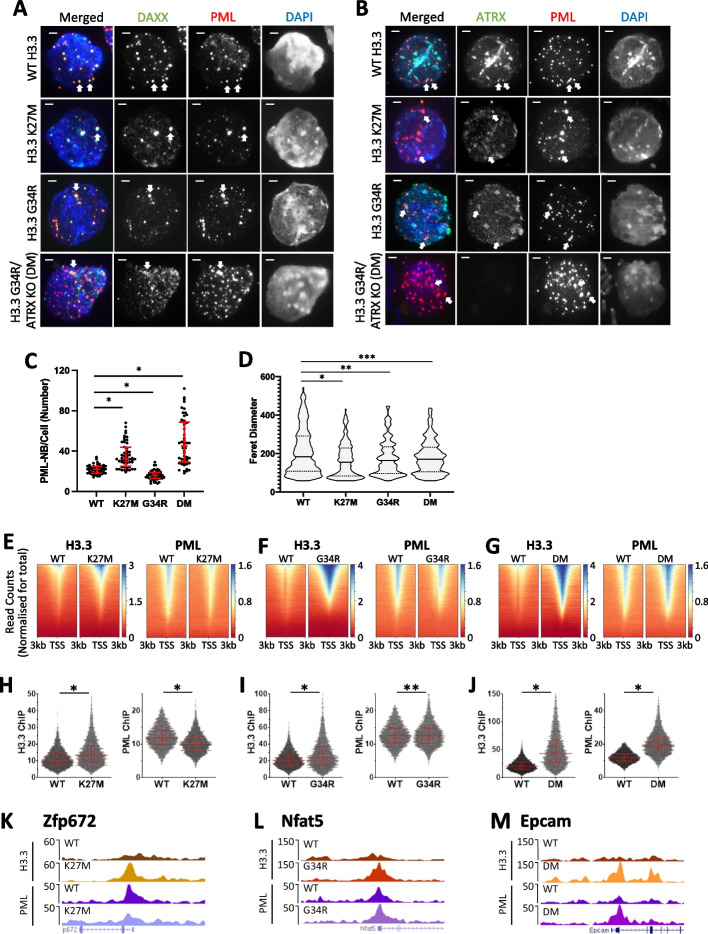
Fig. 2.
PML and H3.3 profiles in mouse ES cells with single-copy H3.3 (K27M/G34R) point mutations, and ATRX KO mutations. Immunofluorescence of A PML (red) and DAXX (green) and B PML (red) and ATRX (green), in WT, H3.3 K27M, H3.3 G34R, and H3.3 G34R/ATRX KO (double mutant; DM) mouse ES cell lines. Scale bar 2 µM. C Quantitation of PML-NBs per cell. Fifty nuclei (n = 50) were counted from three independent immunofluorescence experiments per cell line. Dots represent counts per cell (WT; H3.3 K27M; H3.3 G34R; DM). P-values were calculated using two-tailed Student’s t test (*P < 0.0001). D dSTORM quantitation of Feret diameter (size) of PML-NBs dies in WT (n = 292), H3.3 K27M (n = 200), H3.3 G34R (n = 276), and H3.3 G34R/ATRX KO (DM) (n = 289) cells. 25th, median, and 75th percentiles are shown. P-values were calculated using a two-tailed Mann–Whitney U test (*P < 0.0005, **P < 0.01, **P < 0.05). E–G ChIP-seq read count density heatmaps (normalized for total read counts) of H3.3 (left panel) and PML (right panel) ChIP-seq at transcriptional start sites (TSS) ± 3 kb, sorted by H3.3 enrichment in E H3.3 K27M, F H3.3 G34R, and G DM mutant cell lines, compared to WT cells. H,I H3.3 (left panel) and PML (right panel) ChIP-seq read counts at H3K4me3 enriched promoters (TSS ± 1 kb) in WT and H H3.3 K27M, I H3.3 G34R, and J DM cell lines. P-values were calculated using two-tailed Student’s t test (*P < 0.0001, **P < 0.0005). 25th, median, and 75th percentiles are shown. K–M Representative UCSC genome browser profiles of H3.3 and PML ChIP-seq in WT, K H3.3 K27M (Zfp672; mm9; chr11:58,105,009 – 58,165,027), L H3.3 G34R (Nfat5; mm9; chr8:109,772,915 – 109,862,824), and M DM cell lines (Epcam1; mm9; chr17:87,986,869–88,086,768)