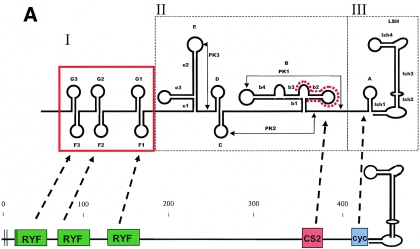
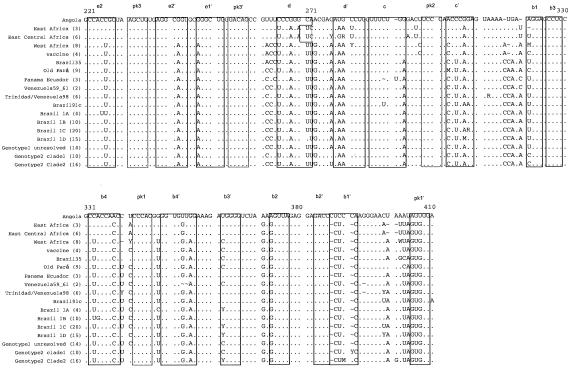
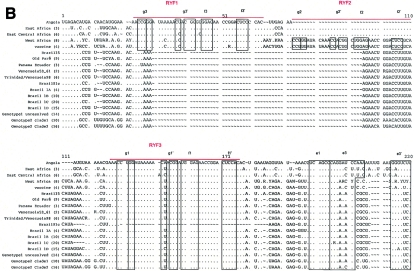
FIG. 1.
(A) Schematic diagram highlighting conserved features of the prototype YFV 3′ NCR (modified from reference 28). RYF, repeated dual F-G hairpins; CS2, ∼24 nt comprising part of dumbbell B; cyc, conserved cyclization domain; LSH, long stable hairpin. (B) Consensus alignment of YFV 3′ NCR sequences (domains I and II). The numbers of isolates used to build the consensus sequences are provided in parentheses (details in Table 1). Nucleotides identical to the reference Angola71 isolate are indicated by dots. Alignment was optimized by the insertion of gaps (∼). Ambiguity codes are as follows: M = A or C; R = A or G; W = A or T; S = C or G; Y = C or T; K = G or T; V = A, C,or G; H = A, C, or T; D = A, G, or T; B = C, G, or T. Regions involved in basepairing are boxed and labeled to correspond with stem structures as indicated in the schematic in panel A.