Abstract
Two-pore K+ (TPK) channels are voltage-independent and involved in stress response in plants. Herein, we identified 12 TaTPK genes located on nine chromosomes in the Triticum aestivum genome. The majority of TaTPK genes comprised two exons. Each TaTPK channel comprised four transmembrane (TM) helices, N- and C-terminal ion-channel domains, two EF-hand domains and one 14-3-3 binding site. Additionally, highly conserved ‘GYGD’ motif responsible for K+ ion specificity, was found in between the TMs in both the ion-channel domains. Nine TaTPK channels were predicted to be localised at the plasma membrane, while three were vacuolar. The protein-protein and protein-chemical interactions indicated the coordinated functioning of the TaTPK channels with the other K+ transporters and their possible interaction with the Ca2+-signaling pathway. Expression studies suggested their importance in both vegetative and reproductive tissues development. Significantly modulated expression of various TaTPK genes during heat, drought, combined heat and drought and salt stresses, and after fungal infestation, depicted their function in stress responses. The miRNAs and transcription factors interaction analyses suggested their role in the hormone, light, growth and development-related, and stress-responsive signaling cascades. The current study suggested vital functions of various TaTPK genes, especially in stress response, and would provide an opportunity for their detailed characterization in future studies.
Keywords: Ca2+ signalling, EF-Hand, Expression, Stress response, TPK channel
1. Introduction
Potassium ion (K+) is an important micronutrient as it regulates various physiological phenomenon such as membrane potential, cell elongation, osmoregulation, and turgor generation in plants [1]. K+ is also engaged in pH equilibrium and required for the activation of a diverse group of enzymes [2]. Additionally, it is also an important regulator in conferring tolerance against numerous abiotic and biotic stress conditions [[3], [4], [5], [6]]). In plants, K+ is present in various cellular compartments such as cytosol, mitochondria, vacuole, chloroplast and nucleus, in which cytosol consists of higher concentration (60–160 mM) and therefore, needs regulated homeostasis for appropriate functioning [7,8]. A complex transport system including carrier and channel proteins play a vital role in K+ ions homeostasis. The K+ channels are multimeric transmembrane proteins consisting of pore domains and highly conserved motif GYGD/E. These channels are classified as voltage-gated shaker channels, tandem or two-pore K+ (TPK) channels and K+-inward rectifier (Kir)-like channel (KCO) [[9], [10], [11]].
The TPK channels consist of two ion transporter domains (PF07885) lying at N- and C-terminus, a binding site for 14-3-3 proteins at N-terminus and one or two Ca2+ binding EF-hand domains at C-terminus [12]). They are voltage-independent membrane channels, but depend upon cytosolic pH and Ca2+ for their functioning [13,14]. Moreover, phosphorylation, 14-3-3 proteins, membrane stretching, and osmotic disturbances also affect the functioning of TPK channels [15,16]. The TPK1 channel of Nicotiana tabacum also showed sensitivity to spermine and spermidine [17]. TPK channels have been identified and characterized in various plant species including Arabidopsis thaliana, Oryza sativa, Solanum tuberosum, Cicer arietinum, N. tabacum, Cajanus cajan, Gossypium raimondii, etc. [[17], [18], [19], [20], [21], [22], [23], [24]].
In A. thaliana, a total of five TPK isoforms (AtTPK1-AtTPK5) have been identified [18]. Through the knockout and overexpressing approaches, the role of AtTPK1 has been established in intracellular K+ homeostasis. It also promotes vacuolar K+ release at the time of stomatal closure and reported to be involved in seed germination at later stages [13,25]. Another study suggested the involvement of AtTPK4 in the regulation of membrane voltage of pollen tube and K+ homeostasis [26].
In O. sativa, three TPK isoforms such as OsTPKa, OsTPKb, and OsTPKc are reported. The OsTPKa and OsTPKb are located in the lytic and storage vacuoles, respectively [20]. The transgenic lines with over-expressed OsTPKa and OsTPKb exhibit improved K+ homeostasis and higher tolerance during osmotic and salt stress conditions [27,28]. Similarly, both NtTPK1a and NtTPK1b of N. tabacum are involved in hypoosmotic stress responses [17,19]. However, these channels have not been explored in T. aestivum.
In our current analysis, we have identified and characterized TPK channels in the genome of T. aestivum L. The identified channels were characterised for their chromosomal localization, gene and protein structure, and phylogenetic relationship. The expression patterns of TaTPKs were analysed to predict their functions in tissue developmental stages and stress conditions. In addition, the other functions of TaTPKs were explored by performing miRNAs interaction, transcription factor binding, protein-protein, and protein-chemical interaction studies. This study would provide insight into the various characteristics of TPK channels, and facilitate an avenue for detailed functional characterization in future studies.
2. Materials and methods
2.1. Identification and nomenclature of TaTPK genes
The BLASTp search approach at the Ensembl Plant database was used to scan the T. aestivum genome for the identification of TaTPKs using the known TPK sequences of A. thaliana and O. sativa at e-value e−10 as done in earlier studies [[29], [30], [31]]. The HMMER and Pfam database searches were also carried out to validate the identification of retrieved sequences through domain confirmation [32]. Furthermore, the presence of characteristic domains of the TPK proteins was reconfirmed by the Simpler Modular Architecture Research Tool (SMART) and the NCBI conserved domain database (CDD) searches [33,34]. The homeologous grouping of identified genes was done on the basis of ≥90% similarity and subsequently, the nomenclature was given following the rules specified by international rules for gene symbolization in T. aestivum (http://wheat.pw.usda.gov/ggpages/wgc/98/Intro.htm). The duplicate event analysis has been performed by predicting the ≥80% sequence similarity between TaTPK genes. The existence of identified TaTPK genes at transcript level was confirmed by TSA (transcriptome shotgun assembly) BLAST at the NCBI-TSA database. Some of the incomplete sequences at the Ensembl Plants were also completed using TSA sequences as described in the earlier studies [[35], [36], [37]].
2.2. Multiple sequence alignment and phylogenetic studies
The TPK protein sequences of A. thaliana, O. sativa and T. aestivum were used for the multiple sequence alignment in order to study the various conserved residues and regions using the MultAlin and MUSCLE programs at default parameters [38]. Moreover, the full-length protein sequences were also used for the construction of a phylogenetic tree using the Molecular Evolutionary Genetics Analysis (MEGA) 11 software along with the parameters like Neighbor-joining method and 1000 bootstrap value [39].
2.3. Gene structure analysis
The TaTPK exon/intron analysis methodology included the alignment of genomic and its respective coding sequences (CDS) sequences. The pictorial representation of gene architecture including intron phase distribution and exon-intron patterns was prepared through the Gene Structure Display Server (GSDS 2.0) server [40].
2.4. Physicochemical properties and motif analyses
The various features, like peptide length, MW (molecular weight), pI (Isoelectric point), aliphatic index, instability index, and GRAVY (Grand average of hydropathy) values were traced out using the Expasy tool [41]. Moreover, the subcellular localization and presence of transmembrane (TM) helices and the signal peptide were confirmed by WoLF PSORT, TMHMM 2.0 (https://services.healthtech.dtu.dk/service.php?TMHMM-2.0), and SignalP 5.0 0 (https://services.healthtech.dtu.dk/service.php?TMHMM-2.0) tools, respectively. In the case of the motif study, the TaTPK peptide sequences were scanned for the identification of the 15 most conserved motifs, and a pictorial representation was made using MEME 5.1.1 (Multiple Expectation Maximization for Motif Elicitation 5.1.1) suite servers [42].
2.5. Expression profiling in tissues developmental stages
The expression study is an important way to understand the functionality of genes. Therefore, the TaTPK genes were analysed for expression profiling using high throughput RNA-sequence (RNA-seq) data, downloaded from the publicly available URGI database (https://urgi.versailles.inra.fr/files/RNASeqWheat). In our analysis, we have used the RNA-seq data generated under three developmental stages of five different tissues of T. aestivum. These tissues include the root, stem, leaf, spike, and grain. The used RNA-seq data was generated in the biological replicates [43,44]. Subsequently, the Trinity package was used to calculate the Fragments Per Kilobase of transcript per Million mapped reads (FPKM) values from available RNA-seq data and further used for the heat map preparation through the Hierarchical Clustering Explorer (HCE.3.5) tool [45,46].
2.6. Expression profiling under abiotic and biotic stress conditions
The expression profiling was also performed under various abiotic and biotic stress conditions to elucidate their possible involvement in stress resistance. In the case of abiotic stress, biologically replicated RNA-seq data (accession number SRP045409) was generated from the leaf tissue after the 1 and 6 h of heat stress (HS) (40 °C), drought stress (DS) (20 % of PEG 6000), and their combined heat drought (HD) treatments, and used for the expression analysis [47]. However, under salt stress, used high throughput RNA-seq data (accession number SRP062745) was generated in triplicates from the root tissues, after the intervals of 6, 12, 24, and 48 h of salt treatment (150 Mm NaCl) [48].
Moreover, in the case of biotic stress, we have used the RNA-seq data (PRJNA243835), which was generated in triplicates from the leaf tissue of seven days old seedlings after the infestation of two fungal pathogens, Blumaria graminis f. sp. tritici (Bgt) and Puccinia striiformis f. sp. tritici (Pst). The high throughput RNA-seq data was generated after 24, 48, and 72 h of infestation [49]. The heat map generation of both abiotic and biotic stress conditions was carried out using HCE 3.5 tool.
2.7. qRT-PCR analysis
The qRT-PCR was performed following the method established in our laboratory [50,51]. Firstly, the sterilization of seeds of T. aestivum cv Chinese spring was done using sodium hypochlorite (1.2%) and 10% ethanol, further washed with double autoclaved water. Seeds were stratified by keeping them at 4 °C overnight. Next day, seeds wereallowed to grow as hydroponics at room temperature, 16 and 8h light/dark period and 60% relative humidity. To analysis the effects of DS, the seven-day old seedlings were treated with 20% Polyethylene glycol (PEG). The control and treated plants were collected after 1h and 6h in liquid nitrogen. The Trizol method was employed for the extraction of total RNA from both control and treated samples, which was further purified using the TURBO DNA-free TM kit (Invitrogen USA). The quality of RNA samples was assessed through agarose gel electrophoresis and concentration through Nanodrop spectrophotometer. The cDNA was prepared using the RevertAid First Strand cDNA synthesis kit (Thermo Scientific™) from 1ug of RNA sample. The qRT-PCR experiment was performed in triplicates at CFX96 Real-Time PCR (BioRad, USA) using SYBR Green master mix, gene specific primers (Supplementary Table S6) and internal control (ADP ribosylation factor, TaARF) [52]. Fold change was calculated by delta-delta CT (2−ΔΔCT) method.
2.8. Interactions analyses
The interacting partners of TaTPK proteins were identified by means of protein-protein, protein-chemical, protein-transcription factor and protein-miRNA interactions. The STITCH and STRING servers were used for the protein-chemical and protein-protein interactions, respectively. The interacting transcription factors of TaTPK genes were predicted using the PlantRegMap database (http://plantregmap.gao-lab.org/). Cytoscape (https://cytoscape. org/download.html) tool was used for the web network preparation [53,54]. Furthermore, to understand the regulatory aspects of TaTPK, the interacting miRNA partners were identified by the use of known miRNA sequences of T. aestivum through the psRNATarget server. The interaction web was prepared using Gephi 0.9.1 tool [55].
3. Results and discussion
3.1. Identification and genome-wide distribution of TaTPK genes
In the T. aestivum genome, 12 TaTPK genes were identified in four homeologous groups and named as, TaTPK1-A, TaTPK1-B, TaTPK1-D, TaTPK2-A, TaTPK2-B, TaTPK2-D, TaTPK3-A, TaTPK3-B, TaTPK3-D, TaTPK4-A, TaTPK4-B and TaTPK4-D following the standard guidelines (Table 1, Supplementary Table S1). The occurrence of these genes were also confirmed by BLASTn search against the transcriptome shotgun assembly (TSA) sequences available at the NCBI. The majority of the sequences showed 100 % query cover with more than 98% similarity with various TSA sequences, which confirmed their existence at transcript level. Further, the sequence for TaTPK3-A that was incomplete in the Ensembl Plant database, was retrived and completed using the TSA database. Besides, TaTPK4-A gene was lacking in the high confidance gene sequences reported at the Ensembl Plant database,. It was identified as TraesCS5A02G541400LC.1, reported as low confidance gene sequence. Moreover, it was confirmed at TSA datasets, and by domain analysis. According to previous studies, three TPK genes are reported in the O. sativa, two in each of N. tabacum and Saccharum spontaneum, four in Vigna radiata, and five in each of A. thaliana and C. arietinum L. All of them are diploid species. However, 9 and 10 TPK genes are reported in tetraploid Glycine max and triploid Populus tomentosa [17,18,21,22,[56], [57], [58]]. These reports revealed the occurrence of a variable number of TPK genes in various diploid and polyploid plant species. Therefore, the occurrence of 12 TaTPK genes in the allohexaploid genome of T. aestivum wasas per the expectation and in agreement with the previous reports representing the higher number of genes in other families of T. aestivum including various channels [12,31,[59], [60], [61]].
Table 1.
Physico-chemical characteristics of TaTPKs of Triticum aestivum.
| Gene ID | Chromosome | Strand | Exons | Peptide length | MW(kDa) | Isoelectric point | Sub-cellular localization |
|---|---|---|---|---|---|---|---|
| TaTPK1-A | 3A | Forward | 2 | 404 | 44.6 | 8.69 | Plasma membrane |
| TaTPK1-B | 3B | Forward | 2 | 404 | 44.7 | 8.69 | Plasma membrane |
| TaTPK1-D | 3D | Forward | 2 | 404 | 44.6 | 8.79 | Plasma membrane |
| TaTPK2-A | 4A | Reverse | 1 | 366 | 39.1 | 6.97 | Vacuolar |
| TaTPK2-B | 4B | Reverse | 1 | 366 | 39.2 | 6.61 | Vacuolar |
| TaTPK2-D | 4D | Forward | 1 | 366 | 39.2 | 7.66 | Vacuolar |
| TaTPK3-A | 5A | Reverse | 3 | 459 | 48 | 8.91 | Plasma membrane |
| TaTPK3-B | 5B | Reverse | 2 | 458 | 50.1 | 9.11 | Plasma membrane |
| TaTPK3-D | 5D | Reverse | 2 | 458 | 50 | 9.11 | Plasma membrane |
| TaTPK4-A | 5A | Reverse | 3 | 349 | 39.4 | 8.7 | Plasma membrane |
| TaTPK4-B | 5B | Reverse | 3 | 349 | 39.4 | 8.7 | Plasma membrane |
| TaTPK4-D | 5D | Reverse | 2 | 349 | 39.3 | 8.58 | Plasma membrane |
In all the homoeologous groups of TaTPKs, the genes were derived from each A, B and D-subgenome of T. aestivum (Fig. 1A). The number of TaTPK homeologous groups wasconsistent with the number of TPK genes in the diploid plant species [18,21,22]. The TaTPK genes were localized on nine different chromosomes (i.e., 3A, 3B, 3D, 4A, 4B, 4D, 5A, 5B and 5D), the first six of them comprised of one gene each, while each of chromosomes 5A, 5B and 5D consisted of two genes (Fig. 1A). The TaTPK genes, TaTPK1-A, TaTPK1-B, TaTPK1-D and TaTPK2-D were present at the forward strand, while other genes were located on the reverse strand (Supplementary Table S2). The distribution of TPK genes at diverse chromosomes wasalso reported in other crop species such as C. arietinum L., C. cajan, V. radiata and S. spontaneum [21,24,58,62]. In our study, no paralogous genes were identified in TaTPK gene family.
Fig. 1.
Chromosomal localization, gene structure and motif analyses. A. Figure represents the localization of TaTPK genes on chromosomes 3A, 3B, 3D, 4A, 4B, 4D, 5A 5B and 5D. B. Figure shows the pattern of exon/intron distribution and intron phase of TaTPKs.C. Figure shows the 15 conserved motifs of TaTPK proteins predicted by MEME Suite 5.1.1. Each motif is represented by different colours. (For interpretation of the references to colour in this figure legend, the reader is referred to the Web version of this article.)
3.2. Gene structure, exon-intron distribution and intron phase analysis
To understand the structural divergence across the various TaTPK genes, their coding sequence (CDS) length, number and length of exons, exon-intron pattern and intron phase analyses were carried out. The CDS length varied from 1047 to 1374 base pairs (bp), while exons length ranged from 206 to 312 bp. The largest and smallest exons were identified in TaTPK2-B and TaTPK4-B genes, respectively (Supplementary Table S2). TaTPK1 and TaTPK2 group genes consisted of two and one exons, respectively. However, in the case of TaTPK3 and TaTPK4 group genes, TaTPK3-A, TaTPK4-A and TaTPK4-B comprised three exons, and TaTPK3-B, TaTPK3-D and TaTPK4-D enclosed two exons (Fig. 1B). The exon-intron patterns were alike in the majority of homoeologous group genes. Further, the number of exons and exon-intron pattern in TaTPK genes werecomparable to other crop species including C. arietinum L., A. thaliana, C. Cajan, V. radiata, Brassica rapa ssp. pekinensis etc. [[21], [22], [23], [24],63]. Intron phase distribution governs the exon shuffling process that ultimately led to protein domain shuffling during evolution. During the current analysis, the majority of introns of TaTPK genes were found in intron phase 2 (Fig. 1B), which signifies the insertion of introns between the second and third bases of triplet codons.
3.3. Physicochemical features, domains and motifs analyses
Various bioinformatic approaches were used to get insight into the characteristics of TaTPK proteins, which are responsible for their stability and functioning. The length of TaTPK proteins ranged from 349 to 459 amino acid (AA) residues. The largest TaTPK protein was TaTPK3-A, while the smallest were TaTPK4-A, TaTPK4-B and TaTPK4-D. The MW of TaTPKs ranged from 39.1 to 50.2 kDa (kDa), whilepI) varied from 6.6 to 9.2 (Table 1). The majority TaTPKs showed more than 7 pI, suggesting their basic nature. GRAVY, instability and aliphatic index of TaTPK proteins varied from −0.045 to 0.14, 33.57 to 47.31 and 84.59 to 99.43, respectively (Supplementary Table S2). The protein length, MW and pI of TaTPKs wereconsistent with the previously published reports [[21], [22], [23], [24],58,63]. The occurrence of ⁓40 computed instability index of TaTPK proteins indicated their stable architecture. A higher aliphatic index (above 80) suggested their thermo-stable nature. The negative GRAVY values indicated the hydrophilic nature of TaTPK1-D, TaTPK2-D, TaTPK3-A, TaTPK3-B and TaTPK3-D, while positive GRAVY values of the remaining TaTPK proteins reveal their hydrophobic nature. Moreover, the analysed GRAVY value indicated the overall nature of TaTPK proteins, not for a specific domain/portion. Since all these channels are located in the membrane, they will have both hydrophobic and hydrophilic portions in the TM and cytosolic portions, respectively.
The subcellular localization analysis predicted the plasma membrane localization for the members of TaTPK1, TaTPK3 and TaTPK4 group channels, and vacuolar membrane for TaTPK2 group proteins (Table 1). In the case of A. thaliana, AtTPK1, AtTPK2, AtTPK3, and AtTPK5 are localized on the vacuolar membrane, while AtTPK4 on the plasma membrane [64]. In O. sativa, OsTPKa and OsTPKb have been reported in lytic and storage vacuoles, respectively [20].
The domain composition is a major factor in determining the functionality of any protein. The conserved domains and motifs were analysed in the TaTPK proteins. Two ion channel domains (PF07885) were identified, one each at N- and C-terminal regions in all the TaTPK proteins (Supplementary Table S3). A total of four TM helices, two in each half were present in each TaTPK protein (Fig. 2). The results are in agreement with the earlier reports [[20], [21], [22], [23], [24],58,63]. Additionally, various conserved residues identified using the multiple sequence alignments of TPK protein sequences of A. thaliana, T. aestivum and O. sativa. Two GYGD motifs present between TM1 and TM2, and TM3 and TM4 of the N- and C-terminal domains were detected as highly conserved motifs (Fig. 2A–C). In addition, one 14-3-3 protein binding site and two EF-hand domains were found highly conserved in the N- and C-terminal domains, respectively. Moreover, TaTPK1-D lacked a second EF-hand domain. These conserved domains and motifs play a vital role in determining the function of proteins. The above-identified domains and motifs have also been reported in TPK channels in previous studies [[20], [21], [22], [23], [24],58,63,65].). The conserved GYGD motif is responsible for ion selectivity, where Y and D residues are vital for K+ selectivity and gating [66]. The presence of EF-hand domains in the C-terminal cytoplasmic region revealed the connection of TaTPK channels with the cytoplasmic concentration of Ca2+ as earlier reports suggested that dependence of TPK channel gating on the Ca2+ ions [20]. A few studies suggested the role of the 14-3-3 domain in channel trafficking from the endoplasmic reticulum (ER) to the plasma membrane [[67], [68], [69]]. On the contrary, another report in A. thaliana could not support the above finding in the case of TPK1 and suggested that the 14-3-3 domain has no role in ER and vacuolar trafficking [64]. In another study, domain swapping between TPKa and TPKb of O. sativa suggested that 14-3-3 might be associated with ER release but not play a vital role in vacuolar trafficking [20].
Fig. 2.
Multiple sequence alignment and domain analysis of TaTPK proteins of A. thaliana, O. sativa and T. aestivum. Fig. A represents 14-3-3 proteins bind sites and N-terminal ion channel domain, B shows the C-terminal ion channel domain and EF-hand domains, and C depicts the visual representation of TPK protein's transmembrane domains, conserved GYGD domain, 14-3-3 binding site, and EF-hand domains.
Additionally, the occurenrence of 15 conserved motifs in TaTPK proteins were analysed using the MEME suite. Motifs 1, 2, 3, 4, 5,6, 8, 9 and 14 were conserved in all the TaTPK proteins (Fig. 1C). Motifs 7 and 12 were present in 6, motifs 10 and 11 in 3, and motif 13 in 9 and motif 15 in 2 TaTPK proteins. Motif 1 was identified as a part of Ca2+ binding EF-hand domains. Motif 2 was part of C-terminal ion transporter domain, while motif 3 is a part of the N-terminal ion transporter domain. Motif 4 was recognised as part of the TM domain.
3.4. Protein-protein interaction analysis
The protein-protein interaction study was performed using bioinformatic approaches, to study the coordination of TaTPK channels with other channels and proteins. A total of 18 different proteins showed interaction with TaTPK proteins (Fig. 3A, Supplementary Table S4). Most of the interacting proteins included K+ transporters and channels such as K+ efflux antiporter (KEA), Shaker family K+ channel (KT), High-affinity K+ transporter (HKT), KT2/3 potassium channel, shaker family of voltage-gated potassium channel (KAT), Gated outwardly-rectifying K+ channel (GORK), and Kup system potassium uptake protein (KUP). They also showed interaction with other ion transporters such as two-pore calcium channel protein (TPC), magnesium/proton exchanger (MHX) and chloride channel protein. In addition, calcium-dependent protein kinase (CDPK), 14-3-3-like protein GF14 lambda (RCI), 14-3-3-like protein GF14 Ω and thiamine-phosphate diphosphorylase also showed interaction with TaTPK channels. The interaction of TaTPK channels with other K+ channels and transporters suggested their co-ordinated function in K+ homeostasis and abiotic stress response such as drought, salt, cold etc. [70,71]. The interaction of TaTPK channels with TPC and CDPK, and the presence of EF-hand domains suggested their regulation by Ca2+-related signaling pathways [72,73]. Since, TaTPK channels consisted of 14-3-3 binding sites, their interaction with 14-3-3 proteins was obvious. Moreover, 14-3-3 proteins play diverse roles in plants from growth to abiotic stress response [74], which indicated the involvement of TaTPKs in diverse functions.
Fig. 3.
Protein-protein and protein-chemical interaction analyses. The figure demonstrates the interaction network of TaTPK proteins with (A) other proteins and (B) chemical compounds.
3.5. Protein-chemical interaction analysis
To study the involvement of TaTPK proteins in catalytic mechanisms and their regulation by other biochemicals, their interaction with various chemical compounds and metal ions was studied. TaTPKs showed interaction with nine different biochemical molecules such as diphosphopyridine nucleotide (DPN), adenosine monophosphate (AMP), cyclic AMP (cAMP), MgADP, reduced glutathione and glutamic acid along with various metal ions including Na+, Ca2+ and K+ (Fig. 3B–Supplementary Table S4). DPN might act as a co-enzyme, AMP and ADP interactions indicate energy-driven active transport by TaTPKs, cAMP might play regulatory role, while interactions with reduced glutathione and glutamic acid depicted their role in stress responses [[75], [76], [77], [78]]. However, these interactions need to be experimentally validated in future studies. Interaction with K+ further depicted their involvement in K+ transport, while Ca2+ interaction showed the putative functioning of EF-hand domain in TPK channels and their Ca2+−mediated regulation, while Na+ interaction indicated their role in salt stress response.
3.6. Phylogenetic analysis
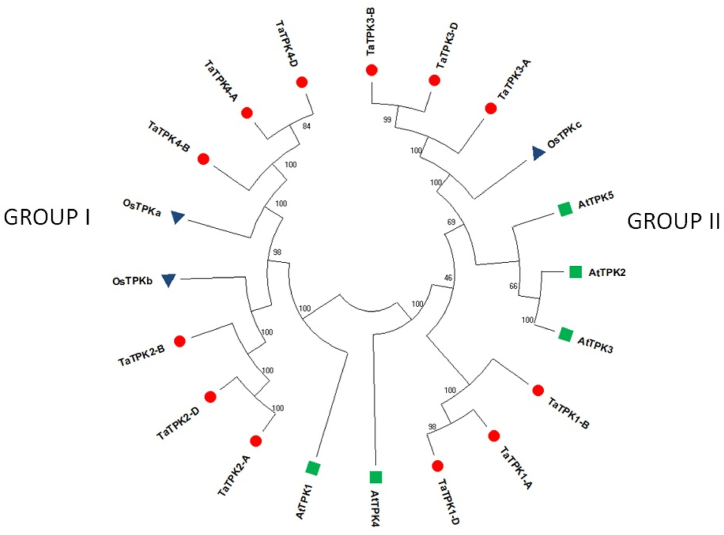
Evolutionary connection was analysed among the TPK proteins of A. thaliana, O. sativa and T. aestivum through the phylogenetic tree (Fig. 4). The TPK proteins were clustered into two groups (I and II) as reported in earlier studies [[21], [22], [23],62,79]. Group I and II comprised 9 and 11 TPK proteins, respectively. A total of six TPKs of T. aestivum, two of O. sativa (TaTPKa and TaTPKb) and one of A. thaliana (AtTPK1) closely clustered in group I, while six TPKs of T. aestivum, one of O. sativa and four of A. thaliana clustered in group II of the phylogenetic tree. The TPKs of both monocots; T. aestivum and O. sativa clustered tightly. Additionally, the homeologous TaTPK proteins, for instance, TaTPK1-A, TaTPK1-B and TaTPK1-D; TaTPK2-A, TaTPK2-B; TaTPK2-D; TaTPK4-A, TaTPK4-B and TaTPK4-D etc. were clustered together in the phylogenetic tree, due to a higher sequence similarity among them.
Fig. 4.
Phylogenetic analysis of TPKs of A. thaliana, O. sativa and T. aestivum. The phylogenetic tree displays the clustering of TPK proteins into two groups: I and II. TPKs of each A. thaliana, O.sativa, T. aestivum are denoted by different markers.
3.7. Expression profiling of TaTPKs in tissues developmental stages
High throughput RNA-seq data collected from three developmental stages of the root, stem, leaf, spike, and grain tissues have been used for expression profiling. TaTPK genes are clustered into two groups on the basis of their expression trend (Fig. 5A). In group I, TaTPK4-B and TaTPK4-D showed significant expression in all the tissues and exhibited higher expression in root and grain tissues. TaTPK2-A, TaTPK2-B and TaTPK2-D expressed in later stages of the leaf i.e., leaf_z23 and leaf_z71. In group II, TaTPK1-A, TaTPK1-B and TaTPK1-D showed expression in spike_z65 and all the leaf developmental stages. On the other hand, TaTPK3-A, TaTPK3-B and TaTPK3-D showed expression at spike_z65 and grain_z85 tissues. These results showed that TaTPK genes exhibited differential expression in various stages of vegetative and reproductive tissues. Earlier, TPK1 of Fragaria ananassa was reported to play an important role in fruit development [80]. In A. thaliana, TPK1 was involved in seed germination and stomatal activity [13] and TPK4 regulated the membrane voltage of the rising pollen tube [26]. The specifically higher expression of certain TaTPK genes in spike and grain tissues suggested their role in reproductive development, while others that showed specifically higher levels in various organs suggested roles in vegetative growth and development.
Fig. 5.
Expression analysis of TaTPK genes in different tissues and during abiotic and biotic stresses and qRT-PCR analysis The expression profiles of TaTPK genes are represented in the form of heat maps for (A) different developmental stages, (B) biotic stress including Bgt and Pst infestation for 24, 48 and 72 h, (C) 1 and 6 h of DS, HS and HD, and (D) 6, 12, 24 and 48h of salt stress. Figure (E) show the qRT-PCR results of three TaTPK genes under DS stress at 1 and 6h of treatments. The bar graphs indicate the fold change of each TaTPK gene expression and a vertical line indicate standard deviation.
3.8. Expression profiling of TaTPKs under biotic stress
To study the effect of pathogens on TaTPK genes, high throughput RNA seq data of leaf tissues treated with two fungal pathogens were used for expression profiling. The TaTPK genes clustered in two groups under biotic stress conditions (Fig. 5B). In group I, the TaTPK1-D showed upregulation and TaTPK3-B showed downregulation after each hour (h) of Bgtand Pst infestation. However, TaTPK2-A, TaTPK2-B and TaTPK2-D were upregulated after Pst infestation. In group II, the gene TaTPK3-D showed upregulation after each h of Bgt and Pst inoculation, except 72 h of Pst. TaTPK3-A exhibited upregulation after Bgt infestation only. TaTPK1-A and TaTPK1-B were upregulated after each h of Bgt and Pst inoculation except 24 h of Pst. TaTPK4-B was upregulated after Bgt and 24 h of Pst inoculation, while TaTPK4-D showed upregulation after 48 and 72 h of Bgt infestation. The differential expression of TPK genes of T. aestivum at different hours of Bgt and Pst infestation suggested their role in biotic stress responses. Earlier studies also suggested the role of K+ channels in biotic stress responses [4,81]. For example, a study suggested the role of TPK1 of F. ananassa in plant resistance against the pathogen Botrytis cinerea [80].
3.9. Expression analysis of TaTPKs under abiotic stresses
Abiotic stresses are a major concern to wheat production as they cause an average of 50% losses to wheat yield [82]. Therefore, to understand the role of the TaTPK family under abiotic stresses, the expression analysis is carried out under DSHS), HD and salt stress conditions using high throughput RNA-seq data. Based on expression data, two main groups were formed in heat maps under DS, HS and HD (Fig. 5C). In group I, the majority of TaTPK genes were highly upregulated at both 1 and 6 h of DS, except TaTPK4-D. On the other hand, the group II genes showed upregulated expression at 6h of HS and some of the TaTPKs, for instance, TaTPK4-B and TaTPK3-D showed higher expression at later h of HD.
Similarly, using high throughput RNA-seq data, the expression of TaTPKs was analysed under salt stress. Here also two groups were formed based on expression trend (Fig. 5D). The group I genes such as TaTPK2-B, TaTPK2-D and TaTPK3-A were upregulated at 24 h of salt stress. However, in group II, TaTPK1-A and TaTPK3-D genes exhibited higher expression at 6 h of salt stress, and TaTPK3-B at 12 h of treatment. Most of the genes in group II demonstrated downregulated expression under salt stress.
The qRT-PCR analysis of TaTPK1-A, TaTPK3-A and TaTPK4-B was carried out at 1 and 6 h of DS (Fig. 5E). The expression of TaTPK1-A and TaTPK4-B was comparable to the high throughput RNA-seq data. the TaTPK1-A showed downregulation at both 1h and 6h of DS. TaTPK4-B showed downregulation at 1 h and upregulation at 6 h of DS. However, TaTPK3-A showed downregulation at both 1 h and 6 h of DS (Fig. 5E).
Similarly, the expression of NtTPK1 also increased about 2-fold under osmotic and salinity stress [17]. In another report, it has been shown that in C. arietinum, the CarTPK3.2 was slightly up-regulated under HS [20]. In addition, the OsTPKb overexpressing transgenic lines showed better growth under drought stress than non-transgenic lines [28]. Likewise, in V. radiata, VrTPK3 gene has upregulated under DS, HS and salt stresses [62]. Furthermore, in G.raimondii, three TPK genes, GrTPK5.2, GrTPK1.1, GrTPK5.1 showed upregulation under salt stress [23]. The modulated expression of TaTPKs under various stress conditions revealed their role in abiotic stress response in T. aestivum, as reported in other crop species [23,28].
3.10. Gene regulatory networks analyses
To understand the regulatory mechanism of TaTPK genes, their interaction with miRNAs and transcription factors was analysed. The miRNA binding analysis suggested the interaction of 21 miRNAs with TaTPKs of T. aestivum (Fig. 6A). The majority of the TaTPKs were targeted via a cleavage process followed by translation inhibition (Supplementary Table S5). Six miRNAs targeted the TaTPKs via translation, while 15 miRNAs targeted the TaTPKs via cleavage. Some of the miRNAs targeted the homoeologous genes, for instance, ta-miR170a targeted the TaTPK1-A and TaTPK1-B, ta-miR012a targeted the TaTPK3-A, TaTPK3-B and TaTPK3-D, tae-miR1536a targated TaTPK4-B, TaTPK4-D etc.
Fig. 6.
miRNA and transcription factor interaction analyses. A. Figure represents the interaction network of TaTPK genes with known miRNA of T. aestivum. B. Figure shows the transcription factors interaction network generated by Cytoscape. The probable regulating transcription factors of TaTPKs are predicted by the PlantRegMap database.
A total of 38 transcription factors showed interaction with TaTPK genes (Fig. 6B). Most of the identified transcription factors were related to light, hormone, stress responses and growth and development. Hormone-responsive interacting transcription factors were BES1, ERF, EIL and ARF etc., while stress-responsive interacting transcription factors were WRKY, MYB_related, CAMTA, MYB, C3H etc. The growth and development-related interacting transcription factors were LFY, WOX, TALE, SBP, CPP, LBD, AP2, YABBY etc (Fig. 6B).
The role of interacting tae-miR159 in salinity, cold and wound stress response further indicates the involvement of TaTPKs in abiotic stress responses [83]. The miR159 and miR164 are key players in the hormone-responsive signalling cascade. miR159 regulated the abscisic acid (ABA) and gibberellin signalling via MYC transcription factor, while miRNA164 regulated the auxin signalling process via ARF transcription factor [[84], [85], [86]]. The interaction of TaTPKs with these miRNAs, and transcription factor MYC and ARF revealed the regulation of TaTPKs through hormone-related signalling processes. The interacting tae-miR395a is reported to be significantly upregulated in K+ deficient conditions which indicated the ′role of TaTPKs in plant survival at the time of K+ ion deficit [87]. Another study in T. aestivum reported an H2O2-mediated response of tae-miR395a which indicates ROS-mediated regulation [88]. Furthermore, the involvement of miR1136a in drought stress revealed the role of interacting TaTPKs in drought response [89]. The identified transcription factor suggested the role of TaTPK genes in light, hormone, stress responses, and growth and development. The interaction of TaTPK proteins with CAMTA and MYC transcription factors suggested their putative functioning in drought and salt tolerance [90,91].
4. Conclusions
In the current study, we explored various features of TaTPKs, an important K+ ion channel, in a staple crop T. aestivum. The interaction of TaTPKs with miRNA, transcription factors, and bio-chemicals suggested their connection with hormone-responsive signaling, drought stress, salt stress response and plant growth and development. Protein-protein interaction analysis indicated coordinated functioning of TaTPKs with other K+ channels and transporters during K+ homeostasis and might be regulated by Ca2+ signaling cascades. The transcript abundance of TaTPKs in roots, grains and leaves suggested their role in both vegetative and reproductive tissues. Their modulated expression suggested an implicit role in biotic and abiotic stress response. Moreover, the specific functions of each TaTPK should be revealed in future studies. This study would facilitate the functional characterization of important TaTPK genes in future studies, which might also be useful in crop improvement programs.
Data availability statement
All the data are available within the manuscript and supplementary files.
Consent for publication
All the authors provided their consent for publication.
CRediT authorship contribution statement
Amandeep Kaur: Writing – original draft, Data curation. Madhu: Methodology, Data curation. Alok Sharma: Methodology, Data curation. Kashmir Singh: Resources. Santosh Kumar Upadhyay: Writing – review & editing, Validation, Supervision, Resources, Investigation, Funding acquisition, Formal analysis, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
The authors are grateful to the Panjab University, Chandigarh, India for research facilities. SKU is grateful to the Science and Engineering Research Board (SERB), Government of India for the Core Research Grant (CRG/2021/000040). M is grateful to UGC, while AS and AK are thankful to CSIR for their respective fellowships.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2024.e27814.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
figs1
References
- 1.Hawkesford M., Horst W., Kichey T., Lambers H., Schjoerring J., Møller I.S., et al. Marschner's Mineral Nutrition of Higher Plants. Academic press; 2012. Functions of macronutrients; pp. 135–189. [Google Scholar]
- 2.Véry A.A., Sentenac H. Molecular mechanisms and regulation of K+ transport in higher plants. Annu. Rev. Plant Biol. 2003;54:575–603. doi: 10.1146/annurev.arplant.54.031902.134831. [DOI] [PubMed] [Google Scholar]
- 3.Amtmann A., Troufflard S., Armengaud P. The effect of potassium nutrition on pest and disease resistance in plants. Physiol. Plantarum. 2008;133:682–691. doi: 10.1111/j.1399-3054.2008.01075.x. [DOI] [PubMed] [Google Scholar]
- 4.Shabala S., Pottosin I. Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol. Plantarum. 2014;151:257–279. doi: 10.1111/ppl.12165. [DOI] [PubMed] [Google Scholar]
- 5.Wu H., Zhang X., Giraldo J.P., Shabala S. It is not all about sodium: revealing tissue specificity and signalling roles of potassium in plant responses to salt stress. Plant Soil. 2018;431:1–17. [Google Scholar]
- 6.Isayenkov S.V., Maathuis F.J. Plant salinity stress: many unanswered questions remain. Front. Plant Sci. 2019;10:80. doi: 10.3389/fpls.2019.00080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Schroeder J.I., Ward J.M., Gassmann W. Perspectives on the physiology and structure of inward-rectifying K+ channels in higher plants: biophysical implications for K+ uptake. Annu. Rev. Biophys. Biomol. Struct. 1994;23:441–471. doi: 10.1146/annurev.bb.23.060194.002301. [DOI] [PubMed] [Google Scholar]
- 8.Leigh R.A. Potassium homeostasis and membrane transport. J. Plant Nutr. Soil Sci. 2001;164:193–198. [Google Scholar]
- 9.Demidchik V., Maathuis F.J. Physiological roles of nonselective cation channels in plants: from salt stress to signalling and development. New Phytol. 2007;175:387–404. doi: 10.1111/j.1469-8137.2007.02128.x. [DOI] [PubMed] [Google Scholar]
- 10.Lebaudy A., Véry A.A., Sentenac H. K+ channel activity in plants: genes, regulations and functions. FEBS Lett. 2007;581:2357–2366. doi: 10.1016/j.febslet.2007.03.058. [DOI] [PubMed] [Google Scholar]
- 11.Sharma T., Dreyer I., Riedelsberger J. The role of K+ channels in uptake and redistribution of potassium in the model plant Arabidopsis thaliana. Front. Plant Sci. 2013;4:224. doi: 10.3389/fpls.2013.00224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kaur A., Sharma A., Dixit S., Singh K., Upadhyay S.K., S. K OSCA genes in bread wheat: molecular characterization, expression profiling, and interaction analyses indicated their diverse roles during development and stress response. Int. J. Mol. Sci. 2022;23 doi: 10.3390/ijms232314867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gobert A., Isayenkov S., Voelker C., Czempinski K., Maathuis F.J. The two-pore channel TPK1 gene encodes the vacuolar K+ conductance and plays a role in K+ homeostasis. Proc. Natl. Acad. Sci. U.S.A. 2007;104:10726–10731. doi: 10.1073/pnas.0702595104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hartley T.N., Maathuis F.J. Allelic variation in the vacuolar TPK 1 channel affects its calcium dependence and may impact on stomatal conductance. FEBS Lett. 2016;590:110–117. doi: 10.1002/1873-3468.12035. [DOI] [PubMed] [Google Scholar]
- 15.Latz A., Becker D., Hekman M., Müller T., Beyhl D., Marten I., et al. TPK1, a Ca2+‐regulated Arabidopsis vacuole two‐pore K+ channel is activated by 14‐3‐3 proteins. Plant J. 2007;52:449–459. doi: 10.1111/j.1365-313X.2007.03255.x. [DOI] [PubMed] [Google Scholar]
- 16.Maathuis F.J. Vacuolar two‐pore K+ channels act as vacuolar osmosensors. New Phytol. 2011;191:84–91. doi: 10.1111/j.1469-8137.2011.03664.x. [DOI] [PubMed] [Google Scholar]
- 17.Hamamoto S., Marui J., Matsuoka K., Higashi K., Igarashi K., Nakagawa T., et al. Characterization of a tobacco TPK-type K+ channel as a novel tonoplast K+ channel using yeast tonoplasts. J. Biol. Chem. 2008;283:1911–1920. doi: 10.1074/jbc.M708213200. [DOI] [PubMed] [Google Scholar]
- 18.Voelker C., Schmidt D., Mueller‐Roeber B., Czempinski K. Members of the Arabidopsis AtTPK/KCO family form homomeric vacuolar channels in planta. Plant J. 2006;48:296–306. doi: 10.1111/j.1365-313X.2006.02868.x. [DOI] [PubMed] [Google Scholar]
- 19.Sano T., Kutsuna N., Becker D., Hedrich R., Hasezawa S. Outward‐rectifying K+ channel activities regulate cell elongation and cell division of tobacco BY‐2 cells. Plant J. 2009;57:55–64. doi: 10.1111/j.1365-313X.2008.03672.x. [DOI] [PubMed] [Google Scholar]
- 20.Isayenkov S., Isner J.C., Maathuis F.J. Rice two-pore K+ channels are expressed in different types of vacuoles. Plant Cell. 2011;23:756–768. doi: 10.1105/tpc.110.081463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Azeem F., Ahmad B., Atif R.M., Ali M.A., Nadeem H., Hussain S., et al. Genome-wide analysis of potassium transport-related genes in chickpea (Cicer arietinum L.) and their role in abiotic stress responses. Plant Mol. Biol. Rep. 2018;36:451–468. [Google Scholar]
- 22.Azeem F., Hussain M., Hussain S., Zubair M., Nadeem H., Ali M.A., et al. Genome-wide analysis and expression profiling of potassium transport related genes in Solanum tuberosum. Pakistan J. Agric. Sci. 2021;58:81–94. [Google Scholar]
- 23.Azeem F., Zameer R., Rashid M.A.R., Rasul I., Ul-Allah S., Siddique M.H., et al. Genome-wide analysis of potassium transport genes in Gossypium raimondii suggest a role of GrHAK/KUP/KT8, GrAKT2. 1 and GrAKT1. 1 in response to abiotic stress. Plant Physiol. Biochem. 2022;170:110–122. doi: 10.1016/j.plaphy.2021.11.038. [DOI] [PubMed] [Google Scholar]
- 24.Siddique M.H., Babar N.I., Zameer R., Muzammil S., Nahid N., Ijaz U., et al. Genome-wide identification, genomic organization, and characterization of potassium transport-related genes in cajanus cajan and their role in abiotic stress. Plants. 2021;10:2238. doi: 10.3390/plants10112238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Czempinski K., Frachisse J.M., Maurel C., Barbier‐Brygoo H., Mueller‐Roeber B. Vacuolar membrane localization of the Arabidopsis ‘two‐pore’K+ channel KCO1. Plant J. 2002;29:809–820. doi: 10.1046/j.1365-313x.2002.01260.x. [DOI] [PubMed] [Google Scholar]
- 26.Becker D., Geiger D., Dunkel M., Roller A., Bertl A., Latz A., et al. AtTPK4, an Arabidopsis tandem-pore K+ channel, poised to control the pollen membrane voltage in a pH-and Ca2+-dependent manner. Proc. Natl. Acad. Sci. U.S.A. 2004;101:15621–15626. doi: 10.1073/pnas.0401502101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Isayenkov S.V., Mian A., Maathuis F.J.M. The overexpression of gene encoding rice potassium channel—OsТРКa increases the salt and drought tolerance of plants. Biotechnologia Acta. 2015;8:78–83. [Google Scholar]
- 28.Ahmad I., Devonshire J., Mohamed R., Schultze M., Maathuis F.J. Overexpression of the potassium channel TPK b in small vacuoles confers osmotic and drought tolerance to rice. New Phytol. 2016;209:1040–1048. doi: 10.1111/nph.13708. [DOI] [PubMed] [Google Scholar]
- 29.Madhu, Kaur A., Singh K., Upadhyay S.K. Ascorbate oxidases in bread wheat: gene regulatory network, transcripts profiling, and interaction analyses provide insight into their role in plant development and stress response. Plant Growth Regul. 2023:1–16. [Google Scholar]
- 30.Madhu A. Sharma, Kaur A., Singh K., Upadhyay S.K. Modulation in gene expression and enzyme activity suggested the roles of monodehydroascorbate reductase in development and stress response in bread wheat. Plant Sci. 2024;338 doi: 10.1016/j.plantsci.2023.111902. [DOI] [PubMed] [Google Scholar]
- 31.Kaur A., Madhu, Sharma A., Singh K., Upadhyay S.K. Exploration of Piezo channels in bread wheat (Triticum aestivum L.) Agriculture. 2023;13:783. [Google Scholar]
- 32.Mistry J., Chuguransky S., Williams L., Qureshi M., Salazar G.A., Sonnhammer E.L., et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 2021;49:D412–D419. doi: 10.1093/nar/gkaa913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Letunic I., Doerks T., Bork P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 2015;43:D257–D260. doi: 10.1093/nar/gku949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Marchler-Bauer A., Bo Y., Han L., He J., Lanczycki C.J., Lu S., et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017;45:D200–D203. doi: 10.1093/nar/gkw1129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Khalil H.B., Brunetti S.C., Pham U.M., Maret D., Laroche A., Gulick P.J. Characterization of the caleosin gene family in the Triticeae. BMC Genom. 2014;15:239. doi: 10.1186/1471-2164-15-239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Brunetti S.C., Arseneault M.K.M., Gulick P.J. Characterization of the Esi3/RCI2/PMP3 gene family in the Triticeae. BMC Genom. 2018;19:898. doi: 10.1186/s12864-018-5311-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Brunetti S.C., Arseneault M.K.M., Gulick P.J. Characterization and expression analysis of the Pirin gene family in Triticum aestivum. Genom. 2022;65:349–362. doi: 10.1139/gen-2021-0094. [DOI] [PubMed] [Google Scholar]
- 38.Edgar R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tamura K., Stecher G., Kumar S. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 2021;38:3022–3027. doi: 10.1093/molbev/msab120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hu B., Jin J., Guo A.Y., Zhang H., Luo J., Gao G.G.S.D.S.2. 0: an upgraded gene features visualization server. Bioinformatics. 2015;31:1296–1297. doi: 10.1093/bioinformatics/btu817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Gasteiger E., Hoogland C., Gattiker A., Duvaud S.E., Wilkins M.R., Appel R.D., et al. Humana press; 2005. Protein Identification and Analysis Tools on the ExPASy Server; pp. 571–607. [DOI] [PubMed] [Google Scholar]
- 42.Bailey T.L., Boden M., Buske F.A., Frith M., Grant C.E., Clementi L. Meme suite: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–W208. doi: 10.1093/nar/gkp335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Choulet F., Alberti A., Theil S., Glover N., Barbe V., V, Daron J. Structural and functional partitioning of bread wheat chromosome 3B. Science. 2014;345 doi: 10.1126/science.1249721. [DOI] [PubMed] [Google Scholar]
- 44.Pingault L., Choulet F., Alberti A., Glover N., Wincker P., Feuillet C., et al. Deep transcriptome sequencing provides new insights into the structural and functional organization of the wheat genome. Genome Biol. 2015;16:1–15. doi: 10.1186/s13059-015-0601-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Haas B.J., Papanicolaou A., Yassour M., Grabherr M., Blood P.D., Bowden J. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 2013;8:1494–1512. doi: 10.1038/nprot.2013.084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Seo J., Gordish-Dressman H., Hoffman E.P. An interactive power analysis tool for microarray hypothesis testing and generation. Bioinformatics. 2006;22:808–814. doi: 10.1093/bioinformatics/btk052. [DOI] [PubMed] [Google Scholar]
- 47.Liu Z., Xin M., Qin J., Peng H., Ni Z., Yao Y., et al. Temporal transcriptome profiling reveals expression partitioning of homeologous genes contributing to heat and drought acclimation in wheat (Triticum aestivum L.) BMC Plant Biol. 2015;15:1–20. doi: 10.1186/s12870-015-0511-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhang Y., Liu Z., Khan A.A., Lin Q., Han Y., Mu P., et al. Expression partitioning of homeologs and tandem duplications contribute to salt tolerance in wheat (Triticum aestivum L.) Sci. Rep. 2016;6:1–10. doi: 10.1038/srep21476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang H., Yang Y., Wang C., Liu M., Li H., Fu Y., et al. Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genom. 2014;15:1–14. doi: 10.1186/1471-2164-15-898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Shumayla, Tyagi S., Sharma Y., Sharma A., Madhu, Pandey A., et al. Expression of TaNCL2-A ameliorates cadmium toxicity by increasing calcium and enzymatic antioxidants activities in Arabidopsis. Chemosphere. 2023;29 doi: 10.1016/j.chemosphere.2023.138636. [DOI] [PubMed] [Google Scholar]
- 51.Tyagi S., Shumayla, Sharma Y., Madhu A. Sharma, Pandey A., et al. TaGPX1-D overexpression provides salinity and osmotic stress tolerance in Arabidopsis. Plant Sci. 2023;337 doi: 10.1016/j.plantsci.2023.111881. [DOI] [PubMed] [Google Scholar]
- 52.Sharma Y., Dixit S., Singh K., Upadhyay S.K. Decoding the features and potential roles of respiratory burst oxidase homologs in bread wheat. Curr. Plant Biol. 2024;37 [Google Scholar]
- 53.Franceschini A., Szklarczyk D., Frankild S., Kuhn M., Simonovic M., Roth A., et al. STRING v9. 1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2012;41:D808–D815. doi: 10.1093/nar/gks1094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kuhn M., Szklarczyk D., Franceschini A., Von Mering C., Jensen L.J., Bork P. Stitch 3: zooming in on protein–chemical interactions. Nucleic Acids Res. 2012;40:D876–D880. doi: 10.1093/nar/gkr1011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Bastian M., Heymann S., Jacomy M. Gephi: an open-source software for exploring and manipulating networks. Proceedings of the international AAAI conference on web and social media. 2009;3:361–362. [Google Scholar]
- 56.Gomez-Porras J.L., Riaño-Pachón D.M., Benito B., Haro R., Sklodowski K., Rodríguez-Navarro A., et al. Phylogenetic analysis of K+ transporters in bryophytes, lycophytes, and flowering plants indicates a specialization of vascular plants. Front. Plant Sci. 2012;3:167. doi: 10.3389/fpls.2012.00167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Rehman H.M., Nawaz M.A., Shah Z.H., Daur I., Khatoon S., Yang S.H., Chung G. In-depth genomic and transcriptomic analysis of five K+ transporter gene families in soybean confirm their differential expression for nodulation. Front. Plant Sci. 2017;8:804. doi: 10.3389/fpls.2017.00804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Feng X., Wang Y., Zhang N., Wu Z., Zeng Q., Wu J., et al. Genome-wide systematic characterization of the HAK/KUP/KT gene family and its expression profile during plant growth and in response to low-K+ stress in Saccharum. BMC Plant Biol. 2020;20:1–17. doi: 10.1186/s12870-019-2227-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kaur A., Taneja M., Tyagi S., Sharma A., Singh K., Upadhyay S.K. Genome-wide characterization and expression analysis suggested diverse functions of the mechanosensitive channel of small conductance-like (MSL) genes in cereal crops. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-73627-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Sharma H., Taneja M., Upadhyay S.K. Identification, characterization and expression profiling of cation-proton antiporter superfamily in Triticum aestivum L. and functional analysis of TaNHX4-B. Genomics. 2020;112:356–370. doi: 10.1016/j.ygeno.2019.02.015. [DOI] [PubMed] [Google Scholar]
- 61.Sharma H., Sharma A., Rajput R., Sidhu S., Dhillon H., Verma P.C., et al. Molecular characterization, evolutionary analysis, and expression profiling of BOR genes in important Cereals. Plants. 2022;11:911. doi: 10.3390/plants11070911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Azeem F., Ijaz U., Ali M.A., Hussain S., Zubair M., Manzoor H., et al. Genome-wide identification and expression profiling of potassium transport-related genes in Vigna radiata under abiotic stresses. Plants. 2021;11:2. doi: 10.3390/plants11010002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Shen C., Yuan J. Genome-wide investigation and expression analysis of K+-transport-related gene families in Chinese cabbage (Brassica rapa ssp. pekinensis) Biochem. Genet. 2021;59:256–282. doi: 10.1007/s10528-020-10004-z. [DOI] [PubMed] [Google Scholar]
- 64.Dunkel M., Latz A., Schumacher K., Müller T., Becker D., Hedrich R. Targeting of vacuolar membrane localized members of the TPK channel family. Mol. Plant. 2008;1:938–949. doi: 10.1093/mp/ssn064. [DOI] [PubMed] [Google Scholar]
- 65.Dabravolski S.A., Isayenkov S.V. New insights into plant TPK ion channel evolution. Plants. 2021;10:2328. doi: 10.3390/plants10112328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Chapman M.L., Krovetz H.S., VanDongen A.M. GYGD pore motifs in neighbouring potassium channel subunits interact to determine ion selectivity. J. Plant Physiol. 2001;530:21. doi: 10.1111/j.1469-7793.2001.0021m.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Coblitz B., Shikano S., Wu M., Gabelli S.B., Cockrell L.M., Spieker M., et al. C-terminal recognition by 14-3-3 proteins for surface expression of membrane receptors. J. Biol. Chem. 2005;280:36263–36272. doi: 10.1074/jbc.M507559200. [DOI] [PubMed] [Google Scholar]
- 68.Shikano S., Coblitz B., Sun H., Li M. Genetic isolation of transport signals directing cell surface expression. Nat. Cell Biol. 2005;7:985–992. doi: 10.1038/ncb1297. [DOI] [PubMed] [Google Scholar]
- 69.O'Kelly I., Goldstein S.A. Forward transport of K2P3. 1: mediation by 14‐3‐3 and COPI, modulation by p11. Traffic. 2008;9:72–78. doi: 10.1111/j.1600-0854.2007.00663.x. [DOI] [PubMed] [Google Scholar]
- 70.Becker D., Hoth S., Ache P., Wenkel S., Roelfsema M.R.G., Meyerhoff O., et al. Regulation of the ABA-sensitive Arabidopsis potassium channel gene GORK in response to water stress. FEBS Lett. 2003;554:119–126. doi: 10.1016/s0014-5793(03)01118-9. [DOI] [PubMed] [Google Scholar]
- 71.Liang M., Gao Y., Mao T., Zhang X., Zhang S., Zhang H., et al. Characterization and expression of KT/HAK/KUP transporter family genes in willow under potassium deficiency, drought, and salt stresses. BioMed Res. Int. 2020;2020 doi: 10.1155/2020/2690760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Patel S., Marchant J.S., Brailoiu E. Two-pore channels: regulation by NAADP and customized roles in triggering calcium signals. Cell Calcium. 2010;47:480–490. doi: 10.1016/j.ceca.2010.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Shabbir R., Javed T., Hussain S., Ahmar S., Naz M., Zafar H., et al. Calcium homeostasis and potential roles to combat environmental stresses in plants. South Afr. J. Bot. 2022;148:683–693. [Google Scholar]
- 74.Huang Y., Wang W., Yu H., Peng J., Hu Z., Chen L. The role of 14-3-3 proteins in plant growth and response to abiotic stress. Plant Cell Rep. 2022;41:1–20. doi: 10.1007/s00299-021-02803-4. [DOI] [PubMed] [Google Scholar]
- 75.Yadav S.K., Singla-Pareek S.L., Reddy M.K., Sopory S.K. Transgenic tobacco plants overexpressing glyoxalase enzymes resist an increase in methylglyoxal and maintain higher reduced glutathione levels under salinity stress. FEBS Lett. 2005;579:6265–6271. doi: 10.1016/j.febslet.2005.10.006. [DOI] [PubMed] [Google Scholar]
- 76.Liu C., Zhao L., Yu G. The dominant glutamic acid metabolic flux to produce γ‐amino butyric acid over proline in Nicotiana tabacum leaves under water stress relates to its significant role in antioxidant activity. J. Integr. Plant Biol. 2011;53:608–618. doi: 10.1111/j.1744-7909.2011.01049.x. [DOI] [PubMed] [Google Scholar]
- 77.Chen J.H., Jiang H.W., Hsieh E.J., Chen H.Y., Chien C.T., Hsieh H.L., et al. Drought and salt stress tolerance of an Arabidopsis glutathione S-transferase U17 knockout mutant are attributed to the combined effect of glutathione and abscisic acid. Plant Physiol. 2012;158:340–351. doi: 10.1104/pp.111.181875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Dubreuil-Maurizi C., Poinssot B. Role of glutathione in plant signaling under biotic stress. Plant Signal. Behav. 2012;7:210–212. doi: 10.4161/psb.18831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Lhamo D., Wang C., Gao Q., Luan S. Recent advances in genome-wide analyses of plant potassium transporter families. Curr. Genom. 2021;22:164. doi: 10.2174/1389202922666210225083634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wang S., Song M., Guo J., Huang Y., Zhang F., Xu C., et al. The potassium channel Fa TPK 1 plays a critical role in fruit quality formation in strawberry (Fragaria× ananassa) Plant Biotechnol. J. 2018;16:737–748. doi: 10.1111/pbi.12824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Musavizadeh Z., Najafi-Zarrini H., Kazemitabar S.K., Hashemi S.H., Faraji S., Barcaccia G. Genome-wide analysis of potassium channel genes in rice: expression of the OsAKT and OsKAT genes under salt stress. Genes. 2021;12:784. doi: 10.3390/genes12050784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kajla M., Yadav V.K., Chhokar R.S., Sharma R.K. Management practices to mitigate the impact of high temperature on wheat. J. Wheat Res. 2015:1–12. [Google Scholar]
- 83.Wang B., Sun Y.F., Song N., Wei J.P., Wang X.J., Feng H., et al. MicroRNAs involving in cold, wounding and salt stresses in Triticum aestivum L. Plant Physiol. Biochem. 2014;80:90–96. doi: 10.1016/j.plaphy.2014.03.020. [DOI] [PubMed] [Google Scholar]
- 84.Rhoades M.W., Reinhart B.J., Lim L.P., Burge C.B., Bartel B., Bartel D.P. Prediction of plant microRNA targets. Cell. 2002;110:513–520. doi: 10.1016/s0092-8674(02)00863-2. [DOI] [PubMed] [Google Scholar]
- 85.Achard P., Herr A., Baulcombe D.C., Harberd N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development. 2004;131:3357–3365. doi: 10.1242/dev.01206. [DOI] [PubMed] [Google Scholar]
- 86.Schwab R., Palatnik J.F., Riester M., Schommer C., Schmid M., Weigel D. Specific effects of microRNAs on the plant transcriptome. Dev. Cell. 2005;8:517–527. doi: 10.1016/j.devcel.2005.01.018. [DOI] [PubMed] [Google Scholar]
- 87.Zhao Y., Xu K., Liu G., Li S., Zhao S., Liu X., et al. Global identification and characterization of miRNA family members responsive to potassium deprivation in wheat (Triticum aestivum L.) Sci. Rep. 2020;10 doi: 10.1038/s41598-020-72642-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Cao J., Gulyás Z., Kalapos B., Boldizsár A., Liu X., Pál M., et al. Identification of a redox-dependent regulatory network of miRNAs and their targets in wheat. J. Exp. Bot. 2019;70:85–99. doi: 10.1093/jxb/ery339. [DOI] [PubMed] [Google Scholar]
- 89.Akpinar B.A., Kantar M., Budak H. Root precursors of microRNAs in wild emmer and modern wheats show major differences in response to drought stress. Funct. Integr. Genomics. 2015;15:587–598. doi: 10.1007/s10142-015-0453-0. [DOI] [PubMed] [Google Scholar]
- 90.Yanfang Y., Kaikai Z., Liying Y., Xing L., Ying W., Hongwei L., et al. Identification and characterization of MYC transcription factors in Taxus sp. Gene. 2018;675:1–8. doi: 10.1016/j.gene.2018.06.065. [DOI] [PubMed] [Google Scholar]
- 91.Srivastava V., Mishra S., Mehrotra S., Upadhyay S.K. Plant Transcription Factors: Contribution in Development, Metabolism, and Environmental Stress. Elsevier Science; 2022. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All the data are available within the manuscript and supplementary files.