Fig. 1.
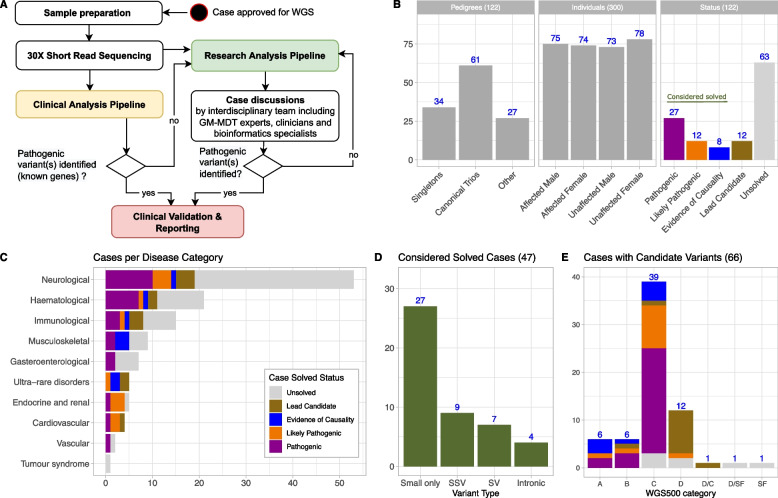
Overview of OxClinWGS study: workflow, clinical cases and results. A Case selection and referral was mainly done by the Oxford Genomic Medicine Multidisciplinary Team (GM-MDT); a detailed description of this process is provided elsewhere [37]. Selected samples were whole-genome sequenced and analysed by a clinical (yellow) as well as a research pipeline (green). Identified pathogenic candidate variants were validated and reported back to referring clinicians. Unsolved cases were iteratively investigated by a research pipeline incorporating the latest methods for in silico analysis of WGS data. Resulting novel disease candidate variants were regularly discussed by an interdisciplinary expert team and either rejected or forwarded to (functional) validation. B Core statistics of considered pedigrees (n = 122) and individuals (n = 300). Note that some families had more than one affected individual. Criteria for the shown classification of pathogenicity are discussed in the main text. C Considered disease categories, coloured by case status. D Variant types for considered solved cases (including pathogenic, likely pathogenic and cases with evidence of causality, see main text). Small only: all causative variants are SNVs or small INDELs, SSV: at least one causative variant is a splice site variant, SV: at least one causative variant is a large structural variant, Intronic: at least one causative variant is (deep) intronic. E Classification of cases using a previously introduced schema published in [23]: A: variant in novel gene for phenotype with additional (genetic) evidence; B: novel (mechanism) for phenotype; C: known gene for phenotype; D: variant in novel gene for phenotype, further genetic and functional validation studies in progress; SF: secondary finding. Note that two cases have candidate variants in two categories. A detailed description of the categories is provided in Additional file 3: Table S3. Abbreviations: SV, structural variant; SSV, splice site variant