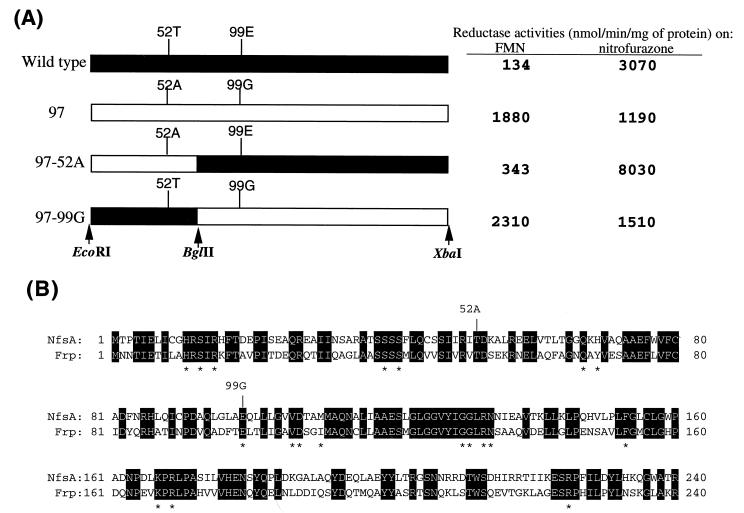
FIG. 1.
Amino acid changes in NfsA-97, a mutant showing a high level of FMN reductase activity. (A) Structures of wild-type NfsA, NfsA-97, and their chimeras (NfsA-97-52A and -99G). Filled boxes, sequences derived from wild-type NfsA; open boxes, sequences derived from NfsA-97. Cleavage sites for restriction endonucleases used for chimera formation are indicated by vertical arrows. Values for NADPH-FMN and nitrofurazone reductase activities in cell extracts are shown on the right. (B) Amino acid sequence homology between NfsA and Frp. Invariant amino acids are shown by white letters in black boxes. The locations of two amino acid substitutions in NfsA-97 are indicated by vertical lines labeled with the mutated amino acids. According to Tanner et al. (10), H-11, S-13, R-15, S-39, Q-67, G-131, K-167, and R-169 are involved in stabilizing the FMN cofactor. R-225 and R-133 may be required for properly steering substrates to the active center (FMN cofactor). Note that R-225 and R-133 have hydrogen bonds to E-99, which is replaced by a glycine residue in NfsA-97. Asterisks show invariant active center amino acids between Frp and NfsA (see Fig. 2).