Abstract
A growing body of evidence exists supporting the role that genetic variation plays in athletic performance and injury. This study sought to identify genetic variants associated with performance and lower limb musculoskeletal injury in a high-level athletic cohort. A total of 126 Estonian National Team members (Olympic athletes and participants of International Championships) (104 males, 82.5%) underwent a genome-wide association analysis between 2017 and 2018, to identify single-nucleotide polymorphisms (SNPs) associated with performance and/or injury. The athletic cohort was stratified within each sport based on performance and whether they were a medalist (n = 29) or not (n = 97), whether they sustained an injury (n = 47) or not (n = 79), and the type of injury (patella tendinopathy n = 22, Achilles tendinopathy n = 17, hamstring injury n = 3, anterior cruciate ligament rupture n = 6). Three SNPs demonstrated strong genome-wide association with athletic performance (podium/medalist versus not), including DSG1 (rs10502567, OR 14.3) and DSG4 (rs73410248, OR 17.4), while 76 SNPs demonstrated suggestive significance. Overall, 37 SNPs gave genome-wide suggestive association with any type of injury, including PAPPA2 (rs11580456, OR 13.8) and MAS1 (rs220735, rs170219, OR 3.1) which demonstrated positive signal with multiple SNPs. Several genes demonstrated positive association for the specific injury types, including COL22A1 (rs3924862) and PLXNA2 (rs11799530), as well as PAPPA2 (rs11580456), DOK5 (rs73142922), GNG12 (rs28435277), and DAP (rs267959, rs2930047, rs1080440, rs267939). The current study identified genetic variants associated with high-level athletic performance and musculoskeletal injury. Further work is required to permit integration of this and future knowledge into individualized training practices, as well as injury mitigation and rehabilitation programs.
Keywords: Genome-wide association, genetics, DNA, lower limb musculoskeletal injury
Impact Statement
This study identified genetic variants in a high-level athletic cohort that were associated with athletic performance and commonly encountered lower limb soft tissue musculoskeletal injuries. Single-nucleotide polymorphisms (SNPs) including DSG1 and DSG4 demonstrated strong genome-wide association with athletic performance. While several genes demonstrated positive association for specific injury types, including COL22A1, PLXNA2, PAPPA2, DOK5, GNG12, and DAP, numerous SNPs gave genome-wide suggestive association with any type of injury, including PAPPA2 and MAS1. Alongside further research, the current study provides ongoing guidance toward individualized training practices, as well as injury mitigation and rehabilitation programs, in the presence of the identified variants.
Introduction
Acute and chronic musculoskeletal injuries are commonly encountered in a sporting environment. While an individual’s susceptibility to injury is multifactorial, genetic variation may play a role in the etiology of these injuries, further contributing to the interindividual variation in the underlying structure and associated functional properties of musculoskeletal soft tissues (such as tendons and ligaments), combined with their response mechanical loading. 1 For this reason, single-nucleotide polymorphisms (SNPs) have been identified and studied as potential risk factors for these commonly encountered soft tissue injuries. While several reviews have provided support to the association between genetic variation and an increased susceptibility to various soft tissue injuries,1–7 variation in genes responsible for coding the structural components of soft tissues have been linked with (though not limited to) anterior cruciate ligament (ACL) rupture,8–18 Achilles pathology (tendinopathy and/or rupture),9,19–25 and rotator cuff disease.26–28
The main structural component of tendons and ligaments is type I collagen. Therefore, with respect to specific gene mutations and their association with injury risk, genes more widely studied include those encoding the basic structural components of tendons, ligaments, and other connective tissue structures. 1 Specific to tendon and ligament injuries, a review by Vaughn et al. 7 reported that genetic variation with the strongest evidence of association with tendon injury was that involving type V collagen A1, tenascin-C, matrix metalloproteinase-3, and estrogen-related receptor beta, with the most convincing genetic association between tendon injury and COL5A1. The majority of studies that were included in this review were linked with Achilles pathology. A recent systematic review undertaken by Longo et al. 3 investigated the genetic basis of rotator cuff disease, identifying an association between rotator cuff tears and variation in several genes including DEFB1, FGFR1, FGFR3, ESRRB, FGF10, MMP-1, TNC, FCRL3, SASH1, SAP30BP, and rs71404070 located next to cadherin 8. While several studies as outlined above have reported an association between gene variation and ACL rupture risk, a recent review reported a relative lack of association between ACL injury risk and genetic variation. 29 However, it was also suggested that further research with larger samples, phenotype homogeneity, and less study bias was required. 29
The current study aimed to explore the whole-genome scale genetic variations within a high-level athletic population and ascertain whether any of these polymorphisms were associated with performance (being a medalist or not) and/or musculoskeletal injury.
Materials and methods
Athletes
This study followed a case-control genetic association study approach (Level of evidence III). The athletic cohort being investigated consisted of Estonian National Team members (Olympic athletes and participants of International Championships) during the period 2017 to 2018 who agreed to participate in the study. Over this period, a total of 176 athletes were identified, of which 3 athletes did not agree to participate and a further 47 athletes were excluded from the study due to incomplete data (mainly blood samples for genotyping). Therefore, 126 athletes (104 males, 82.5%) were included that underwent genotyping as outlined below. The cohort was subsequently stratified within each sport by whether they reached the podium or not in the Olympic Games, or the World or European Championships (medalist = 29, no medal = 97). Subsequently, athletes were further categorized by whether they then subsequently sustained a lower limb musculoskeletal injury or not (injury = 47, no injury = 79) following testing, while the types of injuries were also reported which included patella tendinopathy (PT = 22), Achilles tendinopathy (AT = 16), hamstring injury (HS = 3), and anterior cruciate ligament rupture (ACL = 6). An overview of the athletic cohort stratified into sport, whether they were a Medalist (or not) within that sport, whether they sustained an injury, and the type of injury, is outlined in Table 1. The study was approved by the relevant Human Research Ethics Committees (HRECs).
Table 1.
A synopsis of the athletic cohort that underwent genotyping, stratified into sport, whether they were a medalist (or not) within that sport, whether they sustained an injury, and the type of injury.
| Sport, n | Medalist (or no medal), n | Injury, n | Injury type(s) | Sport, n | Medalist (or no medal), n | Injury, n | Injury type(s) |
|---|---|---|---|---|---|---|---|
| Basketball, n = 10 | Medalist, n = 0 | Injury, n = 0 | Boxing, n = 3 | Medalist, n = 1 | Injury, n = 0 | ||
| No medal, n = 10 | Injury, n = 6 | PT = 2, AT = 3, ACL = 1 | No medal, n = 2 | Injury, n = 0 | |||
| Canoeing, n = 7 | Medalist, n = 0 | Injury, n = 0 | Cycling, n = 5 | Medalist, n = 0 | Injury, n = 0 | ||
| No medal, n = 7 | Injury, n = 2 | PT = 2 | No medal, n = 5 | Injury, n = 1 | PT = 1 | ||
| Cross country skiing, n = 31 | Medalist, n = 3 | Injury, n = 1 | PT = 1 | Decathlon, n = 1 | Medalist, n = 1 | Injury, n = 1 | PT = 1 |
| No medal, n = 28 | Injury, n = 5 | AT = 3, PT = 1, HS = 1 | No medal, n = 0 | Injury, n = 0 | |||
| Discus, n = 3 | Medalist, n = 3 | Injury, n = 0 | Fencing, n = 2 | Medalist, n = 2 | Injury, n = 1 | ACL = 1 | |
| No medal, n = 0 | Injury, n = 0 | No medal, n = 0 | Injury, n = 0 | ||||
| Figure skating, n = 2 | Medalist, n = 0 | Injury, n = 0 | Javelin, n = 1 | Medalist, n = 1 | Injury, n = 1 | ACL = 1 | |
| No medal, n = 2 | Injury, n = 0 | No medal, n = 0 | Injury, n = 0 | ||||
| Judo, n = 4 | Medalist, n = 3 | Injury, n = 1 | HS = 1 | Rowing, n = 8 | Medalist, n = 7 | Injury, n = 0 | |
| No medal, n = 1 | Injury, n = 1 | ACL = 1 | No medal, n = 2 | Injury, n = 0 | |||
| Running, n = 8 | Medalist, n = 1 | Injury, n = 1 | HS = 1 | Sailing, n = 3 | Medalist, n = 0 | Injury, n = 0 | |
| No medal, n = 7 | Injury, n = 5 | AT = 4, ACL = 1 | No medal, n = 3 | Injury, n = 0 | |||
| Ski jump, n = 6 | Medalist, n = 0 | Injury, n = 0 | Speed skating, n = 1 | Medalist, n = 1 | Injury, n = 1 | AT = 1 | |
| No medal, n = 6 | Injury, n = 1 | PT = 1 | No medal, n = 0 | Injury, n = 0 | |||
| Swimming, n = 5 | Medalist, n = 1 | Injury, n = 1 | PT = 1 | Triathlon, n = 4 | Medalist, n = 1 | Injury, n = 1 | AT = 1 |
| No medal, n = 4 | Injury, n = 0 | No medal, n = 3 | Injury, n = 3 | AT = 3 | |||
| Volleyball, n = 15 | Medalist, n = 0 | Injury, n = 0 | Weightlifting, n = 2 | Medalist, n = 1 | Injury, n = 1 | PT = 1 | |
| No medal, n = 15 | Injury, n = 11 | PT = 10, AT = 1 | No medal, n = 1 | Injury, n = 1 | PT = 1 | ||
| Wrestling, n = 5 | Medalist, n = 3 | Injury, n = 1 | ACL = 1 | ||||
| No medal, n = 2 | Injury, n = 0 |
PT: patella tendinopathy; AT: Achilles tendinopathy; ACL: anterior cruciate ligament; HS: hamstring injury.
DNA extraction and genotyping methodology
DNA was extracted from the blood using Gentra Puregene Blood kit. Genotyping was performed using Affymetrix Axiom Precision Medicine Research Arrays (PMRA) and GeneTitan Multi-Channel Instrument. Genotypes were called using Genotyping Console Software and association analysis was performed using Plink software (https://www.cog-genomics.org/plink/).
Data and statistical analysis
PMRA genotype calls were merged and loaded onto Plink. After filtering for Hardy–Weinberg equilibrium (0.001), for low genotypes (0.01), and for missingness, 736,926 genetic markers were used for the association analysis using chi-square test. First, performance associations were analyzed and sought to stratify athletes by whether they reached the podium in their respective sport or not (i.e. medalists versus non-medalists). Second, the presence of injury and its association with a different genetic profile was analyzed, comparing athletes with injuries versus those without. Manhattan plots were employed to represent the p values of the entire genome-wide association study (GWAS), for each of the aforementioned analyses. Statistically significant SNPs were reported at p < 10−5 (suggestive significance) and p < 10−8 (strong genome-wide significance).
Results
Athletic performance
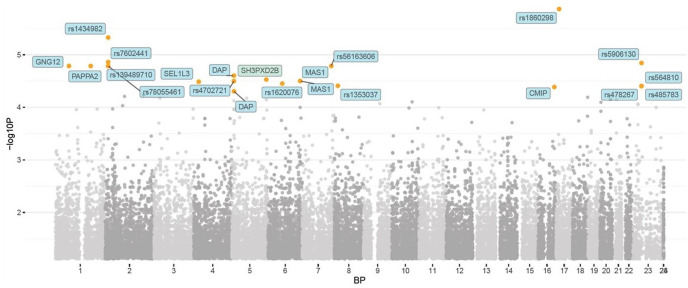
Overall, 3 SNPs were identified that were associated with athletic performance (podium/medalist versus not) with genome-wide significance, while 76 SNPs demonstrated suggestive significance (Figure 1). SNPs associated with the highest performance were from the genes desmoglein 1 (DSG1) (rs10502567, OR 14.3) and desmoglein 4 (DSG4) (rs73410248, OR 17.4) (Appendix A, Supplementary Table). Synaptoporin (SYNPR) (rs58424366, rs2167353, rs938925) and FA complementation group M (FANCM) (rs4900664, rs79199322, rs17115860) genes gave positive signal with multiple SNPs.
Figure 1.
Manhattan plot representing the p values of the genome-wide association in reaching the podium (medalist) or not.
The orange dots represent p < 10−5 while the red dots represent p < 10−8 (i.e. strong genome-wide significance).
Sports injuries
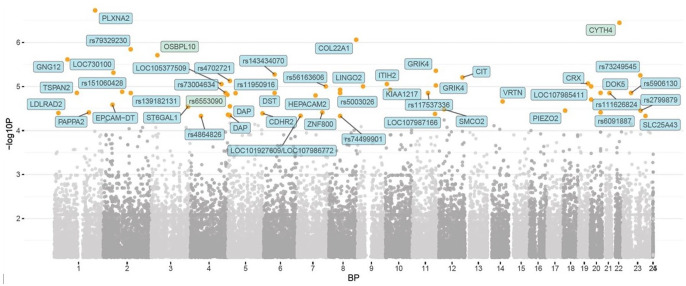
When analyzing sports injuries, 37 SNPs gave suggestively significant genome-wide association (Figure 2). Among the top genes significantly associated with injury was pappalysin 2 (PAPPA2) (rs11580456, OR 13.8), while MAS1 proto-oncogene, G protein-coupled receptor (MAS1) (rs220735, rs170219, OR 3.1) gave positive signal with multiple SNPs (Appendix B, Supplementary Table).
Figure 2.
Manhattan plot representing the p values of the genome-wide association in being injured or not.
The orange dots represent p < 10−5 while the red dots (N/A) represent p < 10−8 (i.e. strong genome-wide significance).
Injury types
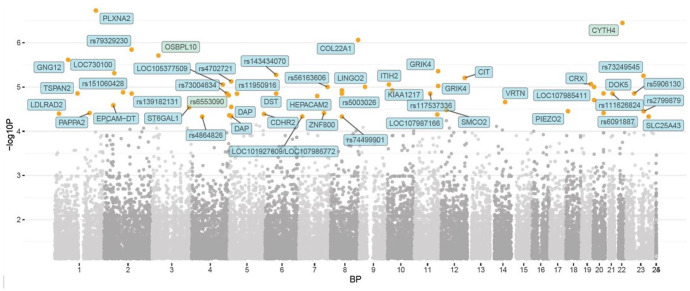
Injuries were divided into four different categories, whereby an attempt was made to identify potential genetic markers for the specific injury types (Figure 3). A range of genes demonstrated positive signals, including collagen type XXII alpha 1 chain (COL22A1) (rs3924862) and plexin A2 (PLXNA2) (rs11799530), as well as PAPPA2 (rs11580456), docking protein 5 (DOK5) (rs73142922), G protein subunit gamma 12 (GNG12) (rs28435277), and death-associated protein (DAP) (rs267959, rs2930047, rs1080440, rs267939) (Appendix C, Supplementary Table). This part of the analysis studied the genetic association with the change in diagnosis or injury type (PT, AT, HS, ACL). As a result, the genetic loci that determine the different types of the injuries were identified, though the genetic variants that are specific for any of these four injuries could not be identified.
Figure 3.
Manhattan plot representing the p values of the genome-wide association in identifying genetic markers for the different injury types.
The orange dots represent p < 10−5 while the red dots (N/A) represent p < 10−8 (i.e. strong genome-wide significance).
Discussion
The current exploratory study identified SNPs that were significantly associated with better performance as a top-level athlete (medalist or not) and experiencing lower limb musculoskeletal injury.
When considering athletic performance, the current study demonstrated genome-wide associations with both DSG1 and DSG4, with a range of SNPs demonstrating suggestive significance (including SYNPR and FANCM). DSG1 and DSG4 are major components of desmosomes that help attach cells to one another. 30 Desmosomes also play a role in the intercellular communications and mutations, with DSG1 and DSG4 related to variable diseases caused by cell adhesion deficiency. 31 However, a range of other genetic markers have previously been linked with elite athletic status, 32 with several reviews commonly reporting two other gene sequence variants associated with performance including the angiotensin-1 converting enzyme insertion/deletion (ACE I/D) and α-actinin-3 (ACTN3) R577X polymorphisms.33 –36 While the ACTN3 genotype has been associated with power-oriented athletic performance, ACE I/I has been associated with endurance performance, with ACE D/D associated with strength/power performance. While the current study stratified elite athletic performance or success based on whether the athlete reached the podium or not, as outlined in a review by Guth and Roth, 34 a particular challenge when investigating the influence of genetic factors is the multifactorial nature of athletic performance and individual characteristics and demands of each sport. Ahmetov et al. 32 identified gene variants specifically associated with both endurance and power/strength athlete status, though they also reported a lack of research identifying genetic markers associated with other sport-related phenotypes such as flexibility, coordination, and athletic temperament. Medalists in the current cohort came from a range of sports, with varied athletic demands spanning endurance, strength/power, speed, and coordination, including cross country skiing, running, swimming, rowing, triathlon, decathlon, discus, javelin, judo, wrestling, boxing, fencing, weightlifting, and speed skating.
Injuries reported in the current study were specific to PT, AT, HS, and ACL. Previous reviews have reported the association between genetic variation and tendon (and/or ligament) injury in general,31,38 while others have reported its association to specific conditions such as Achilles pathology 4 and ACL rupture risk. 29 However, limited (if any) research exists with respect to the risk of HS and PT injury. In the current study, there were no variants that demonstrated strong genome-wide significance with injury, though a range of SNPs demonstrated suggestive significance in association with the injury type (including PAPPA2 and MAS1). PAPPA2 is involved in musculoskeletal diseases and development, and is a metzincin metalloproteinase that cleaves insulin-like growth factor (IGF)-binding protein 5 and regulates the bioavailability of the IGF, also contributing to bone mass and structure formation, with deficiency of PAPPA2 associated with short stature, growth problems developmental hip dysplasia.37 –41 PAPPA2-deficient patients have growth failure with the elevated IGF-1 and IGF-2, while PAPPA2 deficiency is accompanied by impaired glucose metabolism and bone mineral density. 42 MAS1 is a receptor for angiotensin 1-7 and is involved in smooth muscle relaxation, cardioprotection, and regulation of blood pressure.43–45 Angiotensin 1-7 is a product of the ACE2, with ACE2 and angiotensin 1-7 forming a counteracting system against the renin–angiotensin system (RAS) and their primary effect is the lowering of the blood pressure, with MAS1 a receptor for this effect. MAS1 is related to cardiomyocyte signaling and in thermogenesis.43,46
With respect to the analyzed injury categories (AT, PT, HS, ACL), PAPPA2, DOK5, DAP, and GNG12 exhibited positive associations, as did PLXNA2 and COL22A1. COL22A1 is responsible for producing collagen, stabilizing myotendinous junctions and strengthening skeletal muscle attachments during contractions.47,48 These associations demonstrate that these markers are related to the different sensitivity for different injury types, but the current study was unable to identify injury-specific (e.g. ACL-specific) genetic markers due to the limited sample size. However, identification of the loci that show unspecific susceptibility for the injuries provides good rationale and important work for further studies. These genes have a functional impact on the integrity and strength of the skeletomuscular system.
While prior research is lacking with respect to HS injury and PT, DNA sequence variation within the TIMP2, MMP3, MPP7, FBN2, and COL5A1 genes have previously demonstrated an association with AT and/or Achilles tendon pathology.9,19–25 Further to this, Saunders et al. 24 reported significant genetic interactions between variants within genes encoding structural components of the extracellular matrix (TNC, COL27A1) and matrix signaling pathways (IL-6, IL-1b, CASP8) that may collectively contribute to the genetic risk of AT. Sequence variation within COL1A1, COL12A1, COL3A1, COL5A1, and FBN2, across a range of recreational and professional sporting cohorts, have previously demonstrated an association with ACL injury.8 –18 O’Connell et al. 11 reported significant genetic interactions between variants of COL5A1 and COL12A1 and the risk of ACL injury in females, while Rahim et al. 14 reported that regions within VEGFA and KDR may be implicated in the pathophysiology of ACL ruptures. However, Sivertsen et al. 49 could not demonstrate an association between an array of single nucleotide variant in genes (COL1A1, COL3A1, COL5A1, COL12A1) encoding for collagen and the risk of ACL injury in a population of elite female athletes from high-risk team sports in Norway and Finland.
A number of limitations should be acknowledged in the current study. First, while a strength is the high-performance nature of the cohort reviewed, this also limited the sample size and larger, multicenter cohorts that may seek to pool information across several elite-level National teams would provide a wider scope for analysis. Furthermore, larger studies across different populations may be investigated in time to further ascertain variants associated with both performance and lower (and upper) limb musculoskeletal injury risk. Nonetheless, the ORs reported in the current study still demonstrate relatively large effects. The current high-level athletic cohort was heavily biased toward males, though it was heterogeneous given the variety of sports (and therefore varied demands of each sport) included in the cohort. While many studies may investigate injury risk across a broader population, other studies have investigated the risk of a particular injury within a chosen sport, such as ACL injury risk in soccer players8,10 or skiers.15–18 Again, future studies may seek collaboration and multicenter pooling of data to permit gender- and sport-specific analysis. Finally, in the current study, no information was available on the nature of the injury (such as whether the ACL rupture was non-contact or contact), while there was no control group.
Conclusions
The current study identified genetic variants associated with high-level athletic performance, as well as musculoskeletal injury. However, the complexity of human performance and the multifactorial nature of sports, sport performance, and musculoskeletal injury must be appreciated, while despite the current findings and existing literature demonstrating an association between genetic variation and performance/injury, these associations cannot be employed as predictive tools. Further work is required to permit integration of this and future knowledge into talent identification and improved training practices, as well as injury mitigation and rehabilitation programs specific to the individual’s genetic profile.
Supplemental Material
Supplemental material, sj-pdf-1-ebm-10.1177_15353702231198068 for Genome-wide association study identifying variants related to performance and injury in high-performance athletes by Jay R Ebert, Agnes Magi, Eve Unt, Ele Prans, David J Wood and Sulev Koks in Experimental Biology and Medicine
Acknowledgments
The Estonian Research Council and the Estonian Olympic Committee provided funding grants to assist with the GWAS analysis.
Footnotes
Authors’ Contributions: The following authors have conceived and designed the study (JRE, AM, EU, EP, DJW, SK), supervised the conduct of the study and analyzed the data (JRE, AM, EU, EP, DJW, SK), wrote the initial drafts (JRE, DJW, SK), critically revised the manuscript (JRE, AM, EU, EP, DJW, SK), and ensure the accuracy of the data and analysis (JRE, EU, EP, DJW, SK).
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: GWAS analysis was assisted by funding grants provided by the Estonian Research Council (No. IUT 02-7 and No. PRG 435) and the Estonian Olympic Committee (No. SIHT549119).
Ethical Approval: The study was approved by the Research Ethics Committee of the University of Tartu (protocol No. 196/M-30, No. 207/M-9, and No. 224/M-17).
ORCID iDs: Jay R Ebert  https://orcid.org/0000-0001-5843-6243
https://orcid.org/0000-0001-5843-6243
Sulev Koks  https://orcid.org/0000-0001-6087-6643
https://orcid.org/0000-0001-6087-6643
Supplemental Material: Supplemental material for this article is available online.
References
- 1. Collins M, September AV, Posthumus M. Biological variation in musculoskeletal injuries: current knowledge, future research and practical implications. Br J Sports Med 2015;49:1497–503 [DOI] [PubMed] [Google Scholar]
- 2. Collins M, Raleigh SM. Genetic risk factors for musculoskeletal soft tissue injuries. Med Sport Sci 2009;54:136–49 [DOI] [PubMed] [Google Scholar]
- 3. Longo UG, Candela V, Berton A, Salvatore G, Guarnieri A, DeAngelis J, Nazarian A, Denaro V. Genetic basis of rotator cuff injury: a systematic review. BMC Med Genet 2019;20:149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Ribbans WJ, Collins M. Pathology of the tendo Achillis: do our genes contribute? Bone Joint J 2013;95-B:305–13 [DOI] [PubMed] [Google Scholar]
- 5. September AV, Posthumus M, Collins M. Application of genomics in the prevention, treatment and management of Achilles tendinopathy and anterior cruciate ligament ruptures. Recent Pat DNA Gene Seq 2012;6:216–23 [DOI] [PubMed] [Google Scholar]
- 6. September AV, Schwellnus MP, Collins M. Tendon and ligament injuries: the genetic component. Br J Sports Med 2007;41:241–6. Discussion 246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Vaughn NH, Stepanyan H, Gallo RA, Dhawan A. Genetic factors in tendon injury: a systematic review of the literature. Orthop J Sports Med 2017;5:2325967117724416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Ficek K, Cieszczyk P, Kaczmarczyk M, Maciejewska-Karłowska A, Sawczuk M, Cholewinski J, Leonska-Duniec A, Stepien-Slodkowska M, Zarebska A, Stepto NK, Bishop DJ, Eynon N. Gene variants within the COL1A1 gene are associated with reduced anterior cruciate ligament injury in professional soccer players. J Sci Med Sport 2013;16:396–400 [DOI] [PubMed] [Google Scholar]
- 9. Khoury LE, Posthumus M, Collins M, van der Merwe W, Handley C, Cook J, Raleigh SM. ELN and FBN2 gene variants as risk factors for two sports-related musculoskeletal injuries. Int J Sports Med 2015;36:333–7 [DOI] [PubMed] [Google Scholar]
- 10. Lulińska-Kuklik E, Rahim M, Domańska-Senderowska D, Ficek K, Michałowska-Sawczyn M, Moska W, Kaczmarczyk M, Brzeziański M, Brzeziańska-Lasota E, Cięszczyk P, September AV. Interactions between COL5A1 gene and risk of the anterior cruciate ligament rupture. J Hum Kinet 2018;62:65–71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. O’Connell K, Knight H, Ficek K, Leonska-Duniec A, Maciejewska-Karlowska A, Sawczuk M, Stepien-Slodkowska M, O’Cuinneagain D, van der Merwe W, Posthumus M, Cieszczyk P, Collins M. Interactions between collagen gene variants and risk of anterior cruciate ligament rupture. Eur J Sport Sci 2015;15:341–50 [DOI] [PubMed] [Google Scholar]
- 12. Posthumus M, September AV, Keegan M, O’Cuinneagain D, Van der Merwe W, Schwellnus MP, Collins M. Genetic risk factors for anterior cruciate ligament ruptures: COL1A1 gene variant. Br J Sports Med 2009;43:352–6 [DOI] [PubMed] [Google Scholar]
- 13. Posthumus M, September AV, O’Cuinneagain D, van der Merwe W, Schwellnus MP, Collins M. The association between the COL12A1 gene and anterior cruciate ligament ruptures. Br J Sports Med 2010;44:1160–5 [DOI] [PubMed] [Google Scholar]
- 14. Rahim M, Gibbon A, Hobbs H, van der Merwe W, Posthumus M, Collins M, September AV. The association of genes involved in the angiogenesis-associated signaling pathway with risk of anterior cruciate ligament rupture. J Orthop Res 2014;32:1612–8 [DOI] [PubMed] [Google Scholar]
- 15. Stępien-Słodkowska M, Ficek K, Eider J, Leońska-Duniec A, Maciejewska-Karłowska A, Sawczuk M, Zarębska A, Jastrzębski Z, Grenda A, Kotarska K, Cięszczyk P. The +1245g/t polymorphisms in the collagen type I alpha 1 (col1a1) gene in polish skiers with anterior cruciate ligament injury. Biol Sport 2013;30:57–60 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Stepien-Slodkowska M, Ficek K, Kaczmarczyk M, Maciejewska-Karlowska A, Sawczuk M, Leonska-Duniec A, Stepinski M, Zietek P, Krol P, Chudecka M, Cieszczyk P. The variants within the COL5A1 gene are associated with reduced risk of anterior cruciate ligament injury in skiers. J Hum Kinet 2015;45:103–11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Stępień-Słodkowska M, Ficek K, Maciejewska-Karłowska A, Sawczuk M, Ziętek P, Król P, Zmijewski P, Pokrywka A, Cięszczyk P. Overrepresentation of the COL3A1 AA genotype in Polish skiers with anterior cruciate ligament injury. Biol Sport 2015;32:143–7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Stepien-Slodkowska M, Ficek K, Zietek P, Kaczmarczyk M, Lubkowska W, Szark-Eckardt M, Cieszczyk P. Is the combination of COL1A1 gene polymorphisms a marker of injury risk? J Sport Rehabil 2017;26:234–8 [DOI] [PubMed] [Google Scholar]
- 19. El Khoury L, Posthumus M, Collins M, Handley CJ, Cook J, Raleigh SM. Polymorphic variation within the ADAMTS2, ADAMTS14, ADAMTS5, ADAM12 and TIMP2 genes and the risk of Achilles tendon pathology: a genetic association study. J Sci Med Sport 2013;16:493–8 [DOI] [PubMed] [Google Scholar]
- 20. El Khoury L, Ribbans WJ, Raleigh SM. MMP3 and TIMP2 gene variants as predisposing factors for Achilles tendon pathologies: attempted replication study in a British case-control cohort. Meta Gene 2016;9:52–5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Gibbon A, Hobbs H, van der Merwe W, Raleigh SM, Cook J, Handley CJ, Posthumus M, Collins M, September AV. The MMP3 gene in musculoskeletal soft tissue injury risk profiling: a study in two independent sample groups. J Sports Sci 2017;35:655–62 [DOI] [PubMed] [Google Scholar]
- 22. Gibbon A, Raleigh SM, Ribbans WJ, Posthumus M, Collins M, September AV. Functional COL1A1 variants are associated with the risk of acute musculoskeletal soft tissue injuries. J Orthop Res 2020;38:2290–8 [DOI] [PubMed] [Google Scholar]
- 23. Kang X, Tian B, Zhang L, Ge Z, Zhao Y, Zhang Y. Relationship of common variants in MPP7, TIMP2 and CASP8 genes with the risk of chronic achilles tendinopathy. Sci Rep 2019;9:17627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Saunders CJ, van der Merwe L, Cook J, Handley CJ, Collins M, September AV. Extracellular matrix proteins interact with cell-signaling pathways in modifying risk of achilles tendinopathy. J Orthop Res 2015;33:898–903 [DOI] [PubMed] [Google Scholar]
- 25. September AV, Cook J, Handley CJ, van der Merwe L, Schwellnus MP, Collins M. Variants within the COL5A1 gene are associated with Achilles tendinopathy in two populations. Br J Sports Med 2009;43:357–65 [DOI] [PubMed] [Google Scholar]
- 26. Motta R, Amaral MV, Rezende E, Pitta R, Vieira TC, Duarte ME, Vieira AR, Casado PL. Evidence of genetic variations associated with rotator cuff disease. J Shoulder Elbow Surg 2014;23:227–35 [DOI] [PubMed] [Google Scholar]
- 27. Peach CA, Zhang Y, Dunford JE, Brown MA, Carr AJ. Cuff tear arthropathy: evidence of functional variation in pyrophosphate metabolism genes. Clin Orthop Relat Res 2007;462:67–72 [DOI] [PubMed] [Google Scholar]
- 28. Teerlink CC, Cannon-Albright LA, Tashjian RZ. Significant association of full-thickness rotator cuff tears and estrogen-related receptor-beta (ESRRB). J Shoulder Elbow Surg 2015;24:e31–5 [DOI] [PubMed] [Google Scholar]
- 29. Kaynak M, Nijman F, van Meurs J, Reijman M, Meuffels DE. Genetic variants and anterior cruciate ligament rupture: a systematic review. Sports Med 2017;47:1637–50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Wang Y, Amagai M, Minoshima S, Sakai K, Green KJ, Nishikawa T, Shimizu N. The human genes for desmogleins (DSG1 and DSG3) are located in a small region on chromosome 18q12. Genomics 1994;20:492–5 [DOI] [PubMed] [Google Scholar]
- 31. Schaffer JV, Bazzi H, Vitebsky A, Witkiewicz A, Kovich OI, Kamino H, Shapiro LS, Amin SP, Orlow SJ, Christiano AM. Mutations in the desmoglein 4 gene underlie localized autosomal recessive hypotrichosis with monilethrix hairs and congenital scalp erosions. J Invest Dermatol 2006;126:1286–91 [DOI] [PubMed] [Google Scholar]
- 32. Ahmetov II, Egorova ES, Gabdrakhmanova LJ, Fedotovskaya ON. Genes and athletic performance: an update. Med Sport Sci 2016;61:41–54 [DOI] [PubMed] [Google Scholar]
- 33. Eynon N, Hanson ED, Lucia A, Houweling PJ, Garton F, North KN, Bishop DJ. Genes for elite power and sprint performance: ACTN3 leads the way. Sports Med 2013;43:803–17 [DOI] [PubMed] [Google Scholar]
- 34. Guth LM, Roth SM. Genetic influence on athletic performance. Curr Opin Pediatr 2013;25:653–8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Ma F, Yang Y, Li X, Zhou F, Gao C, Li M, Gao L. The association of sport performance with ACE and ACTN3 genetic polymorphisms: a systematic review and meta-analysis. PLoS ONE 2013;8:e54685 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Puthucheary Z, Skipworth JR, Rawal J, Loosemore M, Van Someren K, Montgomery HE. Genetic influences in sport and physical performance. Sports Med 2011;41:845–59 [DOI] [PubMed] [Google Scholar]
- 37. Harsanyi S, Zamborsky R, Krajciova L, Kokavec M, Danisovic L. Genetic study of IL6, GDF5 and PAPPA2 in association with developmental dysplasia of the hip. Genes 2021;12:986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Christians JK, Amiri N, Schipilow JD, Zhang SW, May-Rashke KI. Pappa2 deletion has sex- and age-specific effects on bone in mice. Growth Horm IGF Res 2019;44:6–10 [DOI] [PubMed] [Google Scholar]
- 39. Overgaard MT, Boldt HB, Laursen LS, Sottrup-Jensen L, Conover CA, Oxvig C. Pregnancy-associated plasma protein-A2 (PAPP-A2), a novel insulin-like growth factor-binding protein-5 proteinase. J Biol Chem 2001;276:21849–53 [DOI] [PubMed] [Google Scholar]
- 40. Farr M, Strube J, Geppert HG, Kocourek A, Mahne M, Tschesche H. Pregnancy-associated plasma protein-E (PAPP-E). Biochim Biophys Acta 2000;1493:356–62 [DOI] [PubMed] [Google Scholar]
- 41. Standker L, Wobst P, Mark S, Forssmann WG. Isolation and characterization of circulating 13-kDa C-terminal fragments of human insulin-like growth factor binding protein-5. FEBS Lett 1998;441:281–6 [DOI] [PubMed] [Google Scholar]
- 42. Fujimoto M, Andrew M, Dauber A. Disorders caused by genetic defects associated with GH-dependent genes: PAPPA2 defects. Mol Cell Endocrinol 2020;518:110967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Vargas-Castillo A, Tobon-Cornejo S, Del Valle-Mondragon L, Torre-Villalvazo I, Schcolnik-Cabrera A, Guevara-Cruz M, Pichardo-Ontiveros E, Fuentes-Romero R, Bader M, Alenina N, Vidal-Puig A, Hong E, Torres N, Tovar AR. Angiotensin-(1-7) induces beige fat thermogenesis through the Mas receptor. Metabolism 2020;103:154048. [DOI] [PubMed] [Google Scholar]
- 44. Che Mohd Nassir CMN, Zolkefley MKI, Ramli MD, Norman HH, Abdul Hamid H, Mustapha M. Neuroinflammation and COVID-19 ischemic stroke recovery-evolving evidence for the mediating roles of the ACE2/angiotensin-(1-7)/mas receptor axis and NLRP3 inflammasome. Int J Mol Sci 2022;23:3085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Zhou G, Fan L, Li Z, Li J, Kou X, Xiao M, Gao M, Qu X. G protein-coupled receptor MAS1 induces an inhibitory effect on myocardial infarction-induced myocardial injury. Int J Biol Macromol 2022;207:72–80 [DOI] [PubMed] [Google Scholar]
- 46. Gomes ER, Santos RA, Guatimosim S. Angiotensin-(1-7)-mediated signaling in cardiomyocytes. Int J Hypertens 2012;2012:493129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Charvet B, Guiraud A, Malbouyres M, Zwolanek D, Guillon E, Bretaud S, Monnot C, Schulze J, Bader HL, Allard B, Koch M, Ruggiero F. Knockdown of col22a1 gene in zebrafish induces a muscular dystrophy by disruption of the myotendinous junction. Development 2013;140:4602–13 [DOI] [PubMed] [Google Scholar]
- 48. Koch M, Schulze J, Hansen U, Ashwodt T, Keene DR, Brunken WJ, Burgeson RE, Bruckner P, Bruckner-Tuderman L. A novel marker of tissue junctions, collagen XXII. J Biol Chem 2004;279:22514–21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Sivertsen EA, Haug KBF, Kristianslund EK, Trøseid AS, Parkkari J, Lehtimäki T, Mononen N, Pasanen K, Bahr R. No association between risk of anterior cruciate ligament rupture and selected candidate collagen gene variants in female elite athletes from high-risk team sports. Am J Sports Med 2019;47:52–8 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material, sj-pdf-1-ebm-10.1177_15353702231198068 for Genome-wide association study identifying variants related to performance and injury in high-performance athletes by Jay R Ebert, Agnes Magi, Eve Unt, Ele Prans, David J Wood and Sulev Koks in Experimental Biology and Medicine