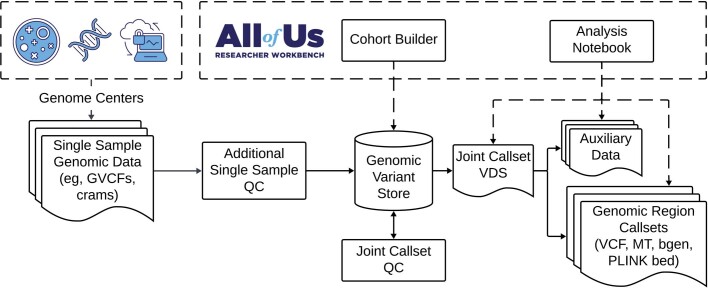
Extended Data Fig. 2. Overview of the Genomic Data Curation Pipeline for WGS samples.
The Data and Research Center (DRC) performs additional single sample quality control (QC) on the data as it arrives from the Genome Centers. The variants from samples that pass this QC are loaded into the Genomic Variant Store (GVS), where we jointly call the variants and apply additional QC. We apply a joint call set QC process, which is stored with the call set. The entire joint call set is rendered as a Hail Variant Dataset (VDS), which can be accessed from the analysis notebooks in the Researcher Workbench. Subsections of the genome are extracted from the VDS and rendered in different formats with all participants. Auxiliary data can also be accessed through the Researcher Workbench. This includes variant functional annotations, joint call set QC results, predicted ancestry, and relatedness. Auxiliary data are derived from GVS (arrow not shown) and the VDS. The Cohort Builder directly queries GVS when researchers request genomic data for subsets of samples. Aligned reads, as cram files, are available in the Researcher Workbench (not shown). The graphics of the dish, gene and computer and the All of Us logo are reproduced with permission of the National Institutes of Health’s All of Us Research Program.