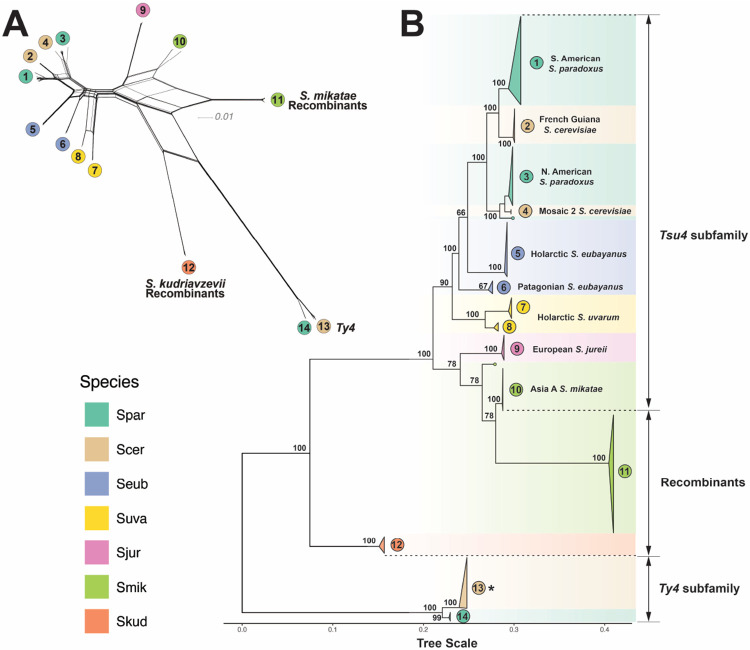
Figure 3: Phylogenetic network and tree of FLEs from the Ty4 family in Saccharomyces.
(A) Phylogenetic network for internal coding regions of Ty4/Tsu4 FLEs based on the NeighborNet algorithm. To simplify visualization, this network only includes Ty4 subfamily FLEs from WGAs reported in Yue et al. (2017). Lineages in the network are labeled according to monophyletic groups identified in panel B. (B) Midpoint rooted ML phylogeny of internal coding regions from Ty4/Tsu4 FLEs. Bootstrap support based on 100 replicates is shown for major nodes. The scale bar for branch lengths is in units of substitutions per site. All monophyletic groups are collapsed as triangles. Two singleton Tsu4 elements (f267 from Hawaiian S. paradoxus strain UWOPS91-917.1 and f256 from S. mikatae strain NBRC 10994) are denoted as dots at tips. Triangles, tip dots, and ranges are colored for each species. Vertical heights of triangles are proportional to the number of taxa. Horizontal widths of triangles are equal to the maximum branch length within the clade. Note that the monophyletic clade for the Ty4 subfamily from S. cerevisiae (annotated with an asterisk) is re-scaled to 5% of the real sample size both horizontally and vertically, due to the large number of Ty4 sequences (n=244) in S. cerevisiae genomes.