A new coronavirus has been identified as the infectious agent of severe acute respiratory syndrome (SARS). Although SARS was successfully contained by quarantine measures, the reconstructed itinerary of the virus through 30 countries and its high mortality rate illustrate the global threat that this newly emerging disease represents (17). During the expression of the SARS coronavirus (SCoV) genome, two viral cysteine proteases, a papain-like protease (PLpro) and a chymotrypsin-like protease (3CLpro), process the encoded polyprotein precursor to release most of the proteins required for virus replication. PLpro refers to a domain of nonstructural protein 3, whose boundaries are defined by homology to the papain-like fold (7). The PLpro domain can be regarded as the catalytic core behind PLpro-mediated cleavages, even though processing by PLpros has been reported to be modulated by additional amino acid residues outside of these boundaries (15, 18). The SCoV utilizes a single PLpro, whereas most coronaviruses contain two paralogous enzymes, termed PL1pro and PL2pro (14). PLpros in general are not as well characterized as 3CLpros and have not generated as much interest as pharmaceutical targets. However, further structure-to-function annotations might refocus the attention on PLpros. For this purpose, we mined the current Protein Data Bank (PDB) content using the Structure Prediction Meta Server (http://www.bioinfo.pl/meta), which assembles state-of-the-art fold recognition methods and provides a consensus sequence-to-structure hyperscore with the 3D-Jury method (4). The only highly reliable prediction (3D-Jury score > 100) obtained for the SCoV PLpro sequence from K1632 to E1847 was the structure of the catalytic core domain of the herpesvirus-associated ubiquitin-specific protease (HAUSP), also known as USP7 (8).
There is compelling evidence for the relevance of a structural relationship between SCoV PLpro and HAUSP (Fig. 1). First, HAUSP is a cysteine protease with a finger domain inserted between the two subdomains of a papain-like fold (8). This structure mirrors the general architecture proposed for the members of the coronaviral PLpro family, which now includes SCoV PLpro, with a Zn ribbon domain inserted in the middle of a papain-like protease domain (7). Our structural bioinformatics data support a circularly permuted rather than a classical Zn ribbon domain for SCoV PLpro, as was recently reported for HAUSP and related enzymes (12). Second, as a deubiquitinating enzyme, HAUSP recognizes the C-terminal ubiquitin sequence, LRGG, which matches the narrow specificity profile of SCoV PLpro (LXGG) derived from the three PLpro-processing sites of the polyprotein (6, 16). The modeled SCoV PLpro binding site is highly complementary to the LXGG sequence and establishes extensive hydrogen bonding with the substrate main chain. In particular, significant occlusions of the S1 subsite (due to N1649 and L1702) and S2 subsite (due to Y1804 and Y1813) account for the strict specificity for diglycine at substrate positions P1 and P2 (Fig. 1C). The structural signatures for strict specificity are present in human coronavirus (HCoV 229E) PL1pro and PL2pro as well as in mouse hepatitis virus (MHV) PL2pro but not in MHV PL1pro (Fig. 1D). This fact correlates with the available specificity data showing a preference for the large arginine residue at the P2 position in the case of MHV PL1pro (2, 3, 9) and with the observation that the irreversible inhibitor E-64d, which contains a bulky P2 residue (leucine), inhibits MHV PL1pro (5, 11) but not MHV PL2pro (10).
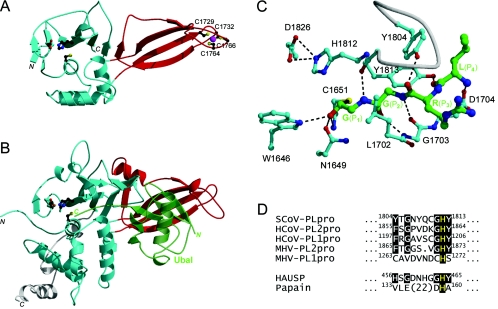
FIG. 1.
Structure-to-function relationships between SCoV PLpro and HAUSP. (A) Three-dimensional model of SCoV PLpro. The protease domain is rendered in cyan, and the circularly permuted Zn ribbon domain is shown in red. The zinc ion is shown as a magenta sphere, and the Zn-chelating cysteines and the catalytic triad are shown in ball-and-stick configurations. The initial homology model was built as a chimera of two templates, the structure of HAUSP complexed with ubiquitin aldehyde (Ubal) (PDB code, 1NBF), with parts of the protease domain outside the binding site based on the structure of the foot-and-mouth disease virus leader protease (PDB code, 1QMY). The model was refined by gradual structural relaxation by using stepwise energy minimization with the AMBER force field. The Ubal C-terminal sequence RLRG-Glycinal, covalently bound at the catalytic C1651 residue as a hemiacetal adduct, was included during the initial stages of structural refinement and replaced with full-length Ubal in the final stage. (B) Crystal structure of the catalytic core domain of HAUSP with bound Ubal. Colors are as described for panel A, except for those for Ubal (semitransparent green) and the C-terminal extension outside the papain-like fold (white). (C) Specific substrate recognition in S1-through-S4 binding site region of SCoV PLpro. The Ubal C-terminal sequence LRG-Glycinal, also corresponding to the SCoV PLpro cleavage motif, is covalently bound at the catalytic C1651 residue, which together with H1812 and D1826 forms the putative catalytic triad. W1646 serves as the putative oxyanion hole residue. The carbon atoms are colored in green for the ligand and in cyan for the binding site. A binding site loop covering the substrate is depicted as a white tube. Hydrogen bonds are shown with dashed lines. (D) Alignment of the S2 subsite signature sequences from selected coronaviral PLpros, HAUSP, and papain. Conserved residues are highlighted on a black background, with the catalytic histidines being shown in yellow. The predicted critical determinants of the S2 subsite specificity correspond to positions 1804 and 1813 of SCoV PLpro.
Structural similarities to HAUSP suggest that, in addition to polyprotein processing activity, SCoV PLpro might possess deubiquitinating activity (including deconjugation of other ubiquitin-like modifiers), as was observed for an adenoviral protease (1). This unexpected activity prediction raises provocative hypotheses regarding the ability of the SARS virus to evade cellular defense mechanisms. For example, it is tempting to speculate that ISG15 deconjugation by PLpro allows the SARS virus to counteract protein ISGylation, an interferon-induced process that may contribute to innate immunity to viral infection (13). The deubiquitination function would greatly impact the value of PLpro as a therapeutic target and provide a framework for the development of antivirals to treat SARS. Strategies for the design of inhibitors of SCoV PLpro must also take into consideration the potentially overlapping specificity of this protease with those of cellular deubiquitinating enzymes. Our finding suggests the performance of follow-up experiments that will increase the understanding of the functional roles of papain-like proteases in the viral life cycles of SCoV and related viruses.
REFERENCES
- 1.Balakirev, M. Y., M. Jaquinod, A. L. Haas, and J. Chroboczek. 2002. Deubiquitinating function of adenovirus proteinase. J. Virol. 76:6323-6331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bonilla, P. J., S. A. Hughes, and S. R. Weiss. 1997. Characterization of a second cleavage site and demonstration of activity in trans by the papain-like proteinase of the murine coronavirus mouse hepatitis virus strain A59. J. Virol. 71:900-909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dong, S., and S. C. Baker. 1994. Determinants of the p28 cleavage site recognized by the first papain-like cysteine proteinase of murine coronavirus. Virology 204:541-549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ginalski, K., A. Elofsson, D. Fischer, and L. Rychlewski. 2003. 3D-Jury: a simple approach to improve protein structure predictions. Bioinformatics 19:1015-1018. [DOI] [PubMed] [Google Scholar]
- 5.Gosert, R., A. Kanjanahaluethai, D. Egger, K. Bienz, and S. C. Baker. 2002. RNA replication of mouse hepatitis virus takes place at double-membrane vesicles. J. Virol. 76:3697-3708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Harcourt, B. H., D. Jukneliene, A. Kanjanahaluethai, J. Bechill, K. M. Severson, C. M. Smith, P. A. Rota, and S. C. Baker. 2004. Identification of severe acute respiratory syndrome coronavirus replicase products and characterization of papain-like protease activity. J. Virol. 78:13600-13612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Herold, J., S. G. Siddell, and A. E. Gorbalenya. 1999. A human RNA viral cysteine proteinase that depends upon a unique Zn2+-binding finger connecting the two domains of a papain-like fold. J. Biol. Chem. 274:14918-14925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hu, M., P. Li, M. Li, W. Li, T. Yao, J. Wu, W. Gu, R. E. Cohen, and Y. Shi. 2002. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111:1041-1054. [DOI] [PubMed] [Google Scholar]
- 9.Hughes, S. A., P. J. Bonilla, and S. R. Weiss. 1995. Identification of the murine coronavirus p28 cleavage site. J. Virol. 69:809-813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kanjanahaluethai, A., D. Jukneliene, and S. C. Baker. 2003. Identification of the murine coronavirus MP1 cleavage site recognized by papain-like proteinase 2. J. Virol. 77:7376-7382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kim, J. C., R. A. Spence, P. F. Currier, X. Lu, and M. R. Denison. 1995. Coronavirus protein processing and RNA synthesis is inhibited by the cysteine proteinase inhibitor E64d. Virology 208:1-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Krishna, S. S., and N. V. Grishin. 2004. The finger domain of the human deubiquitinating enzyme HAUSP is a zinc ribbon. Cell Cycle 3:1046-1049. [PubMed] [Google Scholar]
- 13.Ritchie, K. J., C. S. Hahn, K. I. Kim, M. Yan, D. Rosario, L. Li, J. C. de la Torre, and D. E. Zhang. 2004. Role of ISG15 protease UBP43 (USP18) in innate immunity to viral infection. Nat. Med. 10:1374-1378. [DOI] [PubMed] [Google Scholar]
- 14.Snijder, E. J., P. J. Bredenbeek, J. C. Dobbe, V. Thiel, J. Ziebuhr, L. L. Poon, Y. Guan, M. Rozanov, W. J. Spaan, and A. E. Gorbalenya. 2003. Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage. J. Mol. Biol. 331:991-1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Teng, H., J. D. Piñón, and S. R. Weiss. 1999. Expression of murine coronavirus recombinant papain-like proteinase: efficient cleavage is dependent on the lengths of both the substrate and the proteinase polypeptides. J. Virol. 73:2658-2666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Thiel, V., K. A. Ivanov, A. Putics, T. Hertzig, B. Schelle, S. Bayer, B. Weissbrich, E. J. Snijder, H. Rabenau, H. W. Doerr, A. E. Gorbalenya, and J. Ziebuhr. 2003. Mechanisms and enzymes involved in SARS coronavirus genome expression. J. Gen. Virol. 84:2305-2315. [DOI] [PubMed] [Google Scholar]
- 17.World Health Organization. The World Health Report 2003—shaping the future. [Online.] http://www.who.int/whr/2003.
- 18.Ziebuhr, J., V. Thiel, and A. E. Gorbalenya. 2001. The autocatalytic release of a putative RNA virus transcription factor from its polyprotein precursor involves two paralogous papain-like proteases that cleave the same peptide bond. J. Biol. Chem. 276:33220-33232. [DOI] [PMC free article] [PubMed] [Google Scholar]