELISA Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Cell isolation
For the isolation of renal leukocytes, mouse kidneys were cut into small pieces and digested in complete medium (RPMI 1640, 10% fetal bovine serum, 1% Hepes, 1% penicillin/streptomycin; all Gibco) supplemented with collagenase D (0.4 mg/ml; Roche) and DNase I (100 µg/ml; Roche) for 45 min at 37°C while rotating on a MACSmix tube rotator (Miltenyi) and subsequently further shred using gentleMACS dissociation (Miltenyi). Lungs were cut into small pieces and digested in a complete medium supplemented with Liberase (0.42 mg/ml; Roche) and DNase I (100 µg/ml; Roche) for 45 min at 37°C while rotating. Afterward, the suspension was mashed through a 70-µm nylon strainer. For the isolation of leukocytes from the SILP, cecum, and colon, Peyer’s patches were removed, the intestine was opened longitudinally, and washed in PBS supplemented with 1% FCS, 5 mM EDTA (Sigma-Aldrich), and 1 mM dithiothreitol (Sigma-Aldrich) for 7 min at 37°C on the shaker. After a second round of shaking (without dithiothreitol), epithelial cells were discarded while the remaining intestinal tissue was minced and incubated in complete medium supplemented with 500 μg/ml collagenase VIII (Sigma-Aldrich) and 20 μg/ml DNAse I (Roche) on a shaker for 30 min at 37°C. Then, cells were mashed through a 70-μm nylon strainer. Further, leukocyte purification was achieved by Percoll gradient centrifugation (37.5%; GE Healthcare). After subsequent erythrocyte lysis with ammonium chloride (kidney and lung only), cell suspensions were filtered through a 50-µm nylon strainer and used for further analyses.
Flow cytometry
Nonspecific antibody binding was prevented by incubation with 10% normal mouse serum (Jackson ImmunoResearch Laboratories). To characterize ILC subsets, cell suspensions of mouse kidneys, lungs, and SILP were stained with fluorochrome-coupled antibodies (all from BioLegend, unless otherwise indicated) against CD45 (30-F11), IL-7Rα (CD127; A7R34), Thy1.2 (CD90.2; 30-H12), and a combination of lineage markers (Lin), including CD3 (145-2C11), CD4 (RM4-5), CD5 (53-7.3), CD8 (53-6.7), CD11b (M1/70), CD11c (HL-3; BD), CD19 (6D5), CD49b (HMα2), TCR-β (H57-597), TCR-γδ (GL3), GR-1 (RB6-8C5), and Ter119 (Ter119). For further characterization of ILC surface marker expression, antibodies against CD25 (PC61.5), IL-33R (RMST2-33; Invitrogen), IL-17RB (CD335; 752101; R&D), KLRG1 (2F1/KLRG1), Ly6A/E (Sca-1; D7), ICOS (C398.4A), NKp46 (29A1.4), and CD117 (c-kit; 2B8) were used. Intranuclear staining (INS), using antibodies against GATA-3 (L50-823; BD), T-bet (4B10), ROR-γt (B2D; BD), Eomes (Dan11mag; eBioscience), and Ki67 (B56; eBioscience), was generally performed with the Transcription Factor Staining Buffer Set (eBioscience) according to the manufacturer’s instruction. INS of leukocytes isolated from reporter mice was performed with a special protocol. After staining of the surface markers, isolated leukocytes were fixed with 3.7% paraformaldehyde (Sigma-Aldrich) for 25 min at room temperature and subsequently resuspended in Perm Wash buffer from the above-mentioned Transcription Factor Staining Set supplemented with INS antibodies and incubated overnight at 4°C. For intracellular cytokine staining, isolated leukocytes were restimulated with phorbol 12-myristate 13-acetate (1 µg/ml; Sigma-Aldrich) and ionomycin (1 µg/ml; Calbiochem) in the presence of brefeldin A (10 µg/ml; Sigma-Aldrich) and β-mercaptoethanol (0.05 mM; Thermo Fisher Scientific) for 2.5 h, stained for surface markers as described above, fixed with fixation/permeabilization buffer from the abovementioned Transcription Factor Staining Buffer Set for 25 min at room temperature, and permeabilized with IGEPAL CA-630 (0.1%; Sigma-Aldrich) for 4 min at room temperature. Subsequently, cells were resuspended in Perm Wash buffer for staining with a combination of fluorochrome-coupled antibodies against IL-13 (eBio13A; eBioscience), IL-5 (TRFK5; BioLegend), amphiregulin (Areg; polyclonal Goat IgG; R&D Systems), and IL-4 (11B11; BD) overnight at 4°C. Dead-cell staining was performed using LIVE/DEAD Fixable Read Dead Stain Kit (Invitrogen) or Zombie Dye (BioLegend). The absolute numbers of CD45+ cells in cell suspension were determined by staining with fluorochrome-coupled anti-CD45 combined with cell-count beads (Countbright; Invitrogen). All samples were acquired on a Becton Dickinson LSRII flow cytometer (BD) using the Diva software and analyzed with the FlowJo Software (Treestar, Inc.). Dimensionality reduction and visualization of high-dimensional flow cytometry data were performed using the UMAP plugin of the FlowJo software v10.
Cell sorting
For sorting of ILC2s from the kidneys, cells were isolated as described above. Instead of using a Percoll gradient, leukocytes were enriched with the CD45 microbeads MACS Cell separation technology (Miltenyi) according to the manufacturer’s instruction. For the isolation of lung ILC2s, the EasySep Mouse ILC2 Enrichment Kit (Stemcell) was used prior to flow cytometry–based cell sorting. ILC2s from the small intestine were isolated as described above. For adoptive cell transfer experiments, in vivo IL-33–expanded ILC2s from kidney, lung, and SILP were defined as CD45+Lin−CD127+KLRG1+ and FACS purified from wild type C57BL/6 CD45.1 mice. Transfer of effector ILC2s was performed using IL-33–activated CD45+Lin−CD127+IL-33R+KLRG1+Red5+ ILC2s sorted from the kidneys of C57BL/6 Red5 mice. For adoptive transfers, 5–10 × 104 ILC2s were administered i.v. into Rag2−/−Il2rg−/− recipient mice. For scRNAseq of ILC2s from the kidneys and SILP from BALB/c IL13fm mice, CD45+Lin−CD127+IL13fm+ ILC2s were sorted and further processed. All samples were sorted on a BD FACSAria Fusion.
Bulk RNAseq
RNA from 2 to 3 × 104 ILC2s was isolated using the RNeasy Plus Micro Kit (QIAGEN) according to the manufacturer’s instructions. The quality of prepared RNA was assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies). After library construction, quality control with Qubit 2.0 fluorometer (Life Technologies) was applied. Sequencing was performed by Novogene using the HiSeq 4000 platform with paired-end of 150 bp (PE 150) sequencing strategy (Illumina). Sequence reads were trimmed to remove sequencing adapters by Trimmomatic 0.38 (Bolger et al., 2014), discarding sequences shorter than 36 bps. Trimmed reads were then aligned to the mouse reference assembly (GRCM38.92) using STAR (v2.5.4b; Dobin et al., 2013). Differential expression analysis was carried out using DESeq2 (Love et al., 2014) and R. Significantly differentially expressed genes (false discovery rate < 0.1 and abs. Log2FC > 1) were used in further analysis. Processed and raw data files of the bulk RNAseq data are available from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE242571.
scRNAseq
Single-cell libraries of sorted cells were generated with the 10X Genomics Chromium Single Cell 5′v1.1 reagents kit according to the manufacturer’s protocol. The libraries were sequenced on Illumina NovaSeq 6000 with 150 base pairs and paired-end configurations. Cell Ranger (version 4.0.0) was used to demultiplex cellular barcodes and map reads to the reference genome (mm10-Ensemble 98). Transcriptomic data were added into a single Seurat object (v.4.0.2). After exclusion of low-quality cells (<500 or >2,500 genes expressed), low expressed genes (<3 genes per cell), and cells with high mitochondrial gene number (>8% of all genes), unsupervised clustering was performed using standard workflow by Seurat. UMAP calculated for the first five principal components was used for dimensionality reduction and plotting. Clusters were assigned according to the sample of origin. DE analyses between cell clusters were performed with the FindAllMarkers function of Seurat. For RNA velocity analysis, a loom-file containing the splicing information was generated using velocyto. Using the python (V3.8.8) package “scVelo” (V0.2.5), the data was preprocessed with the function scv.pp.filter_and_normalize() with standard parameters. Then, first- and second-order moments were computed with scv.pp.moments(). RNA velocity was estimated with scv.tl.velocity in the stochastic mode. Plots were generated using the scv.tl.velocity_graph() and then the scv.pl.velocity_embedding_stream() function. To plot gene expression over pseudotime, pseudotime information was calculated with the scv.tl.velocity_pseudotime function. Then, terminal and initial cell states were determined using the cr.tl.terminal_states() and cr.tl.initial_states() function of the “CellRank” package (V1.5.1). Then, general additive models were fitted on gene expression data and transcripts were plotted using the cr.pl.gene_trends() function. The expression modules (SILP program, kidney program) were plotted using the geom_smooth function of the R package ggplot2 with the pseudotime and score values. For GO term analysis, we compared the gene expression of two clusters with a wilcoxauc() test of the presto library (V1.0.0), arranged the genes according to their area under the curve, and performed gene set enrichment using the fgsea() function of the R library fgsea (V. 1.16.0) with nperm = 1,000. Gene sets were downloaded using the msigdbr function of the msigdbr R library (V7.5.1). RA signaling score was generated by combining GO terms for RA receptor binding (GO:0046965), RA receptor signaling pathway (GO:0048384), and response to RA (GO:0032526), and used for analysis within the AddModuleScore function with default parameters. The Notch signaling score was generated similarly using the Notch signaling pathway (KEGG PATHWAY: hsa04330). Overlay of the Tcf7 and Il18r1 mRNA expression was performed using the Seurat FeaturePlot function using the blend parameter. Processed and raw data files of the scRNAseq data are available from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE242571.
Immunohistochemistry
For staining of tdTomato-positive ILC2s in Red5 mice, 2.5-μm-thick paraffin sections from kidney and SILP were heated at 98°C for 40 min in DAKO antigen retrieval buffer (pH 9; Agilent Technologies). Unspecific binding was blocked with 5% horse serum (Vector Laboratories) and 0.05% triton-X100 in PBS. The sections were then incubated with goat anti-tdTomato (1:2,000, orb182397; Biorbyt) in 5% horse serum overnight at 4°C. Next, the sections were incubated with biotinylated rabbit anti-goat antibody (BA-5000; Vector Laboratories) followed by incubation with anti-rabbit polymer (POLAP-006; Zytomed Systems). Finally, the development and nuclear staining of stained sections were performed with New Fuchsin and Hemalaun, respectively. Tissue slices were evaluated with an Axioskop light microscope (Zeiss) and photographed with an Axiocam HRc camera (Zeiss).
Cell culture and cytokine quantification
FACS-purified kidney ILC2s (defined as CD45+Lin−CD127+KLRG1+) from IL-33–treated mice were cultured (∼10,000 cells/well) in a 96-well U-bottom plate in Iscove’s Modified Dulbecco’s Medium containing 5% FCS, 1% penicillin/streptomycin, and 50 µM 2-mercaptoethanol (all Gibco) in the presence of rmIL-2 (10 ng/ml, BioLegend) with or without all trans RA (1 µM; Enzo Life Sciences). In some experiments, the plate was precoated with the Notch ligand DLL4 (2.5 µg/ml; R&D) for at least 3 h at 37°C. Cells were cultured for 4 d and analyzed for surface marker expression using flow cytometry. In some experiments, cells were stimulated with either rmIL-25 (1 ng/ml; R&D) or rmIL-33 (1 ng/ml) on day 3 and culture supernatants were collected 24 h later and analyzed for IL-5 and IL-13 protein with the LEGENDplex bead-based immunoassay according to the manufacturer’s instructions (BioLegend).
Statistics
The two-tailed parametric t test was used for comparison between two groups. In case of three or more groups, ordinary one-way ANOVA with Tukey’s post hoc test was used. A P value of <0.05 was considered to be statistically significant.
Online supplemental material
Fig. S1 characterizes ILC subsets and ILC2s in different organs. Fig. S2 shows gating and sorting strategies for analysis and purification of ILCs from different organs. Fig. S3 depicts the sorting purity and scRNAseq analysis of kidney ILC2s. Fig. S4 illustrates cluster-defining genes and pathway analysis of scRNAseq data. Fig. S5 shows Notch signaling in intestine-specific adaptation of kidney ILC2s in vitro.
Acknowledgments
We thank the FACS Sorting Core Unit, the Single Cell Sequencing Core Unit, the Bioinformatics Facility, and the Research Animal Facility of the Universitätsklinikum Hamburg-Eppendorf for excellent technical support.
This work was supported by a Deutsche Forschungsgemeinschaft (DFG) Emmy Noether Grant (No. 261149925) to J.-E. Turner, by the DFG Priority Programme 1937 (No. 428069668) to J.-E. Turner and C. Wilhelm, and by the DFG Collaborative Research Centre 1192 (No. 272482170) to J.-E. Turner, U. Panzer, C.F. Krebs, and T.B. Huber.
Author contributions: Conceptualization: J.-E. Turner, N. Shaikh, A.-C. Gnirck, M. Becker, M. Breloer, C. Wilhelm, C.F. Krebs, and U. Panzer. Methodology: J.-E. Turner, N. Shaikh, A.-C. Gnirck, A. Waterhölter, M. Becker, V. Adamiak, W. Hartmann, L. Linnemann, M. Breloer, U. Panzer, C.F. Krebs, R.M. Locksley, C. Wilhelm, and T.B. Huber. Investigation: J.-E. Turner, N. Shaikh, A. Waterhölter, A.-C. Gnirck, V. Adamiak, M. Wunderlich, W. Hartmann, L. Linnemann, and L. Henneken. Visualization: J.-E. Turner, N. Shaikh, A. Waterhölter, A.-C. Gnirck, and M. Becker. Funding acquisition: J.-E. Turner, C.F. Krebs, U. Panzer, and T.B. Huber. Supervision: J.-E. Turner. Writing—original draft: J.-E. Turner, N. Shaikh, A. Waterhölter, A.-C. Gnirck, and M. Becker. Writing—review & editing: J.-E. Turner, N. Shaikh, A. Waterhölter, A.-C. Gnirck, U. Panzer, R.M. Locksley, C.F. Krebs, C. Wilhelm, M. Breloer, and T.B. Huber.
Data availability
All the data in the figures are available in the published article and its online supplemental material. Bulk RNAseq and scRNAseq dataset (raw and processed data) are available from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE242571.
References
- Bando, J.K., Liang H.E., and Locksley R.M.. 2015. Identification and distribution of developing innate lymphoid cells in the fetal mouse intestine. Nat. Immunol. 16:153–160. 10.1038/ni.3057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bar-Ephraim, Y.E., Koning J.J., Burniol Ruiz E., Konijn T., Mourits V.P., Lakeman K.A., Boon L., Bögels M., van Maanen J.P., Den Haan J.M.M., et al. 2019. CD62L is a functional and phenotypic marker for circulating innate lymphoid cell precursors. J. Immunol. 202:171–182. 10.4049/jimmunol.1701153 [DOI] [PubMed] [Google Scholar]
- Bartholin, L., Powers S.E., Melhuish T.A., Lasse S., Weinstein M., and Wotton D.. 2006. TGIF inhibits retinoid signaling. Mol. Cell. Biol. 26:990–1001. 10.1128/MCB.26.3.990-1001.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolger, A.M., Lohse M., and Usadel B.. 2014. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics. 30:2114–2120. 10.1093/bioinformatics/btu170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borowczyk, J., Shutova M., Brembilla N.C., and Boehncke W.H.. 2021. IL-25 (IL-17E) in epithelial immunology and pathophysiology. J. Allergy Clin. Immunol. 148:40–52. 10.1016/j.jaci.2020.12.628 [DOI] [PubMed] [Google Scholar]
- Breloer, M., and Abraham D.. 2017. Strongyloides infection in rodents: Immune response and immune regulation. Parasitology. 144:295–315. 10.1017/S0031182016000111 [DOI] [PubMed] [Google Scholar]
- Dobin, A., Davis C.A., Schlesinger F., Drenkow J., Zaleski C., Jha S., Batut P., Chaisson M., and Gingeras T.R.. 2013. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics. 29:15–21. 10.1093/bioinformatics/bts635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Düster, M., Becker M., Gnirck A.C., Wunderlich M., Panzer U., and Turner J.E.. 2018. T cell-derived IFN-γ downregulates protective group 2 innate lymphoid cells in murine lupus erythematosus. Eur. J. Immunol. 48:1364–1375. 10.1002/eji.201747303 [DOI] [PubMed] [Google Scholar]
- Eschbach, M.L., Klemm U., Kolbaum J., Blankenhaus B., Brattig N., and Breloer M.. 2010. Strongyloides ratti infection induces transient nematode-specific Th2 response and reciprocal suppression of IFN-gamma production in mice. Parasite Immunol. 32:370–383. 10.1111/j.1365-3024.2010.01199.x [DOI] [PubMed] [Google Scholar]
- Gadani, S.P., Smirnov I., Wiltbank A.T., Overall C.C., and Kipnis J.. 2017. Characterization of meningeal type 2 innate lymphocytes and their response to CNS injury. J. Exp. Med. 214:285–296. 10.1084/jem.20161982 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasteiger, G., Fan X., Dikiy S., Lee S.Y., and Rudensky A.Y.. 2015. Tissue residency of innate lymphoid cells in lymphoid and nonlymphoid organs. Science. 350:981–985. 10.1126/science.aac9593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerbe, F., Sidot E., Smyth D.J., Ohmoto M., Matsumoto I., Dardalhon V., Cesses P., Garnier L., Pouzolles M., Brulin B., et al. 2016. Intestinal epithelial tuft cells initiate type 2 mucosal immunity to helminth parasites. Nature. 529:226–230. 10.1038/nature16527 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hams, E., Armstrong M.E., Barlow J.L., Saunders S.P., Schwartz C., Cooke G., Fahy R.J., Crotty T.B., Hirani N., Flynn R.J., et al. 2014. IL-25 and type 2 innate lymphoid cells induce pulmonary fibrosis. Proc. Natl. Acad. Sci. USA. 111:367–372. 10.1073/pnas.1315854111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howitt, M.R., Lavoie S., Michaud M., Blum A.M., Tran S.V., Weinstock J.V., Gallini C.A., Redding K., Margolskee R.F., Osborne L.C., et al. 2016. Tuft cells, taste-chemosensory cells, orchestrate parasite type 2 immunity in the gut. Science. 351:1329–1333. 10.1126/science.aaf1648 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang, Y., Mao K., Chen X., Sun M.A., Kawabe T., Li W., Usher N., Zhu J., Urban J.F. Jr, Paul W.E., and Germain R.N.. 2018. S1P-dependent interorgan trafficking of group 2 innate lymphoid cells supports host defense. Science. 359:114–119. 10.1126/science.aam5809 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwata, M. 2009. Retinoic acid production by intestinal dendritic cells and its role in T-cell trafficking. Semin. Immunol. 21:8–13. 10.1016/j.smim.2008.09.002 [DOI] [PubMed] [Google Scholar]
- Jeffrey, K.L., Camps M., Rommel C., and Mackay C.R.. 2007. Targeting dual-specificity phosphatases: Manipulating MAP kinase signalling and immune responses. Nat. Rev. Drug Discov. 6:391–403. 10.1038/nrd2289 [DOI] [PubMed] [Google Scholar]
- Karagiannis, F., and Wilhelm C.. 2018. Innate lymphoid cells-key immune integrators of overall body homeostasis. Semin. Immunopathol. 40:319–330. 10.1007/s00281-018-0684-y [DOI] [PubMed] [Google Scholar]
- Karta, M.R., Rosenthal P.S., Beppu A., Vuong C.Y., Miller M., Das S., Kurten R.C., Doherty T.A., and Broide D.H.. 2018. β2 integrins rather than β1 integrins mediate Alternaria-induced group 2 innate lymphoid cell trafficking to the lung. J. Allergy Clin. Immunol. 141:329–338.e12. 10.1016/j.jaci.2017.03.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim, M.H., Taparowsky E.J., and Kim C.H.. 2015. Retinoic acid differentially regulates the migration of innate lymphoid cell subsets to the Gut. Immunity. 43:107–119. 10.1016/j.immuni.2015.06.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klose, C.S.N., and Artis D.. 2016. Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat. Immunol. 17:765–774. 10.1038/ni.3489 [DOI] [PubMed] [Google Scholar]
- La Manno, G., Soldatov R., Zeisel A., Braun E., Hochgerner H., Petukhov V., Lidschreiber K., Kastriti M.E., Lönnerberg P., Furlan A., et al. 2018. RNA velocity of single cells. Nature. 560:494–498. 10.1038/s41586-018-0414-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lavin, Y., Winter D., Blecher-Gonen R., David E., Keren-Shaul H., Merad M., Jung S., and Amit I.. 2014. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell. 159:1312–1326. 10.1016/j.cell.2014.11.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim, A.I., Li Y., Lopez-Lastra S., Stadhouders R., Paul F., Casrouge A., Serafini N., Puel A., Bustamante J., Surace L., et al. 2017. Systemic human ILC precursors provide a substrate for tissue ILC differentiation. Cell. 168:1086–1100.e10. 10.1016/j.cell.2017.02.021 [DOI] [PubMed] [Google Scholar]
- Love, M.I., Huber W., and Anders S.. 2014. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15:550. 10.1186/s13059-014-0550-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu, T.C., Wang Z., Feng X., Chuang P., Fang W., Chen Y., Neves S., Maayan A., Xiong H., Liu Y., et al. 2008. Retinoic acid utilizes CREB and USF1 in a transcriptional feed-forward loop in order to stimulate MKP1 expression in human immunodeficiency virus-infected podocytes. Mol. Cell. Biol. 28:5785–5794. 10.1128/MCB.00245-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masopust, D., and Soerens A.G.. 2019. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37:521–546. 10.1146/annurev-immunol-042617-053214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McHedlidze, T., Waldner M., Zopf S., Walker J., Rankin A.L., Schuchmann M., Voehringer D., McKenzie A.N.J., Neurath M.F., Pflanz S., and Wirtz S.. 2013. Interleukin-33-dependent innate lymphoid cells mediate hepatic fibrosis. Immunity. 39:357–371. 10.1016/j.immuni.2013.07.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meiners, J., Reitz M., Rüdiger N., Turner J.E., Heepmann L., Rudolf L., Hartmann W., McSorley H.J., and Breloer M.. 2020. IL-33 facilitates rapid expulsion of the parasitic nematode Strongyloides ratti from the intestine via ILC2- and IL-9-driven mast cell activation. PLoS Pathog. 16:e1009121. 10.1371/journal.ppat.1009121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meininger, I., Carrasco A., Rao A., Soini T., Kokkinou E., and Mjösberg J.. 2020. Tissue-specific features of innate lymphoid cells. Trends Immunol. 41:902–917. 10.1016/j.it.2020.08.009 [DOI] [PubMed] [Google Scholar]
- Monticelli, L.A., Osborne L.C., Noti M., Tran S.V., Zaiss D.M.W., and Artis D.. 2015. IL-33 promotes an innate immune pathway of intestinal tissue protection dependent on amphiregulin-EGFR interactions. Proc. Natl. Acad. Sci. USA. 112:10762–10767. 10.1073/pnas.1509070112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monticelli, L.A., Sonnenberg G.F., Abt M.C., Alenghat T., Ziegler C.G.K., Doering T.A., Angelosanto J.M., Laidlaw B.J., Yang C.Y., Sathaliyawala T., et al. 2011. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12:1045–1054. 10.1038/ni.2131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moro, K., Kabata H., Tanabe M., Koga S., Takeno N., Mochizuki M., Fukunaga K., Asano K., Betsuyaku T., and Koyasu S.. 2016. Interferon and IL-27 antagonize the function of group 2 innate lymphoid cells and type 2 innate immune responses. Nat. Immunol. 17:76–86. 10.1038/ni.3309 [DOI] [PubMed] [Google Scholar]
- Neill, D.R., Wong S.H., Bellosi A., Flynn R.J., Daly M., Langford T.K.A., Bucks C., Kane C.M., Fallon P.G., Pannell R., et al. 2010. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature. 464:1367–1370. 10.1038/nature08900 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nussbaum, K., Burkhard S.H., Ohs I., Mair F., Klose C.S.N., Arnold S.J., Diefenbach A., Tugues S., and Becher B.. 2017. Tissue microenvironment dictates the fate and tumor-suppressive function of type 3 ILCs. J. Exp. Med. 214:2331–2347. 10.1084/jem.20162031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penny, H.A., Hodge S.H., and Hepworth M.R.. 2018. Orchestration of intestinal homeostasis and tolerance by group 3 innate lymphoid cells. Semin. Immunopathol. 40:357–370. 10.1007/s00281-018-0687-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitz, M., Hartmann W., Rüdiger N., Orinska Z., Brunn M.L., and Breloer M.. 2018. Interleukin-9 promotes early mast cell-mediated expulsion of Strongyloides ratti but is dispensable for generation of protective memory. Sci. Rep. 8:8636. 10.1038/s41598-018-26907-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ricardo-Gonzalez, R.R., Schneider C., Liao C., Lee J., Liang H.E., and Locksley R.M.. 2020. Tissue-specific pathways extrude activated ILC2s to disseminate type 2 immunity. J. Exp. Med. 217:e20191172. 10.1084/jem.20191172 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ricardo-Gonzalez, R.R., Van Dyken S.J., Schneider C., Lee J., Nussbaum J.C., Liang H.E., Vaka D., Eckalbar W.L., Molofsky A.B., Erle D.J., and Locksley R.M.. 2018. Tissue signals imprint ILC2 identity with anticipatory function. Nat. Immunol. 19:1093–1099. 10.1038/s41590-018-0201-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riedel, J.H., Becker M., Kopp K., Düster M., Brix S.R., Meyer-Schwesinger C., Kluth L.A., Gnirck A.C., Attar M., Krohn S., et al. 2017. IL-33-mediated expansion of type 2 innate lymphoid cells protects from progressive glomerulosclerosis. J. Am. Soc. Nephrol. 28:2068–2080. 10.1681/ASN.2016080877 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roquilly, A., Mintern J.D., and Villadangos J.A.. 2022. Spatiotemporal adaptations of macrophage and dendritic cell development and function. Annu. Rev. Immunol. 40:525–557. 10.1146/annurev-immunol-101320-031931 [DOI] [PubMed] [Google Scholar]
- Rückert, T., Lareau C.A., Mashreghi M.F., Ludwig L.S., and Romagnani C.. 2022. Clonal expansion and epigenetic inheritance of long-lasting NK cell memory. Nat. Immunol. 23:1551–1563. 10.1038/s41590-022-01327-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samarut, E., and Rochette-Egly C.. 2012. Nuclear retinoic acid receptors: Conductors of the retinoic acid symphony during development. Mol. Cell. Endocrinol. 348:348–360. 10.1016/j.mce.2011.03.025 [DOI] [PubMed] [Google Scholar]
- Schneider, C., Lee J., Koga S., Ricardo-Gonzalez R.R., Nussbaum J.C., Smith L.K., Villeda S.A., Liang H.E., and Locksley R.M.. 2019. Tissue-resident group 2 innate lymphoid cells differentiate by layered ontogeny and in situ perinatal priming. Immunity. 50:1425–1438.e5. 10.1016/j.immuni.2019.04.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seehus, C.R., Kadavallore A., Torre B.d. l., Yeckes A.R., Wang Y., Tang J., and Kaye J.. 2017. Alternative activation generates IL-10 producing type 2 innate lymphoid cells. Nat. Commun. 8:1900. 10.1038/s41467-017-02023-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simoni, Y., Fehlings M., Kløverpris H.N., McGovern N., Koo S.L., Loh C.Y., Lim S., Kurioka A., Fergusson J.R., Tang C.L., et al. 2017. Human innate lymphoid cell subsets possess tissue-type based heterogeneity in phenotype and frequency. Immunity. 46:148–161. 10.1016/j.immuni.2016.11.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spencer, S.P., Wilhelm C., Yang Q., Hall J.A., Bouladoux N., Boyd A., Nutman T.B., Urban J.F. Jr, Wang J., Ramalingam T.R., et al. 2014. Adaptation of innate lymphoid cells to a micronutrient deficiency promotes type 2 barrier immunity. Science. 343:432–437. 10.1126/science.1247606 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spits, H., and Mjösberg J.. 2022. Heterogeneity of type 2 innate lymphoid cells. Nat. Rev. Immunol. 22:701–712. 10.1038/s41577-022-00704-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stier, M.T., Zhang J., Goleniewska K., Cephus J.Y., Rusznak M., Wu L., Van Kaer L., Zhou B., Newcomb D.C., and Peebles R.S. Jr. 2018. IL-33 promotes the egress of group 2 innate lymphoid cells from the bone marrow. J. Exp. Med. 215:263–281. 10.1084/jem.20170449 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turner, J.E., Morrison P.J., Wilhelm C., Wilson M., Ahlfors H., Renauld J.C., Panzer U., Helmby H., and Stockinger B.. 2013. IL-9-mediated survival of type 2 innate lymphoid cells promotes damage control in helminth-induced lung inflammation. J. Exp. Med. 210:2951–2965. 10.1084/jem.20130071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Moltke, J., Ji M., Liang H.E., and Locksley R.M.. 2016. Tuft-cell-derived IL-25 regulates an intestinal ILC2-epithelial response circuit. Nature. 529:221–225. 10.1038/nature16161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- White, K.A., Yore M.M., Warburton S.L., Vaseva A.V., Rieder E., Freemantle S.J., and Spinella M.J.. 2003. Negative feedback at the level of nuclear receptor coregulation. Self-limitation of retinoid signaling by RIP140. J. Biol. Chem. 278:43889–43892. 10.1074/jbc.C300374200 [DOI] [PubMed] [Google Scholar]
- Wong, S.H., Walker J.A., Jolin H.E., Drynan L.F., Hams E., Camelo A., Barlow J.L., Neill D.R., Panova V., Koch U., et al. 2012. Transcription factor RORα is critical for nuocyte development. Nat. Immunol. 13:229–236. 10.1038/ni.2208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang, J., Hu S., Zhao L., Kaplan D.H., Perdew G.H., and Xiong N.. 2016. Selective programming of CCR10(+) innate lymphoid cells in skin-draining lymph nodes for cutaneous homeostatic regulation. Nat. Immunol. 17:48–56. 10.1038/ni.3312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeis, P., Lian M., Fan X., Herman J.S., Hernandez D.C., Gentek R., Elias S., Symowski C., Knöpper K., Peltokangas N., et al. 2020. In situ maturation and tissue adaptation of type 2 innate lymphoid cell progenitors. Immunity. 53:775–792.e9. 10.1016/j.immuni.2020.09.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, J., Qiu J., Zhou W., Cao J., Hu X., Mi W., Su B., He B., Qiu J., and Shen L.. 2022. Neuropilin-1 mediates lung tissue-specific control of ILC2 function in type 2 immunity. Nat. Immunol. 23:237–250. 10.1038/s41590-021-01097-8 [DOI] [PubMed] [Google Scholar]
- Zhang, K., Xu X., Pasha M.A., Siebel C.W., Costello A., Haczku A., MacNamara K., Liang T., Zhu J., Bhandoola A., et al. 2017. Cutting edge: Notch signaling promotes the plasticity of group-2 innate lymphoid cells. J. Immunol. 198:1798–1803. 10.4049/jimmunol.1601421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhuang, X., Ma J., Xu S., Zhang M., Xu G., and Sun Z.. 2021. All-trans retinoic acid attenuates blue light-induced apoptosis of retinal photoreceptors by upregulating MKP-1 expression. Mol. Neurobiol. 58:4157–4168. 10.1007/s12035-021-02380-3 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All the data in the figures are available in the published article and its online supplemental material. Bulk RNAseq and scRNAseq dataset (raw and processed data) are available from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE242571.