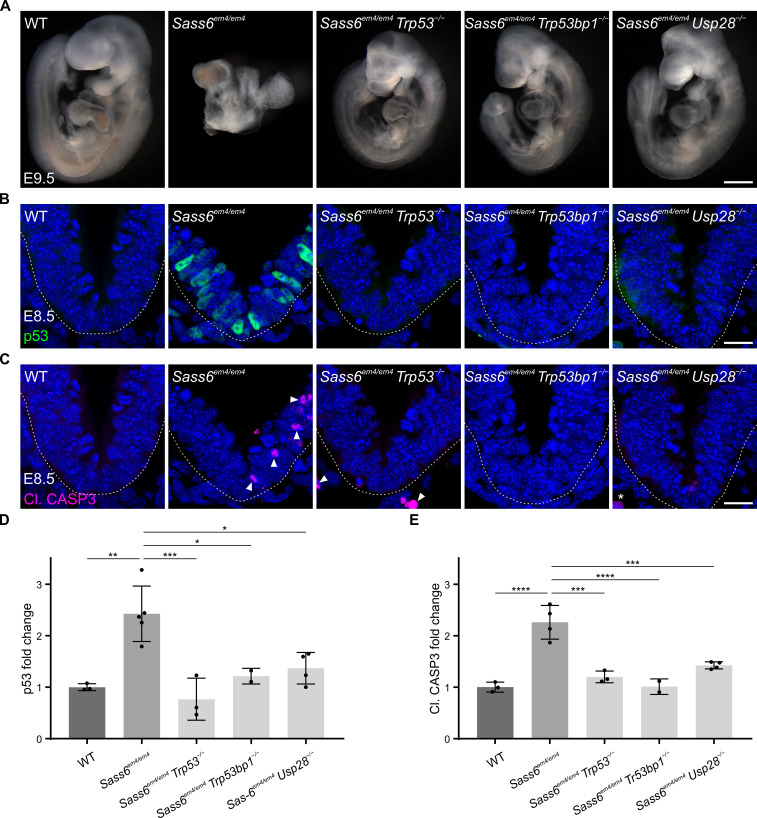
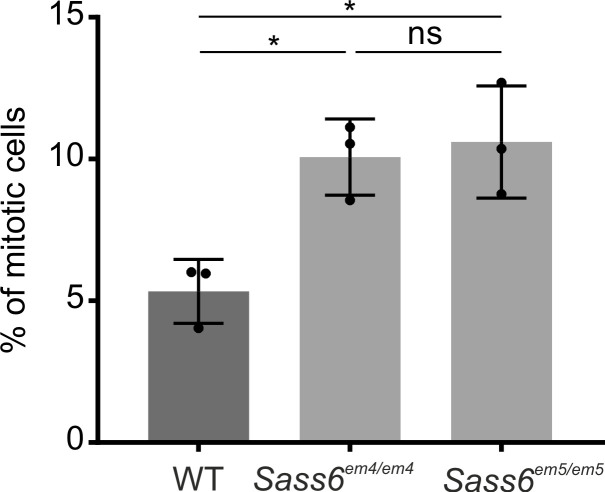
Figure 1. Mutation in mouse Sass6 activates the 53BP1-USP28-p53 mitotic surveillance pathway.
(A) Left-side views of wild-type (WT), Sass6em4/em4, Sass6em4/em4 Trp53−/−, Sass6em4/em4 Trp53bp1−/−, and Sass6em4/em4 Usp28−/− embryos at E9.5. Anterior is up in all images. At least three embryos per genotype showed similar phenotypes. Scale bar = 500 µm. (B) Immunostaining for p53 on transverse sections of WT, Sass6em4/em4, Sass6em4/em4 Trp53−/−, Sass6em4/em4 Trp53bp1−/−, and Sass6em4/em4 Usp28−/− embryos at E8.5. The sections shown encompass the neural plate (top) and mesenchyme (bottom), demarcated by the dashed line. Dorsal is up in all images. Scale bars = 25 µm. (C) Immunostaining for Cleaved-Caspase3 (Cl. CASP3) as mentioned in (B). Arrowheads indicate Cl. CASP3-positive cells, while asterisks mark non-specific staining of blood cells. (D) Quantification of the nuclear p53 in (B). Values were normalized to WT. Error bars represent mean ± SD WT: 1.00 ± 0.06 (n=2582 cells from three embryos); Sass6em4/em4: 2.4 ± 0.5 (n=2372 from four embryos); Sass6em4/em4 Trp53−/−: 0.8 ± 0.3 (n=2379 from three embryos); Sass6em4/em4 Usp28−/−: 1.4 ± 0.3 (n=2775 from four embryos); Sass6em4/em4 Trp53bp1−/−: 1.2 ± 0.1 (n=1840 from two embryos). *p<0.05, **p<0.01, ***p<0.001 (one-way ANOVA with Tukey’s multiple comparisons). (E) Quantification of Cl. CASP3 in (C) as mentioned in (D). WT: 1.00 ± 0.1 (n=2582 cells from three embryos); Sass6em4/em4: 2.3 ± 0.3 (n=2372 from four embryos); Sass6em4/em4 Trp53−/−: 1.2 ± 0.1 (n=2379 from three embryos); Sass6em4/em4 Usp28−/−: 1.4 ± 0.1 (n=2775 from four embryos); Sass6em4/em4 Trp53bp1−/−: 1 ± 0.1 (n=1840 from two embryos). ***p<0.001, ****p<0.0001 (one-way ANOVA with Tukey’s multiple comparisons).