Summary
Antimalarial drug resistance and unavailability of effective vaccine warrant for newer drugs and drug targets. Hence, anti-inflammatory activity of phyto-compound (oleuropein; OLP) was determined in antigen (LPS)-stimulated human THP-1 macrophages (macrophage model of inflammation; MMI). Reduction in the inflammation was controlled by the PI3K-Akt1 signaling to establish the “immune-homeostasis.” Also, OLP treatment influenced the cell death/autophagy axis leading to the modulated inflammation for extended cell survival. The findings with MII prompted us to detect the antimalarial activity of OLP in the wild type (3D7), D10-expressing GFP-Atg18 parasite, and chloroquine-resistant (Dd2) parasite. OLP did not show the parasite inhibition in the routine in vitro culture of P. falciparum whereas OLP increased the antimalarial activity of artesunate. The molecular docking of autophagy-related proteins, investigations with MMI, and parasite inhibition assays indicated that the host activated the autophagy to survive OLP pressure. The challenge model of P. berghei infection showed to induce autophagy for circumventing anti-plasmodial defenses.
Subject areas: Health sciences, Drug delivery system
Graphical abstract
Highlights
-
•
Oleuropein (OLP) modulates inflammation in macrophage model of inflammation (MMI)
-
•
Modulated inflammation is controlled by PI3K-Akt1 signaling to establish homeostasis
-
•
OLP treatment influences cell death/autophagy axis for extended cell survival
-
•
P. falciparum employs autophagy to circumvent anti-plasmodial defenses of OLP
Health sciences; Drug delivery system
Introduction
P. falciparum is one of the leading causes of malaria infection and associated mortality and morbidity.1 Severity of infection depends upon the host-pathogen interactions and the immune status of the host. A conserved catabolic process called macroautophagy/autophagy involves the degradation and sequestration of cytosolic components in the lysosomes through the autophagosomes.2 The autophagy promotes adaptation of cells to survive the physiological stress and maintain cellular homeostasis by the cycled amino acids and other macromolecules within the lysosomes.3 The autophagy is activated by the parasite during the physiological stress conditions such as starvation, hypoxia, high temperatures, differentiation, and protein metabolism.3,4 This self-defense mechanism modulates the immune responses and enables host to survive the pathogenic invasion. Parasite survival is extended by the host-induced autophagy under the physiological stress/drug pressure. Autophagy-driven survival of parasite when submitted to drug pressure gives rise to drug tolerance. Hence, understanding the autophagy-mediated survival and evolution of the drug-tolerant parasite may help us better understand antimalarial drug resistance. This shall, in turn, help devising newer drugs and drug targets to treat malaria infection.
Oleuropein (OLP) possessing anti-inflammatory, anti-oxidants, immunomodulatory activity is known to activate autophagy.5,6,7,8 Hence. human THP-1 macrophage model of inflammation (MMI; antigen [LPS]-stimulated macrophages) was developed to confirm the inflammatory and signaling properties of OLP. Also, OLP drives the death of the stimulated macrophages9 controlled by the CD40-driven PI3K-Akt1 signaling pathway. Further, the downstream effector of PI3K-Akt1 (Akt1) is known to regulate the cytoplasmic nuclear factor κB (NF-κβ) and caspase-310,11 in HepG2 and breast cancer cells (MCF-7, MDA-MB-231). Death of antigen-stimulated macrophages programmed by the OLP treatment prompted us to determine its parasite inhibition activity. Interestingly, the OLP treatment influences the cell death/autophagy axis to inhibit the death of antigen-stimulated cells.11 And, immunological balance is tipped toward autophagy like self-defense mechanism employed by the host to survive the OLP pressure. This was confirmed by the protein expression of the co-stimulatory, inflammatory, pro-apoptotic, and autophagy-related protein (ARP) markers in the MMI receiving OLP treatment. OLP-treated and LPS-inflamed macrophages are shown to bypass the death signals controlled by the reduced expression of phosphorylated NF-κB (Rel) that blocks the apoptotic pathways.12 Moreover, binding affinity-based molecular docking analyses showed the interaction of drugs (OLP, artesunate [ART]) with ARP (LC3II, Atg18) and development-related proteins (DRPs) (Kelch13, Akt1). Autophagy employed by the stimulated and OLP-treated macrophages confirmed our molecular docking (protein with the drugs) results. Hence, host-induced autophagy under OLP pressure survives the death and extended the survival.
No reports suggested the antimalarial activity of OLP in P. falciparum. However, only a few reports showed the clearance of the asexual blood stage (BS) infection of P. berghei-infected mice.13 Therefore, we carried out the in vitro assays with the MMI and docked the proteins (DRP & ARP) with ART and OLP. Hence, we determined the antimalarial activity of OLP in the routine culture of the asexual BS infection of P. falciparum. Autophagy activated by the host under the physiological stress or drug pressure could contribute to antimalarial drug resistance.14,15,16 Therefore, we confirmed the starvation-induced autophagy in all three laboratory strains (3D7, D10 parasite expressing GFP-Atg18, Dd2) of P. falciparum.17,18 OLP alone did not show parasite inhibition but increased the antimalarial activity of ART in combination. Indeed, our data suggest the activation of autophagy by the host to circumvent the therapeutic effect of OLP. Gene transcription and protein expression of ARP confirmed the employment of autophagy to evade the OLP pressure. In vitro findings were validated in the challenge model of P. berghei infection. OLP-treated mice showing greater parasite burden compared to the untreated control confirmed the induction of autophagy enabling parasite to circumvent the anti-plasmodial defense.
Collectively, OLP treatment showed the anti-inflammatory and signaling properties confirmed in the antigen-stimulated human macrophages. PI3K-Akt1 signaling could influence the apoptosis/autophagy axis to survive OLP pressure. In vitro findings were confirmed in the rodent model of P. berghei infection that showed to induce autophagy by the host to circumvent the anti-plasmodial defense. Our finding shall have implications in the evolution of antimalarial drug resistance.
Results
OLP modulates inflammation in the antigen (LPS)-stimulated macrophages
OLP is known to attenuate the inflammation in vitro as well as experimental animal models.5,19,20 Therefore, hTHP-1 macrophages (phorbol 12-myristate 13-acetate [PMA]-differentiated monocytes) were stimulated with the antigen (LPS; 1 μg/mL) to induce the acute inflammation. This in vitro model is referred to as macrophage model of inflammation (MMI). The gene expression of inflammatory (interleukin [IL]-1β, tumor necrosis factor alpha [TNF-α], IL-6, iNOS), immunoregulatory (IL-10), and signaling (CD40, Akt1, NF-κB, Bim) mediators was determined in the OLP-treated MMI (supplementary information; Table S1). LPS-stimulated macrophages were treated with OLP in 1) prophylactic (OLP treatment followed by the LPS stimulation) (Figures 1A) and 2) therapeutic (LPS stimulation followed by the OLP treatment) (Figure 1B) mode of treatment. The fold change regulation of relative mRNA expression of pro-inflammatory (IL-1β, IL-6, TNF-α), immunoregulatory (IL-10), anti-oxidant (iNOS-mediated NO production), and the signaling (CD40, AKT, NF-κB, Bim) markers was assessed as compared to that with the untreated control (Figures 1A and 1B). A significant (p < 0.01) reduction in the CD40 expression was correlated well with Akt1-mediated higher expression of NF-κBp50 and Bim. The latter however did not reach the statistical significance (Figure 1A). This reduced inflammation is correlated well with the clearance of the stimulated macrophages in the prophylactic mode of OLP treatment (Figure 1A).21,22 The reduced mRNA expression of pro-inflammatory cytokines (IL-1β, IL-6, TNF-α), iNOS pathway-mediated decreased NO production, and increased expression of the anti-inflammatory/immunoregulatory cytokine (IL-10) suggest the anti-inflammatory potential of OLP (Figure 1A). Further, therapeutic mode of OLP treatment showing the reduced mRNA expression of downstream effectors of PI3K-Akt1 signaling markers CD40 (p < 0.01; significant), AKT (p < 0.001; highly significant), NF-κB (p < 0.01; significant), and Bim (p < 0.1 moderately significant) indicated that the death of antigen-stimulated macrophages was independent of CD40/NF-κB signaling (Figure 1B). Also, the reduction seen in inflammation upon antigenic stimulation was characterized by the significantly (p < 0.01) reduced expression of pro-inflammatory cytokines (IL-1β, IL-6, TNF-α), and iNOS (p < 0.01)-driven decreased NO production. Additionally, enhanced expression of anti-inflammatory/immunoregulatory cytokine (IL-10) (p < 0.1) suggested the immunomodulatory properties of OLP (Figure 1B).
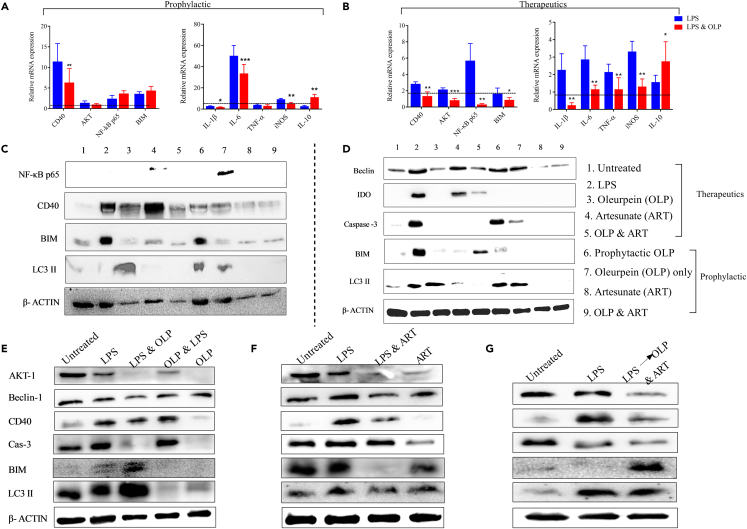
Figure 1.
Determination of immune/signaling mediators in the macrophage model of inflammation
(MMI) (antigen [LPS; 1 μg/mL]-stimulated human macrophages).
(A and B) The human THP-1 monocytes were differentiated to THP-1 macrophages by PMA (100 ng/mL) treatment. Quantification of relative mRNA expression of signaling (CD40, Akt, NF-κB, Bim) (left panel) and inflammatory (IL-1β, IL-6, TNFα, iNOS, IL-10) (right panel) markers in OLP (30 μg/mL)-treated macrophages in (A) prophylactic and (B) therapeutic mode of drug treatment. The protein expression of NF-κB, CD40, Bim, LC3II, IDO1, Beclin-1, caspase-3, and LC3 II was assessed in the OLP- and ART (30 μg/mL)-treated macrophages in both the therapeutic and prophylactic mode of drug treatment (C, D, E, F, G).
(D) Lane 6: prophylactic OLP treatment, Lane 7: OLP treatment. The individual (OLP, ART)- and combination of drugs (OLP & ART)-treated cells were lysed, and proteins were detected by the immunoblotting. The β-actin served as a loading control, and data represent three independent experiments (n = 3) (∗∗∗p < 0.001; highly significant, ∗∗p < 0.01; moderately significant, ∗p < 0.1 significant).
The transcript-level expression of immune mediators was validated by assessing the protein expression through immunoblotting. Protein expression of the PI3K-Akt signaling pathway controlled by the co-stimulatory CD40 in OLP (prophylactic and therapeutic)-treated macrophages (stimulated by the antigen) was determined (Figures 1C and 1D). ART-treated macrophages express CD40 whereas reduced expression was seen by OLP treatment following the prophylactic mode (Figure 1C). Pro-apoptotic protein (Bim) expression was seen independently of CD40 expression in the prophylactic mode of OLP treatment (Figure 1C), whereas stimulated macrophages express caspase-3 upon treatment with OLP prophylactically. It implies that OLP treatment dismantled the cells by programming the cell death of the infectious and inflammatory diseases23 (Figures 1D and 1E). Further, IDO1 in response to the inflammation is known to develop immune tolerance.24 Therefore, macrophages secret heme containing immunoregulatory enzyme Indoleamine 2,3-dioxygenase (IDO1) under ART pressure (Figure 1D). We however did not see the IDO1 expression by the activated macrophages prophylactically treated with OLP (Figure 1D). The equilibrium between the death and survival of antigen-stimulated cells is important to maintain immune-homeostasis.25 Hence, we determined the protein expression of death and autophagy mediators in the stimulated macrophages (Figure 1E, 1F, and 1G). Hardly any difference was seen in the expression of Beclin (BECN1) in the prophylactic and therapeutically treated macrophages with ART & OLP (Figure 1E, 1F, and 1G). OLP treatment showed the induced expression of ARP (LC3II) by the stimulated macrophages (Figure 1E), whereas faint band of LC3 II was seen with ART treatment (Figure 1F). Interestingly, treatment with the combination of ART and OLP exhibited the quantifiable expression of LC3II compared to that with the untreated control (Figure 1G). The gene transcription and protein expression confirmed that OLP influences the death/autophagy axis and hence confers the extended survival on the cells to establish the immune-homeostasis.
OLP activates autophagy in the antigen-stimulated macrophages
Autophagy, a conserved process, confers the survival on cells submitted to the cellular or physiological stress.26,27 The lysosomal turnover of autophagosomal marker LC3 II was shown to induce autophagy in response to the starvation.28,29 Therefore, we decided to perform the confocal microscopy-based immunofluorescence assay (IFA) to detect the expression of ARP to attest the activation of autophagy by the LPS-stimulated macrophages. The induced autophagy by the macrophages enables parasite to stand the pressure of individual (OLP, ART) and combination (OLP & ART) drug treatment (Figures 2 and 3). The qualitative and quantitative (mean fluorescence intensity; MFI) expression of LC3 II & Beclin (Figure 2) and NF-κB & NF-κBp (Figure 3) was determined in the drug-treated macrophages. We observed the significantly (p < 0.0001) higher expression of LC3 II with the OLP treatment compared to the ART-treated macrophages (Figures 2A and 2B). Treatment with the combination of OLP and ART exhibited significantly higher (p < 0.0001) expression of LC3 II (Figures 2A and 2B), whereas no difference was seen in the expression of Beclin/survivin (mammalian ortholog of yeast Atg6) (Figures 2C and 2D). Further, NF-κβ is known to regulate the transcription of genes that control inflammation, development of the immune effector, cell cycle, proliferation, and cell death.30 Therefore, we determined the qualitative (Figures 3A and 3C) and quantitative (Figures 3B and 3D) expression of the nuclear transcription factor (NF-κB & NF-κBp65) in the drug-treated macrophages stimulated with LPS. Higher expression (p < 0.001) of NF-κBp65 was seen in macrophages receiving treatment with the drugs (OLP, ART) as well as their combination (OLP & ART) (Figure 3B), while lesser expression of un-phosphorylated NF-κB was seen with the individual (OLP, ART) (p < 0.001) and combination (OLP & ART) (p < 0.001) of drugs-treated macrophages as compared to the untreated control (Figures 3C and 3D). Our results are in agreement with previously published31,32 findings wherein phosphorylated NF-κB is shown to enter the nucleus upon co-stimulation with CD40. The latter regulates FOXO1 for the controlled expression of pro-apoptotic protein (Bim) and thereby induced cell death.
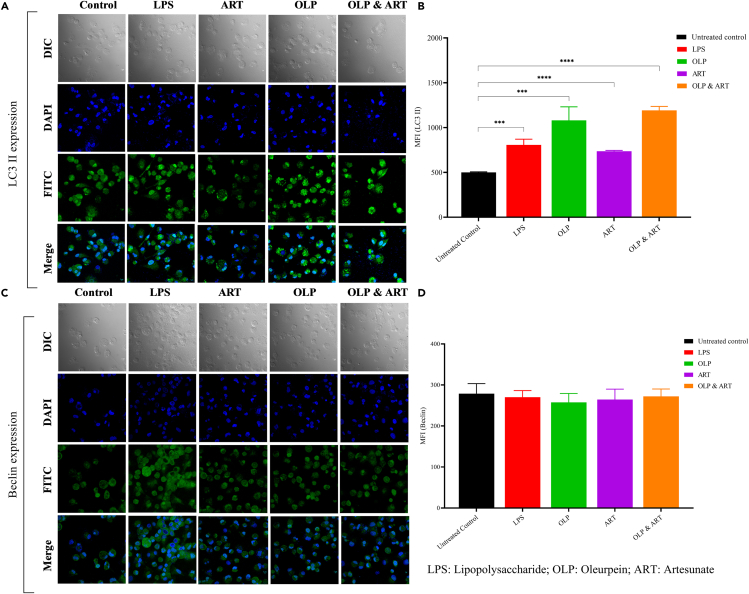
Figure 2.
Expression of autophagy-related protein markers by the confocal microscopy-based immunofluorescence assay
(A) Qualitative and (B) quantitative (mean fluorescence intensity; MFI) expression of LC3 II in the individual (OLP, ART)- and combination of drugs (OLP & ART)-treated and antigen-stimulated macrophages. The qualitative expression of (C) Beclin-1/surviving protein was observed to determine the drug-induced autophagy in the macrophages. Beclin-1 binds phosphatidylinositol 3-kinase (PI3KC3) to form a core PI3KC3 complex that mediates multiple vesicle-trafficking processes to regulate the autophagy.
(D) Quantification (MFI) of Beclin-1 expression did not show any difference with individual and combination drug treatment. The data represent three independent experiments (n = 3). (∗∗∗∗p < 0.0001, highly significant; ∗∗∗p < 0.001, significant).
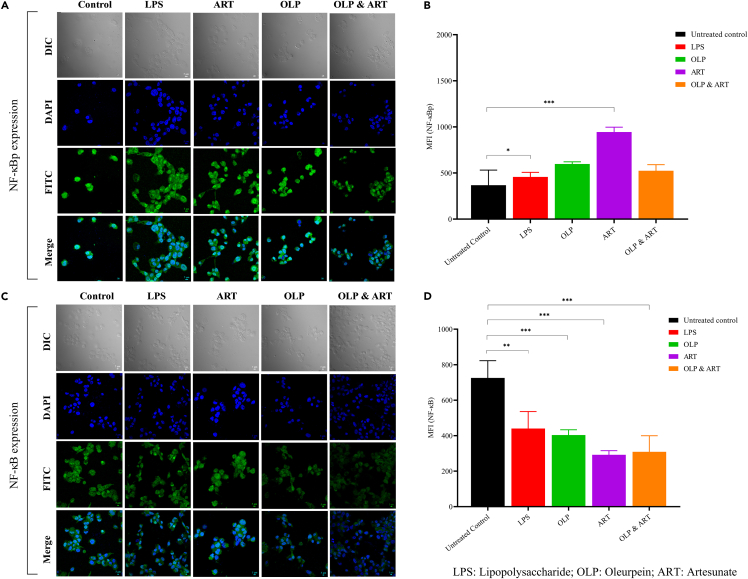
Figure 3.
Drug treatment influences the cell death/autophagy axis and controls the death of macrophages receiving stimulus with antigen
(A–D) Cell death is regulated by the nuclear transcription factor (NF-κBp65 & NF-κB); (A) qualitative and (B) quantitative (MFI) expression of NF-κBp65 in the individual (OLP, ART)- and combination of drugs (OLP & ART)-treated and antigen-stimulated macrophages. The expression of un-phosphorylated NF-κB was determined (C) qualitatively and (D) quantitatively (MFI). The reduced NF-κB expression leads to the reduced cell death and promotes autophagy. NF-κB gets dissociated from its inhibitor (Ikβ) and regulates FOXO1 in the nucleus. The latter controls the expression of Bim.31,32 Death of the antigen stimulated macrophages by binding the phosphatidylinositol 3-kinase (PI3KC3) to form a core PI3KC3 complex that mediates the multiple vesicle-trafficking processes and regulates the autophagy complex. The data represent three independent experiments (n = 3) (∗∗∗p < 0.001, highly significant; ∗∗p < 0.01, significant; ∗p < 0.1, moderately significant).
OLP attenuates free radicals (reactive oxygen species [ROS]) production to escape cell death
OLP exerts the pro- and anti-oxidant properties in a dose-dependent manner in the ovarian cancer cells.33 Higher dose of OLP maintains the delicate balance between cell death and survival, and inhibition in the cell survival is controlled by the antigen stimulation. These findings prompted us to determine the ROS production in the LPS-stimulated macrophages receiving treatment with individual (OLP, ART) and combination of drugs (OLP & ART) (Figure 4A). OLP treatment showed inhibited ROS production compared to that seen with the ART treatment as well as combination of OLP & ART (Figure 4A). ROS production (Annexin V staining) investigation encouraged us to determine the drug-treated macrophages stimulated with the antigen (Figure 4B). Higher cell death (∗p < 0.1) (% Annexin-positive cells) was seen in the cells treated with the combination of drugs compared to those receiving treatment with the individual drugs (OLP, ART) and the untreated control (Figure 4B). Treatment with ART and combination of OLP & ART induced the death of stimulated cells confirmed by the statistically significant (∗p < 0.1) Annexin-positive cells. Our data suggested the shift toward right in the flow cytometry-based histogram during cell count analysis (Figure 4C). Also, the scattered plots of Annexin-positive/negative cells were stained with propidium iodide (PI) (Figure 4D) to confirm the death of the macrophages.
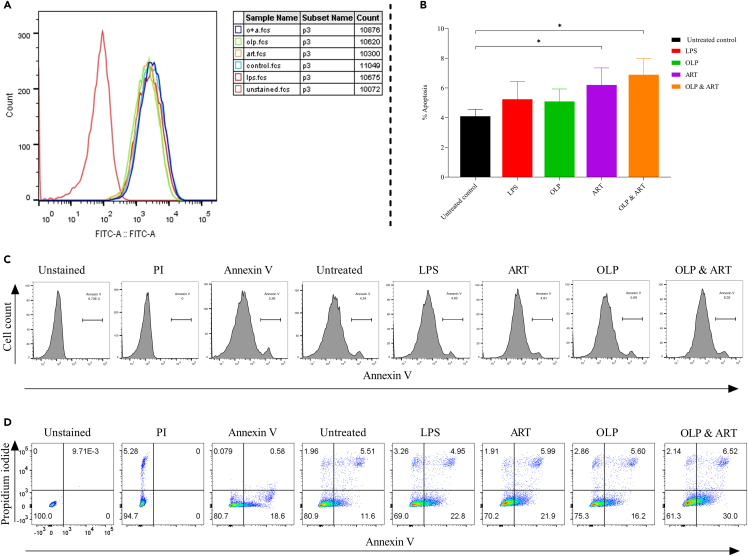
Figure 4.
Oleuropein mediates the survival of the antigen-stimulated human macrophages
(A and B) Cells receiving treatment with individual (OLP, ART) and combination of drugs (OLP & ART) to assess (A) reactive oxygen species (ROS) production and (B) detection of % cell death of LPS-stimulated macrophages by the fluorescently labeled Annexin V staining. 7-aminoactinomycine (7-AAD) or propidium iodide (PI) staining by the C6 Accuri flow cytometer (∗p < 0.05 [significant]).
(C) Histogram of Annexin V and FITC/PI flow cytometry to confirm the cell death of treated and untreated cells.
(D) The scatterplots of annexin V-FITC/PI-stained cells treated with drug (OLP, ART) individually as well in combination. The data represent three independent experiments (n = 3).
Drug and protein interaction and activation of the autophagy
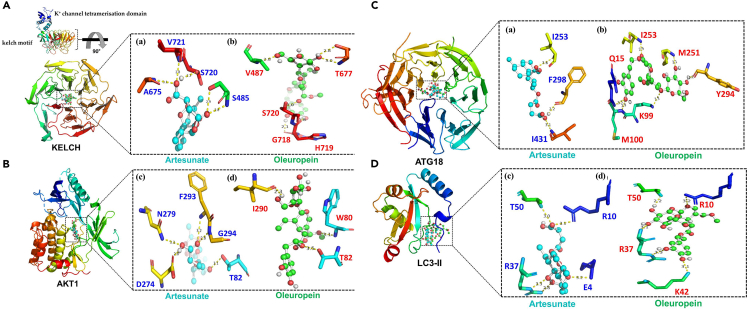
Kelch protein contains K+ channel tetramerization domain and kelch motif. K+ channel tetramerization domain is the N-terminal (distinct group of ion channel family) that establishes and maintains the ionic homeostasis. Kelch motif is constructed from the beta-sheets that fold together to form a large circular solenoid beta-propeller domain. The latter forms homo- or hetero-dimers in order to mediate the protein-protein interaction. OLP (−10.3 kcal/mol) and ART (−10.1 kcal/mol) (Figure S1) showed the similar binding affinity and bind to the active site region (Figure 5). ART binds to the S485, A675, S720, and V721 (Figure 5Aa) amino acids of kelch motif whereas OLP binds to the V487, G718, H719, S720, and T677 amino acids (Figures 5Ab). Our molecular docking analysis suggested that OLP may be an alternative to ART especially against the Kelch protein expressed by the P. falciparum.34
Figure 5.
Binding affinity-based molecular docking analysis to confirm the expression of autophagy-related protein (ARP) upon the drug-protein interaction
The interaction of ART and OLP with the development (A) (Kelch13) (a, b) and cell death/survival (B) (AKT1) (c, d) protein markers. Black box is magnified to show the interaction of the amino acid residues with ART and OLP. ART (cyan) and OLP (green) shared the similar binding region with both the proteins (AKT1, KELCH) (shown in the ball and sticks). The yellow dotted line represents the formation of the hydrogen bond among the selected residues to the corresponding ligands/drugs.
Molecular docking interaction with P. falciparum-specific (C) (PfATG18) (a, b) and autophagy-related protein (D) (LC3 II) (c, d). Black box is magnified to show the amino acid residues showing interaction with ART and OLP. ART (cyan) and OLP (green) shared the similar binding region with both the proteins (ATG18 and LC3II) (shown in the ball and sticks). The yellow dotted line represents the formation of the hydrogen bond among the selected residues with the corresponding drugs.
The investigation for the anti-inflammatory, immunomodulatory, and signaling properties in the MMI (Figure 1) led us to examine the interaction of Akt1 (protein) with the ligands (ART, OLP). Akt1 is a serine/threonin-protein kinase enzyme comprising PH (Pleckstrin homology) and SH2 (Src homology 2-like) domains. Highly conserved PH is involved in the intracellular signaling whereas SH2 domain supports the signal transduction of receptor tyrosine kinase pathways. A critical mediator of the growth factor (Akt1) gets activated by the phosphatidylinositol 3-kinase (PI3K). Cells employ autophagy driven by the activated Akt1 to circumvent the drug pressure and proliferate. In addition, other factors suppress the inactive Akt1 controlling the cell death in a transcription-independent manner. Hence, a delicate balance between the death and cell survival is maintained to achieve the physiological homeostasis. Further, our molecular docking data suggest that ART and OLP share the similar binding pocket of Akt1 for their interactions (Figure 5B). ART binds the region with the higher binding affinity (−10.8 kcal/mol) with T82, D274, N279, F293, and G294 amino acids (Figure 5Bc; Figure S1) whereas OLP exhibited binding with lower-affinity (−9.8 kcal/mol) (T82, W80 and, I290) amino acids (Figures 5Bd; Figure S1). We observed the higher binding affinity of ART that leads to the stronger binding with the Akt1 than OLP.
Autophagy maintains cellular and tissue homeostasis and galvanizes metabolic and immunologic adaptation in response to a highly diverse plethora of stress-inducing agents. ATG18 is an essential protein required for autophagy, vacuole homeostasis, and endosomal functions to establish physiological homeostasis. It is basically a β-propeller, shaped by seven beta-transducing repeat/WD40 domains with conserved Phe-Arg-Arg-Gly (FRRG) motif.35 Our investigation suggested that ART and OLP bind to the similar pocket present in the ATG18 (Figure 5C) with slightly higher propensity of binding affinity of ART (−9.9 kcal/mol) (Figure 5Ca) than OLP (−9.2 kcal/mol) (Figure 5Cb). However, OLP binds to ATG18 with higher number of amino acids (Q15, K99, M100, I253, M251, and Y294) (Figure 5Cb) than ART (I253, F298, I431) (Figure 5Ca) showing the comparable binding affinity.
LC3 (Microtubule-associated protein 1A/1B-light chain 3) is a soluble protein distributed ubiquitously. A cytosolic form (LC3-I) conjugates with phosphatidylethanolamine (PE) (a phospholipid) to form membrane-bound LC3II complex during the activation of autophagy. Succinctly, the intracellular isolation of a double-membrane complex occurs and enwraps specific and selected cargo for the degradation (auto-phagosome formation) upon its fusion to the lysosomes (autophagolysosome). Consequently, it leads to the digestion of previously selected cargo and promotes the nutrients recycling of the organelle turnover. Molecular docking analysis performed with ART and OLP showed that LC3II shares similar binding region for ART & OLP with better binding affinity of ART (−8.4 kcal/mol) due to its small size (Figure 5Dc). Our analysis suggests that R10, R37, and T50 amino acids are common in both the ligands and their interaction with LC3II. E4 interacts with ART, and K42 and R10 with OLP (Figure 5Dd).
Autophagy is crucial for the reinvasion and replication of P. falciparum
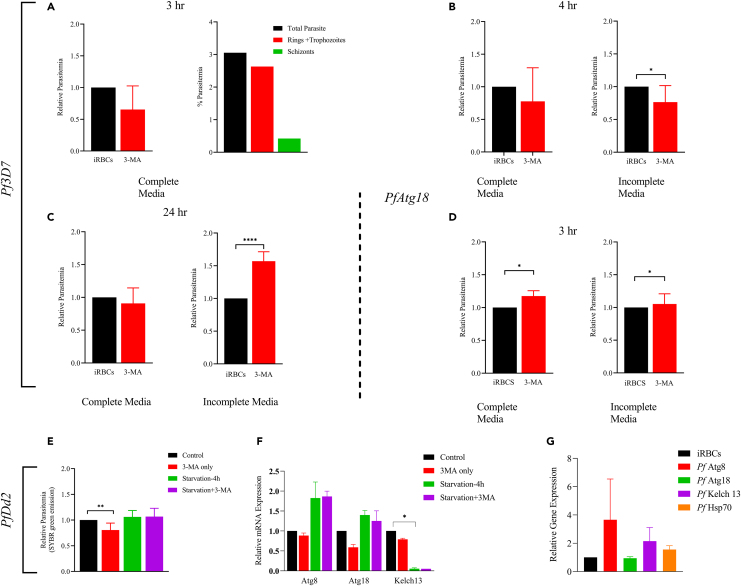
The basal autophagy maintains cellular homeostasis and protects cells from physiological stress in all living organisms including human parasite, P. falciparum.18,36 Therefore, we decided to decipher the role of autophagy employed by the P. falciparum (Pf3D7) to survive the OLP pressure. Parasite was cultured in the Albumax-deficient medium followed by the treatment with 3-MA (autophagy inhibitor) to confirm the activation of autophagy. A comparative assessment of the relative parasitemia was done in order to assess the parasite grown in the nutrient-deficient (incomplete) and nutrient-sufficient (complete) medium (Figure 6). Parasite was starved for 3, 4, and 24 h (Figures 6A, 6B, and 6C) followed by the 3-MA treatment when parasite had 95% schizonts population (Figure 6A). We saw nearly 50% inhibition in the parasite growth (p < 0.01 and ∗∗∗∗p < 0.0001) than untreated control in the SYBR green-based fluorescence assay. 3-MA treatment showed to reduce rings by 40%–50% confirmed by the % parasitemia read on the Giemsa-stained thin blood smears (Figure 6A, right panel). A decrease seen in the relative parasitemia (Figures 6A, 6B, and 6C) (p < 0.01, and ∗∗∗∗p < 0.0001) indicated the employment of the self-defense mechanism by the P. falciparum during the physiological stress. As others,18 we confirmed the inhibited reinvasion of starved P. falciparum when treated with 3-MA. Also, we confirm that P. falciparum activates a “defense” mechanism to survive the physiological stress (Figures 6A, 6B, and 6C).
Figure 6.
Activation of autophagy in response to starvation and drug pressure
(A–C) The asexual BS infection of 3D7P. falciparum was cultivated in the complete (with Albumax) and incomplete (without Albumax) medium for 3 h (A), 4 h (B), and 24 h (C). Parasite cultured in incomplete medium was treated with 3-MA (3 mM), and infectious parasite load (relative parasitemia) was determined by the SYBR green emission assay.
(D) Quantification of parasite load of D10 parasite expressing GFP-Atg18 line of P. falciparum cultivated in the complete and incomplete medium for 3 h and treated with 3-MA. Confirmation of autophagy in chloroquine-resistant (Dd2) P. falciparum under physiological stress.
(E) Quantification of parasite load in the starved (3 h) parasite followed by the 3-MA treatment.
(F) Quantification of the relative gene expression (fold change regulation) of the autophagy and tolerance markers (Atg8, Atg18, Kelch13) by the RT-PCR in the starved (4 h) Dd2 parasite.
(G) Expression profiling of Atg8, Atg18, Kelch13, and Hsp70 in the Dd2 P. falciparum by RT-PCR. (∗p < 0.05 [significant], ∗∗p < 0.01 [moderately significant], and ∗∗∗∗p < 0.0001 [highly significant]). Data represent two independent experiments in triplicates (n = 2).
We next decided to confirm the induction of autophagy in D10 parasite expressing GFP-Atg18 line of P. falciparum (Figure 6D). 3-MA-treated starved parasite did not show the ensuing invasion as shown by the relative parasitemia determined by the SYBR green emission assay (Figure 6D). Interestingly, GFP-Atg18 line of P. falciparum upon starvation showed a significant (∗p < 0.05) increase in parasitemia even when treated with 3-MA.
We then sought to determine the starvation-induced autophagy in the chloroquine (CQ)-resistant P. falciparum (PfDd2) (Figure 6E). The pattern of growth and inhibition in the relative parasitemia in the starved Dd2 was seen to be similar to that in the wild type (Pf3D7) as it showed an increase in the parasitemia which was inhibited (∗∗p < 0.01) following the treatment with the 3-MA (Figure 6E).
We quantified the fold change regulation of autophagy markers (Pfatg8, Pfatg18, Pfkelch13) (primer information; Table S2) when parasite was starved followed by the 3-MA treatment (Figure 6F). Non-significant higher expression of atg8 and atg18 was seen in Dd2 (Figure 6F). It may be attributed to the activation of the autophagy-like self-defense mechanism employed by the parasite to survive the starvation-induced stress. Further, a significantly reduced expression of kelch13 following the 3-MA treatment is suggestive of the activation of autophagy. This implies that reinvasion of parasite induced by autophagy was inhibited by 3-MA (Figure 6F). The extended starvation (24 h) exhibited higher expression of atg8, atg18, and kelch13 in Dd2 P. falciparum. This higher expression could however not reach the statistical significance. Also, no detrimental effect was seen on parasite growth and development (Figure 6G).
Antimalarial activity of drugs
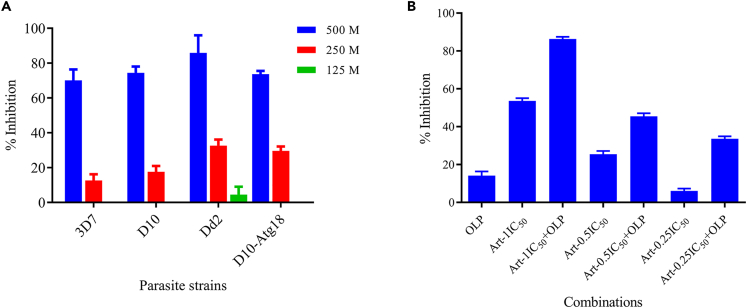
We assessed the OLP and antimalarials (artemisinin and CQ) for inhibition of the asexual erythrocytic stage development of P. falciparum (3D7, D10, Dd2, D10-Atg18) strains. 3D7 and Dd2 are susceptible and moderately resistant strains to CQ,37 respectively. D10 is susceptible to routinely used antimalarials, and D10-Atg18 is D10-based Atg18 expression strain. Artemisinin inhibited the asexual erythrocytic development of all the strains (Table 1), with slightly higher half-maximal inhibitory concentration (IC50) for Dd2 and D10-Atg18 strains as compared with the 3D7 and D10 strains. CQ inhibited Dd2 with much higher IC50 concentrations than those for 3D7, which is consistent with the reported resistant phenotype of Dd2.37 OLP did not completely inhibit the development of any of the parasite strains in the 1st cycle even at 500 μM but showed >75% inhibition of all the strains at 250 μM when parasite incubation with drugs was extended up to 96 h (Figure 7A) indicating that OLP is similarly effective against all the parasite strains used in this study. Although OLP did not have any antimalarial effect in 1 cycle even at 500 μM, it increased antimalarial activity of artemisinin in 1 cycle inhibition assay (Figure 7B), indicating that artemisinin-OLP combination has additive antimalarial effect. This additive effect of OLP may be due to increased autophagy, which has been associated with hemoglobin catabolism in Plasmodium, which provides iron for artemisinin activation.38
Table 2.
Docking of proteins involved in anti-inflammatory, anti-malarial, and PI3K-Akt signaling activity of oleuropein and antimalarial drug, artesunate
| Protein name | Binding affinity (Kcal/mol) |
|
|---|---|---|
| Artesunate | Oleuropein | |
| AKT1 | −10.8 | −9.8 |
| KELCH | −10.3 | −10.1 |
| ATG18 | −9.9 | −9.2 |
| ATP_SYN | −9.0 | −9.6 |
| MDR1 | −8.5 | −8.1 |
| LC3-II | −8.4 | −7.5 |
| HSP70 | −7.8 | −7.2 |
| IL-10 | −7.8 | −7.2 |
| IL-1β | −7.3 | −7.2 |
| NHE | −7.3 | −8.3 |
| TNF | −7.1 | −6.9 |
| BIM | −7.0 | −7.1 |
| NF-κB | −7.0 | −5.8 |
| IL-6 | −6.6 | −7.0 |
| ATG8 | −6.5 | −6.4 |
| CD40 | −6.1 | −6.4 |
Figure 7.
Effect of OLP and combination of OLP and artemisinin on parasite development
(A) P. falciparum 3D7, Dd2, D10, and D10-Atg18 strains were cultured in presence of indicated compounds for 96 h. Each bar shows mean % inhibition with SD error bar at the indicated OLP concentration from three experiments, each with two replicates.
(B) P. falciparum 3D7 were cultured in the presence of artemisinin, OLP, or artemisinin-OLP combination for 50 h. Each bar shows mean % inhibition with SD error bar at different concentration of indicated compounds or artemisinin-OLP combination from three experiments, each with two replicates.
P. falciparum sensitivity and drug toxicity assay
Anti-inflammatory and signaling activity determined in the MMI (Figures 1, 2, 3, and 4) led us to determine parasite clearance and SYBR and DAPI emission when the routine culture of the asexual BS infection of P. falciparum was treated with different concentrations of OLP (Figures S2A, S2B, S2C, and S2D). Also, we decided to determine the toxicity of OLP by MTT assay at 40, 80, 160, and 320 nM in the HepG2 cells (Figure S2E). ART-treated HepG2 cells were considered as a positive control. We observed the similar cell viability upon receiving treatment with OLP and ART (Figure S2E). Our data confirmed the safety of OLP for humans.
P. falciparum survives the drug pressure by the host-activated autophagy and overcomes the anti-plasmodial defenses elicited by the host. We characterized the autophagy-like escape mechanism at gene transcription (Figure 1) and protein levels (Figures 1, 2, 3, and 4) of autophagy mediators, as well as PI3K-Akt signaling pathway in the stimulated macrophages. Our data suggest that the involved CD40-driven signaling during OLP pressure may confer the survival on the parasite (CD40) (Figures 1 and 2). Therefore, we decided to conduct the gene expression profiling to attest that the parasite survival mechanism(s) employed could circumvent the anti-plasmodial defenses under OLP pressure.
Molecular profiling
We determined the relative expression of development-, survival-, and autophagy-related markers (kelch13, mdr1, nhe, atg8, atg18, hsp70, and Atp synthase) (primer; Table S2) in the laboratory strains of P. falciparum (3D7, D10-Atg18, Dd2) (Figures 8, S3, and S4). We determined the gene transcription profiling with the individual (OLP, ART) (Figures S3 and S4) and combination (OLP & ART) drug treatment (Figure 8) in 3D7, D10-Atg18, and Dd2 strains.
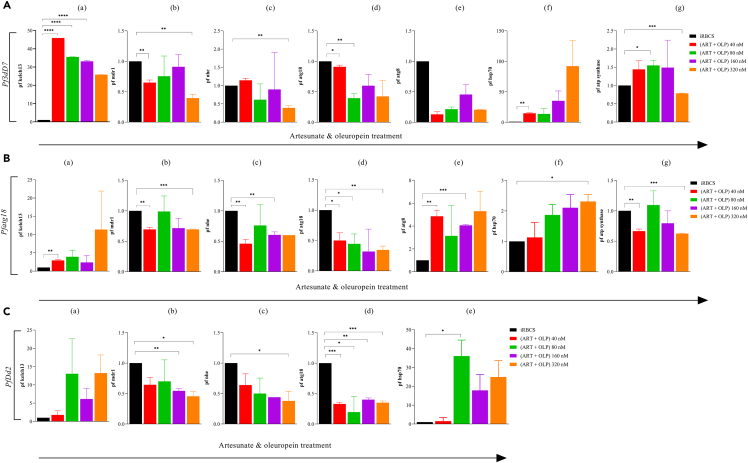
Figure 8.
Determination of gene expression profiling of growth (Kelch13, Atp synthase), resistance/tolerance (mdr1, nhe), and autophagy (Atg8, Atg18) markers under drug pressure by RT-PCR
(A–C) The laboratory strains (A) 3D7 and (B) D10 parasite expressing GFP-Atg18 line and (C) chloroquine-resistant (Dd2) of P. falciparum were treated with the combination of drugs (OLP & ART) at 40, 80, 160, and 320 nM concentrations. iRBCs were taken as an experimental/negative control. The data represent two independent experiments in triplicates (n = 2). (∗p < 0.05 [significant], ∗∗p < 0.01 [moderately significant], and ∗∗∗p < 0.001 [highly significant]).
Gene expression profiling in 3D7
Treatment with the combination of OLP and ART exhibited significantly (p < 0.001) higher expression of Kelch13 at 40, 80, and 160 nM than the infection control. ART treatment did not show the Kelch13 expression in 3D7 (Figure S3Aa) whereas OLP treatment especially at higher concentrations (160 nM, 320 nM) exhibited its expression (Figure S3Ba) that could not reach the statistical significance. Further, the inhibited expression of multi-drug resistance (mdr1)39 (p < 0.001) and Na+/H+ exchanger (nhe)39,40,41 (p < 0.01) by the treatment with the combination of drugs (Figures 8Ab, c) and ART (Figure S3Ab, c) was observed, whereas OLP-treated 3D7 showed the increased expression of mdr1 and inhibited (p < 0.001) expression of nhe (Figure S3Bb, c). Autophagy (atg18, atg8) markers showed the altered expression in the 3D7 parasite treated with the combination of ART and OLP (Figures 8Ad, e). The reduced expression of atg18 was seen with the combination drug regimen (Figures 8Ad). Our finding is of particular relevance since expression of atg18 is shown to facilitate the PI3P protein that plays a crucial role in the expression of Akt1 regulated by the PI3K signaling pathway responsible for the autophagosome formation.42,43 Hence, individual treatment with ART (Figure S3Ad) and OLP (Figure S3Bd) showed increased atg18 expression. The reduced expression of Atg8 with the individual treatment of ART or OLP (Figures S3Ae, Be) and combination of ART & OLP (Figure 8Ae) suggests the activation of Atg8-independent autophagy. Hsp70 is known to support the virulent cyto-adherence and parasite growth during the harsh temperature conditions.44 Our data showed higher expression of hsp70 at all the concentrations of the combination of ART and OLP tested (Figure 8Af). On the contrary, significantly reduced (p < 0.001) expression was seen with ART treatment (Figure S3Af) and reduced expression was seen with the OLP-treated 3D7 (Figure S3Bf). Additionally, we assessed the survival marker45 (Pfatp synthase) (Figure 8Ag) expression in response to the treatment with the combination of drugs. It showed increased expression of Pfatp synthase (p < 0.05) (Figure 8Ag). We saw a significantly higher (p < 0.001) expression of Atp synthase in the ART-treated parasite (Figure S3Ag). Moreover, non-significant increase in the expression of this survival marker in OLP-treated 3D7 (Figure S3Bg) could suggest the employment of autophagy.
Gene expression profiling in D10-Atg18 and Dd2 P. falciparum
We determine the gene expression profiling in the D10-expressing Atg18 falciparum (Figures 8B and S4). We saw the molecular patterns of kelch13, mdr1, and nhe similar to those of the 3D7 strain. Also, treatment with ART showed the mixed expression of kelch13, mdr1, and nhe (Figures S4Aa, b, c). A significant (p < 0.001) reduction in the kelch13 expression was seen in the OLP-treated Atg18 parasite compared to the infection control (Figure S4Ba). D10 parasite expressing Atg18 showed a decreased expression of Atg18 with the drug combination treatment (Figure 8Bd) and individual drugs (ART, OLP) (p < 0.01) (Figures S4Ad, Bd). Significantly enhanced Atg8 expression was seen in the Atg18 parasite upon receiving treatment with combination drug (OLP & ART) treatment (Figure 8Be) and ART treatment (Figure S4Ae). Significantly higher (p < 0.001) expression of Atg8 expression was seen in the OLP-treated parasite (Figure S4Be). The hsp70 (p < 0.05) expression was seen to be significantly elevated with the treatment of the combination of drugs (Figure 8Bf) as well as individual drug (ART, OLP) (p < 0.01)-treated Atg18 parasite (Figures S4Af, Bf). Besides, decreased expression of Atp synthase in Atg18 was observed (Figure 8Bg, Figures S4Ag, Bg) (p < 0.001). In the end, we observed the gene transcription patterns in the Dd2 parasite similar to those of the 3D7 P. falciparum with the combination (Figure 8C) and individual drug treatment (Figures S5A and S5B).
P. falciparum employs autophagy to circumvent anti-plasmodial defenses
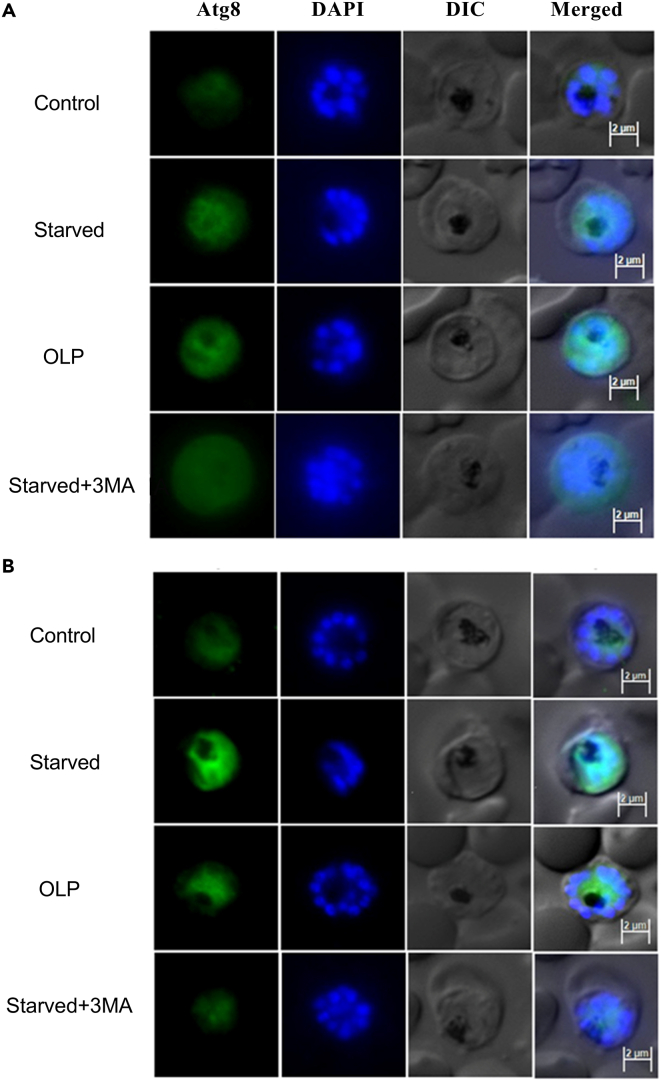
Our findings with the MMI suggested that antigen-stimulated macrophages induce autophagy to survive the OLP pressure (Figures 1, 2, 3, and 4). Also, molecular docking of the ARP and DRP with the drugs (OLP, ART) showed the binding affinity-based interaction (Figure 5). Also, starvation-induced autophagy and gene transcription analysis (Figure 6) drove us to confirm the employed autophagy under the physiological stress/drug pressure in 3D7 (Figure 9A) and D10-Atg18 P falciparum (Figure 9B). In contrast to the inhibition data, treatment of 3D7 and D10-Atg18 trophozoite with non-inhibitory concentration of OLP (320 nM) for 3 h increased Atg8 signal similar to starved parasites when compared to the parasites grown in the complete medium or treated with autophagy inhibitor, 3-MA (Figure 9). These data are consistent with the reported autophagy-inducing property of OLP.3,46
Figure 9.
Atg8 expression in P. falciparum
(A and B) Synchronized late trophozoite stages P. falciparum 3D7 (A) and P. falciparum D10-Atg18 (B) parasites were cultured in complete medium (control), Albumax-deficient medium (Starved), and complete medium with OLP or Albumax-deficient medium with 3-MA for 3 h. The parasites were processed for Atg8 expression using anti-Atg8 antibodies. The panels are for Atg8 expression (Atg8), nucleic acid stain (DAPI), bright field (DIC), and the overlap of all three images (Merged). Scale bar has been shown in the merged panel.
In vivo challenge model of P. berghei infection
tReduced inflammation seen in the OLP-treated stimulated macrophages and parasite inhibition activity in the routine culture of the asexual BS infection of P. falciparum suggested that OLP possesses the anti-inflammatory, controlled signaling and possibly parasite inhibition activity (Figures 1, 2, 3, 4, and 5). Also, LPS-stimulated macrophages overcome the death of stimulated macrophages upon OLP treatment (Figure 4). Transcript and protein level expression of molecular markers in MMI and molecular docking analyses is suggestive of the employment of autophagy-like escape mechanism(s). The activation of autophagy was confirmed in the laboratory strains (3D7, D10-Atg18, Dd2) of P. falciparum in response to the starvation (Figure 6). Further, drug sensitivity assays performed with the routine culture of asexual BS infection of 3D7, Atg18, and Dd2 showing the parasite clearance activity (Table 1; Figure 7), gene transcription profiling (Figures 8, S3, S4, and S5), and protein expression of atg8 (Figure 9) confirmed the activation of autophagy under the drug pressure.
Table 1.
Antimalarial activity of artemisinin and chloroquine
| Parasite strains | Compounds |
|
|---|---|---|
| Artemisinin | Chloroquine | |
| P. falciparum 3D7 | 17.7 ± 3.4 nM | 13.6 ± 2.5 nM |
| P. falciparum D10 | 17.3 ± 2.2 nM | ND |
| P. falciparum Dd2 | 45.6 ± 3.6 nM | 199.0 ± 23.9 nM |
| P. falciparum D10-Atg18 | 29.6 ± 5.0 nM | ND |
IC50 concentrations of artemisinin and chloroquine for inhibition of asexual erythrocytic stage development of different P. falciparum strains. ND indicated “not determined”. The values are mean of 2–3 independent experiments with SD.
Therefore, we developed a challenge model of P. berghei infection to validate the in vitro findings. P. berghei is used as surrogate for P. falciparum.47 Infected BALB/c mice were divided into 7 groups (n = 6). The level of parasitemia reached nearly 5% (Scheme 1) before mice were intraperitoneally injected with CQ (30 and 100 mg/kg), ART (30 mg/kg), and OLP (30, 100, and 500 mg/kg). The drawn thin blood smears were read to determine % parasitemia (Figures 10A, S6, and S7). The parasite was completely cleared from the mouse periphery by the CQ treatment at both the doses tested (30 & 100 mg/kg), whereas ART cleared the parasite significantly but not completely from mouse periphery (Figures 10A, S6, and S7). This delay in parasite clearance by the ART is consistent to our previous finding wherein experimental induction of ART resistance in the P. falciparum-infected immunodeficient (humanized) mice was shown.15 The OLP treatment saw a reduced parasite load for only a few days followed by activation of autophagy-like escape mechanism to circumvent the anti-plasmodial defenses mounted by the host. Therapeutic effect of OLP was evaded by the parasite. This leads to the increase in the parasitemia at all the concentrations tested, but a sharp rise (p < 0.001) in the parasitemia was seen with 30 mg/kg OLP (Figure 10A) than that seen with the untreated control. This rise in the parasitemia could be attributed to the activation of host autophagy to resist the parasite clearance by OLP (Figure 10A).
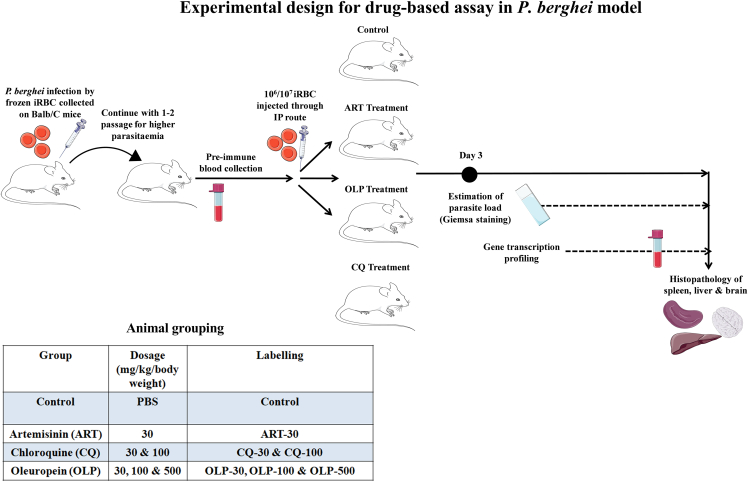
Scheme 1.
Experimental design for drug-based assays in the challenge model of P berghei infection
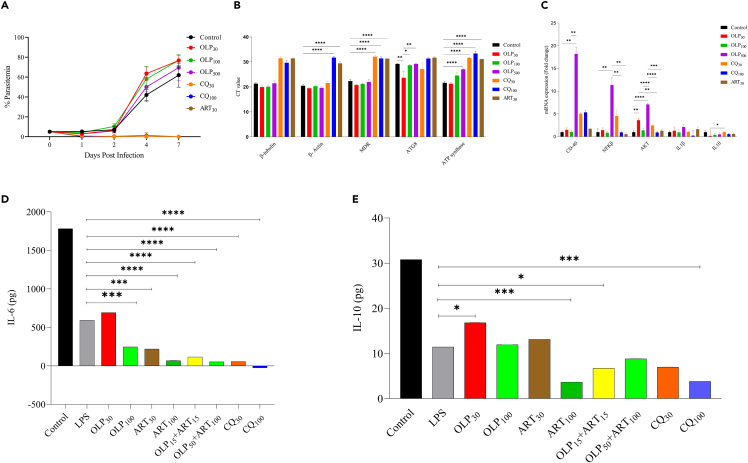
Figure 10.
Infectious challenge model of P. berghei infection in the BALB/c mice to confirm activation of autophagy under OLP pressure
(A) The parasite load (% parasitemia) was determined by the Giemsa-stained thin blood smears drawn from the experimental mice receiving treatment with OLP (30, 100, 500 mg/kg), CQ (30, 100 mg/kg), and ART (30 mg/kg).
(B) Determination of the expression (cT value) of autophagy and development markers of pathogen (P. berghei) (mdr, atg8, Atp synthase) by RT-PCR on the mRNA isolated from blood samples collected on the experimental mice on day 10 post-infection (day 5 post drug treatment).
(C) The relative mRNA expression (fold change regulation) of signaling (CD40, NF-κBp50, Akt1) and inflammatory (IL-1β, IL-10) markers of the host (Balb/C mice). The reduction in the chronic inflammation induced by the LPS (1 μg/mL)-stimulated human macrophages following the OLP treatment was measured by ELISA.
(D and E) The expression of (D) pro-inflammatory (IL-6) and (E) anti-inflammatory (IL-10) cytokines was quantified from the supernatant collected on the OLP-, ART-, and CQ-treated macrophages. Combination of OLP & ART was used at the 30 and 100 μg/ml. The data represent one independent experiment in triplicate (n = 1). (∗p < 0.05 [significant], ∗∗p < 0.01 [moderately significant], ∗∗∗p < 0.001 [highly significant], ∗∗∗∗p < 0.0001 [extremely significant]).
P. berghei activates autophagy under OLP pressure
Parasitemia patterns seen in the P. berghei infected mice treated with the drugs prompted us determine the expression of mdr1, atg8, and Atp synthase from the blood samples collected on the experimental mice (Figure 10B) (Primer information; Table S3). We saw lower cT values (higher expression) of mdr1 at all concentrations of OLP (30, 100, and 500 mg/kg) tested when compared to the untreated control (Figure 10B). The higher expression of mdr1 suggested that the parasite may have well-tolerated the OLP pressure. Moreover, the higher expression of food-vacuole Pbmdr1 indicates the role of autophagy-like escape phenomenon during the P. berghei infection.42 On the contrary, lower expression (higher cT values) of mdr1 observed in the CQ (30 and 100 mg/kg)- and ART (30 mg/kg)-treated animals exhibited the parasite clearance from the mouse circulation (Figure 10B). Further, Atg8 expression was seen to be significantly (p < 0.001) higher with 30 mg/kg OLP concentration. However, CQ and ART treatment did not reach the statistical significance for the expression of Atg8 (Figure 10B).
Next, we determined the gene transcription of the downstream mediators of the PI3K-Akt1 signaling pathway in mice (Figure 10C). We estimated many-fold non-significant relative mRNA expression of co-stimulatory CD40 whereas the nuclear transcription factor (NF-κB) expression was significantly (p < 0.01) higher in the animals treated with OLP (500 mg/kg). Akt1 expression was seen significantly (p < 0.001) higher at both 30 and 500 mg/kg of OLP indicating the clearance of P. berghei by the host immune responses. In brief, gene transcription profiling and parasitemia patterns indicated the dose-dependent activation of autophagy. Additionally, level of parasitemia estimated on Giemsa-stained thin blood smears showing a steep increase at 30 mg/kg of OLP compared to the untreated control confirmed the dose-dependent induction of autophagy, whereas higher OLP concentration allowed the host to clear parasite from circulation (Figure 10C). Hardly any difference was seen in the expression of IL-10 and IL-1β.
OLP modulates the host immune responses
LPS-stimulated macrophages were treated with OLP (30, 100 μg/mL), ART (30, 100 μg/mL), CQ (30, 100 μg/mL), and combination of OLP and ART (30 μg/mL and 100 μg/mL). ELISA was carried out from the cell supernatant to assess the regulation/modulation of the antigen-elicited inflammation (Figures 10D and 10E). The significant (p < 0.0001) reduction in the acute inflammation was characterized by the reduction of the IL-6 when treated with individual drugs (OLP, ART, CQ) and combination of OLP and ART (Figure 10D).48 This reduction in the inflammation confirmed the anti-inflammatory potential of OLP as shown by the quantification of anti-inflammatory/immunoregulatory markers (IL-10)49 (Figure 10E). Significantly (p < 0.001) higher secretion of IL-10 was measured when macrophages were treated with OLP at 30 μg/mL. Collectively, these data suggested that OLP treatment modulates the host-elicited inflammation in response to the antigenic stimulation and induced the autophagy.
Histopathology of deep-seated organs
The experimental mice receiving drug treatment were euthanized, and brain, liver, and spleen were harvested to detect the pathological changes (Figure S8). Also, we kept uninfected and untreated mice as the control (Figures S8 and S9). The sections of the extracted organs were stained with the hematoxylin and eosine (H&E). The brain histopathology of untreated control (infected mice receiving vehicle PBS) group showed vacuolated neutrophils with the increased surrounding space in the brain cortex. 30 mg/kg CQ (CQ30) showed the vacuolization in the neutrophils and congestion in the blood vessel, whereas CQ100 and ART30 showed severe congestion in the leptomeninges and increased hyper-cellularity of neurons. Furthermore, experimental animal receiving OLP30 & OLP100 showed vacuolization in the neutrophils and congestion in the blood vessel in the brain cortex. H&E staining of the brain sections exhibited mild vacuolization in the neutrophils in the brain cortex and no congestion in the blood vessels (Figure S8; upper panel).
Liver histopathology from the untreated control showed severe hemosiderosis, central vein congestion, multifocal necrosis, and Kupffer cell hyperplasia. The experimental animals treated with the CQ30, CQ100, and ART30 showed mild hemosiderosis, vacuolar degeneration, multifocal necrosis, and distorted hepatic cords. OLP30, OLP100, and OLP500 exhibited moderate hemosiderosis, central vein congestion, and multifocal necrosis along with severe Kupffer cell hyperplasia.
In the end, spleen histopathology from the untreated control showed severe depletion of white pulp and presence of hemosiderin pigment. CQ at both the concentrations tested showed moderate depletion of white pulp with the reticuloendothelial (RE) cell hyperplasia. And, animals receiving treatment with ART30 got nearly normal architecture of the spleen with mild depletion of white pulp. On the contrary, OLP30 and OLP100 exhibited mild hemosiderosis and depletion of white pulp of the spleen. The histology data suggested the presence of hemosiderin pigment in the red and white pulp at all the tested OLP concentrations (Figures S8 and S9).
Discussion
As shown in other apicomplexan,4 there are a few reports confirming the activation of autophagy during physiological stress. We first developed a MMI to determine the anti-inflammatory/signaling activity in both the prophylactic and therapeutic mode of treatment of OLP. Our data indicate the inhibition of active NF-κB mediated by the co-stimulatory CD40 and Akt1 in the therapeutic mode of OLP treatment. OLP treatment is shown to modulate the inflammation and signaling pathway such as the PI3K-Akt1 pathway in the stimulated macrophages.5,50 Further, free radical scavenging activity of OLP is shown to exhibit anti-inflammatory and disease-fighting activity.51 The relative mRNA expression of inflammatory immune mediators in the antigen-stimulated macrophages following the drug treatment in prophylactic and therapeutic mode suggested the attenuation of inflammatory responses. Our data confirming the reduction in the inflammation is consistent to the reports published in human coronary artery endothelial cells (HCAEC)52 and rodent plasmodia infection.13 Further, higher expression of anti-inflammatory/immunoregulatory cytokine (IL-10) seen in the in LPS-stimulated macrophages following the drug treatment53 confirmed the anti-inflammatory activity of OLP.
Reduced mRNA and protein level expression of immune-markers related to PI3k-Akt1 signaling and inflammation suggested the altered phenotypic expression of macrophages.54 CD40-dependent and independent cell death is regulated by the transcription nuclear factor (NF-κB) by controlling the expression of pro-apoptotic protein (Bim). The latter regulates the inhibited expression of FOXO1,31,32 an important nuclear marker for apoptosis. CD40-dependent activation of PI3K-Akt1 signaling pathway mediates the survival of antigen-stimulated macrophage by the OLP treatment.55 Furthermore, OLP-induced autophagy drives the cell differentiation and inhibits the default apoptosis.56,57 The protein expression of Beclin-1 and cytoplasmic expression of Akt1, NF-κB, and LC3 II (Figures 1, 2, and 3) suggested that induced autophagy confers tolerance toward the OLP treatment. No pro-apoptotic protein (Bim) expression in OLP- or ART-treated macrophages suggested the activation of autophagy that circumvents OLP pressure and extends survival.
The obligate intracellular parasite, P. falciparum, resides inside human red blood cells (RBCs) (huRBCs) and uses hemoglobin as its primary nutritional source. huRBCs that host P. falciparum are equipped with various intracellular defense mechanism(s) to destroy the invading pathogens.58 Hence, autophagy could be a remarkable tool that pathogens may have to confront upon the host cell invasion. The assays performed with the MMI (Figures 1, 2, 3, and 4) and molecular docking (Figure 5) analyses suggested that the modulated inflammation activates autophagy under OLP pressure. Hence, we then decided to explore the autophagy employed by P. falciparum under OLP pressure. As earlier,18,59 we confirmed the autophagy-mediated tolerance when P. falciparum was submitted to the OLP pressure.
Modulated inflammation and induction of autophagy under OLP pressure suggested maintained eubiosis during P. falciparum infection. Autophagy is reportedly known to repair the disrupted host-parasite interaction that helps survive drug pressure for normal growth and development of the parasite.60 As others,17,59,61 our data suggest the starvation-induced autophagy in wild type (3D7), CQ-resistant (Dd2), and D10 parasite expressing D10-Atg18 line of P. falciparum. The growth of Atg18 parasite remained unaffected by the autophagy activated in response to starvation.61 Absence of full repertoire of proteins responsible for autophagy poses a challenge to dissect the mechanism responsible for autophagy. Bioassays performed with MMI led us to determine the antimalarial activity of OLP in 3D7, D10-Atg18, and DD2 P. falciparum. We observed the parasite clearance activity of OLP when parasite was exposed to the drug for longer durations. Also, OLP leaves an additive antimalarial effect when used in combination with ART. The plasma half-life and bio-distribution study in humans62 suggest the use of OLP as an antimalarial drug.13 We are the first group to report activity of OLP in P. falciparum. Since P. falciparum has been gaining resistance to almost all frontline antimalarials15,63,64,65 as well as artemisinin-based combination treatment (ACT) drug treatment,15,66 OLP could be a possible partner drug for the ACT. Autophagy activated under OLP pressure enables parasite to tolerate and survive the drug pressure. The expression of Atg8 in the starved and OLP-treated 3D7 is in agreement with earlier findings wherein autophagy was activated against drug/physiological pressure by the parasite2,18,67 and other diseases.5,68 We confirmed the in vitro findings in the experimental challenge model of P. berghei infection.69,70 Mice harboring P. berghei infection receiving treatment with OLP showed a decreased parasite load (Figure 10A) for a few days followed by a sharp increase in the parasite infection at all the concentrations of OLP (30,100, 500 mg/kg) tested. The infectious parasite load was seen to be significantly higher at lower concentration of OLP (OLP30) than that seen with the higher OLP concentrations (100, 500 mg/kg) as compared to untreated control. This confirms that parasite activated the dose-dependent autophagy to circumvent the anti-plasmodial defenses. The gene transcription profiling of P. berghei (pathogen) and mice (host) further suggested that the activation of autophagy-like escape mechanism evading the anti-plasmodial defenses could be a double-edged sword6 (Figures 10B and 10C). Autophagy helped parasite escape the anti-plasmodial defenses and withstand drug pressure to confer the extended survival by inhibiting the programmed cell death (Figures 4 and 10C).71 In the end, histopathological observation from deep-seated tissues (spleen, brain, and liver) indicates the lesser cell infiltration suggesting the induction of autophagy for extended survival of malaria parasite. Also, the lower dose of OLP modulates the host immune response by influencing the cell death/autophagy axis (Figures 10D and 10E). ELISA results confirmed the dose-dependent immune response-modulatory effect of OLP to activate autophagy that confers the extended survival on the antigen-stimulated macrophages. Collectively, assays with the MMI, molecular docking, P. falciparum assays, and challenge model of P. berghei study confirmed the activation of autophagy for extended survival of the parasite.
Conclusions and future perspectives
The present work confirmed the induction of autophagy in the MMI, in vitro culture of P. falciparum infection (3D7, D10-Atg18, Dd2), and challenge model of P. berghei infection (Figure 11). This work is of clinical relevance as far as human health is concerned since it could be one of the plausible explanations for the evolution of antimalarial drug resistance. The present work shall help find newer drugs, and OLP could be used in combination with ART to increase the latter’s parasite clearance activity.
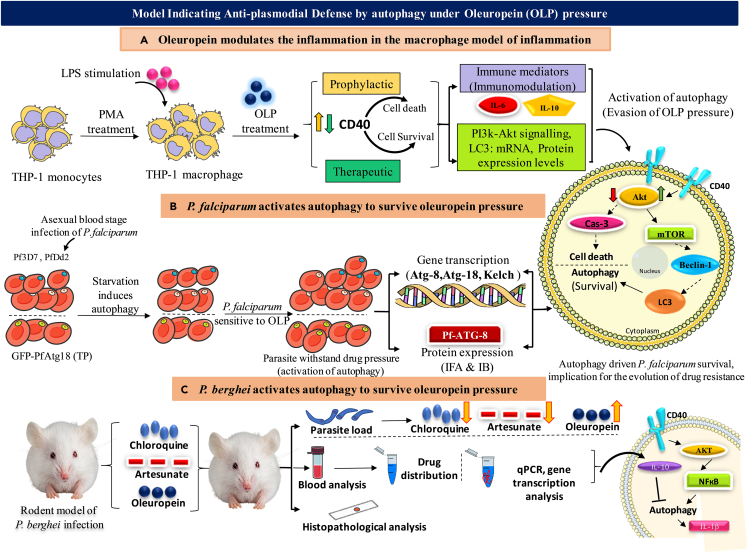
Figure 11.
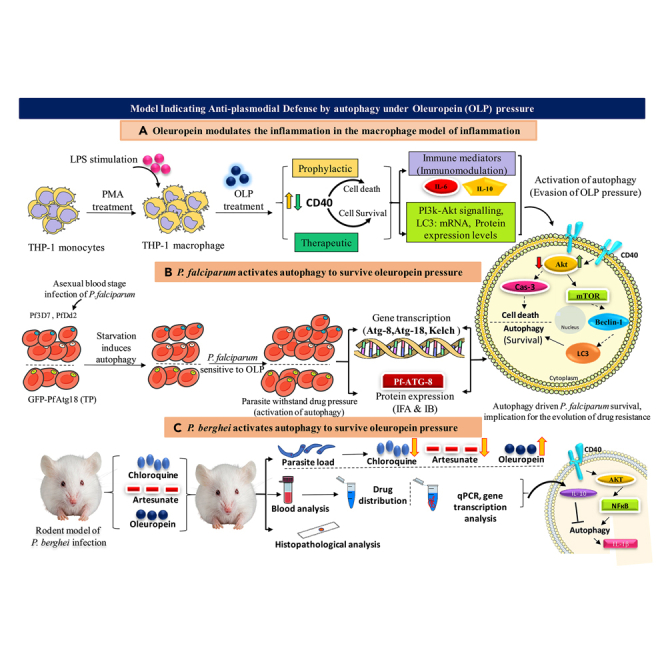
Model indicating the activation of immune response evasion mechanism (autophagy) by the oleuropein (OLP) treatment in the antigen (LPS)-stimulated human macrophages
Antimalarial activity of OLP in the routine culture of P. falciparum (3D7, D10-Atg18, Dd2) and confirmation of the elicitation of the anti-plasmodial defenses driven by autophagy.
(A) Development of LPS-stimulated human THP-1 macrophages (macrophage model of inflammation; MMI) to determine modulation of inflammation controlled by the PI3K-Akt1 signaling to establish the immune homeostasis. OLP treatment circumvents the death of the antigen stimulated macrophages.
(B) Antimalarial activity of OLP in 3D7, D10 parasite expressing GFP-Atg18 line, chloroquine-resistant (Dd2) P. falciparum. Gene transcription and protein level expression, autophagy, and development-related proteins (ARP/DRP) to confirm the inhibited death of the antigen-stimulated macrophages regulated by the Akt1-mediated signaling.
(C) Developed challenge model of P. berghei infection to validate the in vitro findings showing the induction of autophagy under OLP pressure.
Limitation of the study
Present study showing the activation of autophagy in the MMI, laboratory strains of the asexual BS infection of P. falciparum, and challenge model of P. berghei infection is limited due to the use of human RBCs reconstituted NSG (immunodeficient) mice (humanized mice). This finding could be further validated in the humanized mice of P. falciparum infection.
STAR★Method
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| AKT1 | Cell Signaling Technologies (USA) | Cat No: C73H10 |
| pAKT | Cell Signaling Technologies (USA) | Cat No:S473 |
| NF-κB | Cell Signaling Technologies (USA) | Cat No: S536 |
| NFκBp | Cell Signaling Technologies (USA) | Cat No: D14E12 |
| pfAtg8 | Raised in rabbit by Dr. Sijwali, CCMB | – |
| FITC conjugated anti-rabbit IgG | Santa Cruz, USA | Cat No: sc2357 |
| Beclin-1 | Cell Signaling Technologies (USA) | Cat No: D40C5 |
| CD40 | Cell Signaling Technologies (USA) | Cat No: D8W3N |
| Cas-3 | Cell Signaling Technologies (USA) | Cat No: 9662S |
| Cas-8 | Cell Signaling Technologies (USA) | Cat no: D35G2 |
| Bim | Cell Signaling Technologies (USA) | Cat No: C34C5 |
| IDO1 | Cell Signaling Technologies (USA) | Cat No: D5J4E |
| LC-3 | Cell Signaling Technologies (USA) | Cat No: 4108S |
| Alexa Fluor 488-conjugated anti-mouse antibodies | Cell Signaling Technologies (USA) | Cat No: 4408 |
| Bacterial and virus strains | ||
| Plasmodium falciparum 3D7, Dd2 | Received from NIMR, New Delhi | |
| D10 parasite expressing GFP-Atg18 line of P. falciparum | Dr. Puran S. Sijwali, CSIR-CCMB, Hyderabad, India | |
| P. berghei; ANKA strain | Dr. Agam Singh, NII, New Delhi | |
| Biological samples | ||
| Crude extract of oleuropein (OLP) | Prof. B. S. Chandel, Sardarkrushinagar Dantiwada Agricultural University, Gujarat, India | |
| Critical commercial assays | ||
| FITC ANNEXIN V Apoptosis detection kit | Invitrogen™ | Cat No: V13242 |
| CM-H2DCFDA dye (General Oxidative Stress Indicator) | ThermoFisher scientific, USA | Cat No: C6827 |
| ECL reagent (Pierce western blotting reagent) | ThermoFisher scientific, USA | Cat No: 32134 |
| IL-6 Human Uncoated ELISA Kit | ThermoFisher scientific, USA | Cat No: 88-7066-88 |
| IL-10 Human Uncoated ELISA Kit | ThermoFisher scientific, USA | Cat No: 88-7106-88 |
| GeneJET RNA purification kit | ThermoFisher scientific, USA | Cat No: K0731 |
| DNAeasy Blood and tissue kit | Qiagen | Cat No.69504 |
| iScript™ cDNA Synthesis Kit | Bio-Rad, USA | Cat No: 1708890 |
| RevertAid First Strand cDNA Synthesis Kit | ThermoFisher scientific, USA | Cat No.: K1622 |
| Experimental models: Cell lines | ||
| Human monocytic cell (hTHP-1) | ATCC, USA | |
| Software and algorithms | ||
| Oleuropein | PubChem Database | CID_5281544 |
| Artesunate | PubChem Database | CID_6917864 |
| AutoDock Vina 1.1 | Center for computational structure biology, The Scripps Research Institute, USA | |
| Pymol Software | PyMOL is an open source but proprietary molecular visualization system created by Warren Lyford DeLano. | Pymol.org |
| 3-dimensional (3D) structures of ligand (ART and OLP) | Available on Pubchem | Oleuropein (CID_5281544) (artesunate; CID_6917864) |
| Other | ||
| RPMI-1640 media | Gibco Life Technologies, USA | Cat. No. 31800-022 |
| Fetal bovine serum (FBS) | Gibco Life Technologies, USA | Cat. No. 10270106 |
| antibiotic cocktail (Penicillin + streptomycin) | Gibco Life Technologies, USA | Cat No.15070063 |
| β-mercaptoethanol | GCC biotech | Cat No.GPC-009 |
| phorbol 12-Myristate 13-Acetate (PMA) | Sigma | Cat No: 524400 |
| LPS | Sigma | Cat No: L7895 |
| Chloroquine (CQ) | Sigma-Aldrich | Cat No.: C6628 |
| Artemisinin (ART) | Sigma-Aldrich | Cat No.: A3731 |
| 3-methyl adenine (3-MA) | Sigma-Aldrich | Cat No: M9281 |
| Trizol | Invitrogen, US | Cat No 15596026 |
| SYBR Green | Invitrogen, US | Cat No: S7563 |
| HEPES | Himedia Laboratories Pvt. Ltd., India | Cat. No. MB016 |
| NaHCO3 | Himedia Laboratories Pvt. Ltd., India | Cat. No. TC230 |
| Hypoxanthine | Sigma-Aldrich, USA | Cat. No. H9636-5G |
| Albumax | Gibco Life Technologies, USA | Cat. No. 11020-021 |
| Gentamicin | Gibco Life Technologies, USA | Cat. No:15750060 |
| anti-fade | ThermoFisher scientific, USA | Cat. No: P36930 |
| Blasticidin | Gibco, USA | Cat. No: R210-01 |
| GENES2ME Green Eye-Ab universal qPCR Master Mix | Imperial Life Sciences, India | Cat No. SMM03, Genes2me |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Dr. Rajeev K Tyagi (rajeevtyagi@imtech.res.in, rajeev.pasteur@gmail.com).
Material availability
This study did not generate new unique reagents.
Data and code availability
-
•
This study does not report any original code.
-
•
All unique/stable reagents generated in this study are available from the lead contact without restriction.
-
•
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Experimental model and study participant details
All animal experiments were reviewed and approved by the Institutional Animal Ethical Committee (IAEC) (protocol No.: IS/PHD/30/2022/35 & IS/PHD/30/2022/46), Nirma University, Ahmedabad, India. The 6–8 weeks old BALB/c were procured from Zydus Research Center (ZRC), Ahmedabad and housed in pathogen free conditions at the central animal facility of Nirma University. Animals were acclimated for a week before initiating the studies. Animals were provided food and water ad libitum.
Animal model
BALB/c mice.
Sex and age
Male and 8–10-week-old.
Method details
Development of antigen stimulated macrophage model of inflammation (MMI)
Human monocytic cell (hTHP-1) line was procured from ATCC, USA, and cultured in RPMI-1640 medium supplemented/containing 10% heat de-complemented fetal bovine serum (FBS), penicillin and streptomycin (100 U/mL), β-mercaptoethanol (3.5 μL/L) and maintained in a humidified atmosphere of 95% O2, 5% CO2 at 37°C. hTHP-1 monocytes (2 × 105 cells) were differentiated with100 ng/mL phorbol 12-Myristate 13-Acetate (PMA, Sigma, Cat: 524400) for 24 h. Following the cell differentiation, cells were stimulated with the antigen (LPS; 1 μg/mL) for another 24 h followed by the treatment with OLP, ART (30 μg/mL) individually as well as combination for 24 h.72 The antigen (LPS) stimulated cells were then used to study the gene expression profiling through the RT-PCR.
Gene expression by RT-PCR in the MMI and P. falciparum
The drug treated parasites were harvested and the pellet was washed twice with 1X PBS after centrifugation at the 2,000 rpm for 5 min followed by the DNA isolation using the manufacturer’s protocols (DNAeasy Blood and tissue kit, Qiagen (Cat No 69504). The DNA quantity and quality was determined by the Nano-drop, and 50 ng of total DNA was used to perform RT-PCR on CFX96 Connect, Biorad, USA.
Similarly, total mRNA was isolated from hTHP-1 macrophages using Trizol reagent, ThermoFischer scientific, USA (Cat No 15596026) and approximately 400 ng of total mRNA was converted to cDNA using iScript (Cat No: 1708890) Biorad, USA. The expression analysis was performed using SYBR green master mix. Further, the reaction mixture containing 10 μM forward and reverse primers, 2X SYBR green was carried out at 95°C for 3 min, annealing at 55°C for 30 s, and extension at 72°C for 30 s spawning 40 cycles. The human specific gene (CD40, Akt, NF-kβ, Bim, IL-1β, IL-6, TNF-α, iNOS, IL-10) expression was normalized to GAPDH for hTHP-1 and β-Tubulin for P. falciparum73 (SI information, Table S1). The expression for P. falciparum specific genes (Kelch-13, Atp synthase, mdr1, nhe1, Atg8, and Atg18) were quantified with the individual treatment of OLP and ART as well as combination of OLP and ART (SI information, Table S2). The combination of drugs was used as follows. For example; 40 nM concentration of OLP or ART whereas combination was used as 20 nM each of OLP and ART. This methodology was used for all the concentrations tested.
Confocal microscopy based immunofluorescence assay (IFA)
hTHP-1 cells were seeded on to the glass coverslips, and differentiated into the macrophages. Further, the cells were treated with 30 μg/mL of OLP and ART each incubated for 24 h. The media was aspirated and cells were washed with 1X PBS, fixed with 4% para-formaldehyde (PFA) for 5 min followed by the permeabilization with 0.1% Triton X-100 for 3 min. The fixed and permeabilized cells were blocked with 5% BSA reconstituted in PBS for 1 h. The blocked cells were incubated with the primary antibodies (AKT, pAKT, NF-κB, NFκBp & pfAtg8 at a 1:200 dilution) for 2 h. The washed cells were then incubated with the FITC conjugated anti-rabbit IgG for 1 h. The antibody stained cells were placed on slides, washed and mounted with the anti-fade (Thermo Fisher Scientific, USA) mounting medium with DAPI.74 The images were captured using Nikon confocal microscope and data was processed and quantified by the software, ImageJ-win64.
Annexin V/PI staining to detect the death of antigen stimulated macrophages
PMA differentiated and LPS stimulated hTHP macrophages were treated with the drugs for 24 h as mentioned above. The drug treated cells were harvested by the trypsin/EDTA. The pelleted cells were washed with 1X PBS and processed according to the manufacturer’s recommendations (FITC ANNEXIN V Apoptosis detection kit Invitrogen, Cat No: V13242). Briefly, cells were re-suspended in 1X Annexin binding buffer and then stained by incubating with 5 μL of Annexin V dissolved in 1X Annexin binding buffer and 1 μL propidium iodide (PI) for 30 min in the dark at 37°C. Following the incubation, 10,000 events were acquired using BD verse and data was analyzed on FlowJo (FlowJo_v10.8.1).
Assessment of ROS generation
hTHP-1 macrophages were treated and harvested as described above. The harvested cells were washed with 1X PBS buffer and then stained with 1μM CM-H2DCFDA dye (Cat No: C6827, Thermofisher Scientific) at 37°C for 15 min and the fluorescence was recorded by using Flow-cytometer. The 10,000 events were captured and data was analyzed using FlowJo (FlowJo_v10.8.1) software.75
Immunoblot analysis
The antigen stimulated macrophages were harvested once the drugs (OLP, ART and combination of OLP & ART) treatment (24 h) was over. The cells were lysed using radio-immunoprecipitation assay (RIPA) buffer mixed with protease inhibitors for 30 min and vortexed intermittently. The supernatant was clarified by centrifugation at 12,000 rpm for 15 min. Further, protein concentration was quantified by the Bradford reagent and 20 μg protein was loaded on to each well, and subjected to non-reducing SDS-PAGE. The separated proteins were transferred on to the nitrocellulose (NC) membrane for 2 h at 100V. NC membranes transferred with the protein bands were blocked with 5% BSA in PBS for 1 h. Further, proteins transferred NC membranes was incubated overnight with the primary antibodies of NF-kβ, Beclin-3, CD40, Akt1, Cas-3,8, Bim, IDO1, and LC-3 (CST, USA). NC membranes were washed three times with 1X PBST and incubated with the secondary antibody for 1 h followed by three washes with 1X PBST. In the end, blots were developed using ECL reagent (Pierce western blotting reagent, Cat no: 32134, USA) on a Chemi-doc system.
Macrophage supernatant quantification of IL-6 & IL-10
IL-6 (Cat no: 88-7066-88) and IL-10 (Cat no: 88-7106-88) were procured from the Invitrogen, USA. The drug (OLP, ART and OLP & ART) treated supernatant of LPS stimulated macrophages was assessed with respect to the secretion of pro-and anti-inflammatory cytokines. The sensitivity of the ELISA kit was reported 2 pg/mL for both IL-6, and IL-10. Briefly, plates were coated with the coating antibody overnight at 4°C. The solutions for standard and working samples were added to the coated wells, and plates were incubated overnight at 4°C. The plates were washed three times with the washing buffer, and incubated with the biotin-conjugated secondary antibodies for 2 h at room temperature (RT). Following the incubation, washed plates were treated with the streptavidin-HRP enzyme and incubated for 1 h at RT. Following the incubation. TMB substrate was added and incubated for 10 min in the dark for the development of color at RT. In the end, reaction was stopped by adding stop solution (2N H2SO4), and plates were read at the 450 nm at an ELISA plate reader (BioTek, EPOCH/2 microplate reader). The concentration of the immune markers was determined comparing the cytokines with the standards.
Molecular docking to analyze protein-drug interactions
The structure of proteins involved in the autophagy mediated by PI3K-Akt1 signaling pathway was retrieved from Protein DataBank (PDB). Oleuropein (CID_5281544) and a well-known antimalarial drug (artesunate; CID_6917864) were retrieved from PubChem database to dock the proteins with OLP and ART. The proteins (AKT1, KELCH, ATG18, ATP-Synthetase, MDR1, LC3-II, HSP70, IL-10, IL-1B, NHE, TNF, BIM, NF-KB, IL-6, ATG8 and, CD40) responsible for the anti-inflammatory, antimalarial and signaling activity were docked with the artesunate and Oleuropein (Table 2). Based on the binding affinity, AKT1 (PDB: 3096),76 KELCH (PDB: 4ZGC), LC3-II (PDB: 2ZJD_A) and, ATG18 (PDB: 6KYB) were chosen for the detailed molecular docking analyses. The corresponding PDB structure was further energy minimized by Chiron web server,77 and 3-dimensional (3D) structures of ligand (ART and OLP) were obtained from PubChem (https://pubchem.ncbi.nlm.nih.gov/). The molecular docking study was performed by using AutoDock Vina 1.1.78 A grid box with a spacing of 1 Å and a size of 40 × 40 × 40 were built around the center of the binding site as predicted by CB-dock server for AutoDock Vina 1.1.79 Further, other parameters of docking were set to default whereas exhaustiveness value was adjusted to 8 and energy range to 4. The pymol software was used to further analyze the structure.
In vitro culture of asexual blood stage (BS) of P. falciparum
Parasite culture
Plasmodium falciparum 3D7, Dd2, and D10 strains were grown in RPMI 1640 medium (containing 2 g/L glucose, 2 g/L sodium bicarbonate, 300 mg/L glutamine, 25 μg/mL gentamicin and 100 μM hypoxanthine) with 0.5% albumax II and human RBCs at 2 and 5% hematocrit under a gas mixture (5% CO2 + 5% O2 + 90% N2) at 37°C.80,81 P. falciparum D10 strain episomally expressing Atg18 (D10-Atg18) has been described earlier,42 and maintained in the above medium with blasticidin (1 μg/mL), which was excluded during experimental conditions. Synchronization of was carried out by treating parasites with 5% sorbitol treatment when the majority of parasites were in ring stage.82 Parasite growth and development was determined by the microscopic examination of Giemsa-stained thin blood smears drawn on the parasitized RBCs. The D10 parasite expressing GFP-Atg18 line of P. falciparum was obtained from Dr. Puran S. Sijwali, CSIR-CCMB, Hyderabad, India.
In vitro antimalarial drug screening
In vitro antimalarial activity assays with different P. falciparum strains were carried out essentially as has been described previously.80,83 Briefly, parasites were synchronized at ring stage, the compounds were dissolved in DMSO (OLP, artemisinin) or water (chloroquine). The compound stocks were serially diluted 2-fold in 50 μL of culture medium across a 96 well tissue culture plate. DMSO (0.5% final) and chloroquine (500 nM) were added to some of the wells as positive and negative controls, respectively. 50 μL of the ring stage parasite culture (1–2% parasitemia at 4% hematocrit, ≈0.5% parasitemia for OLP) was added to each well, and the plate was incubated at 37°C for 50 h (for pyrimethamine, chloroquine, artemisinin) or 96 h (for OLP). For OLP assay plate, culture media was replaced with fresh culture medium containing the corresponding concentration of OLP or DMSO or chloroquine. At the end of incubation, 100 μL of lysis buffer (20 mM Tris-Cl, 5 mM EDTA, 0.008% saponin, 0.08% Triton X-100, pH 7.5, SYBR green 1 at the manufacturer’s recommended dilution) was added to each well and the plate was incubated at 37°C for 30 min. The fluorescence of each well was measured (λex 485 nm, λem 530 nm) using Spectra Max iD3 multimode microplate reader. The background fluorescence of chloroquine control was subtracted from those of DMSO and test compound wells. The adjusted values were normalized as percentage of the fluorescence value of DMSO control, and plotted as % inhibition versus concentrations using GraphPad Prism software.
For antimalarial activity of OLP-artemisinin combination, OLP or artemisinin or different artemisinin-OLP combinations were added in 50 μL of culture medium in a 96 well tissue culture plate. 50 μL of culture medium with ring stage parasite (≈1.5% parasitemia and 4% hematocrit) was added to each well to achieve 500 μM OLP alone or artemisinin alone (at 1×IC50 (21 nM), 0.5×IC50 (10.5 nM), 0.25×IC50 (5.25 nM)) or artemisinin with 500 μM OLP. The plate was incubated at 37°C for 50 h and fluorescence was read as described above. The adjusted values were normalized as percentage of the fluorescence value of DMSO control, and plotted as % inhibition versus drug concentrations using GraphPad Prism software.
DNA isolation and parasite starvation
P. falciparum at 0.2% ring-stage parasites with 5% hematocrit was treated with the different concentrations of OLP and ART (40, 80, 160, 320 nM) for 72 h. Drug treated parasite were harvested for DNA isolation following the 72 h of incubation. Parasite was cultivated in the Albumax deficient medium followed by the 3-MA treatment18 to confirm the starvation induced autophagy. The schizont stage parasite was treated with 3 mM 3-MA for 3 h followed by the starvation for another 4 h to confirm the induction of autophagy under the physiological stress. The parasite load (relative parasitemia) was determined using SYBR Green-I as reported elsewhere.84 SYBR Green-I was diluted in sterilized distilled water (10X) and 25 μL SYBR Green I (10X) was added to 100 μL sample. Following experimental groups were included in the assay; P. falciparum specific RPMI-1640 medium (background), uninfected RBCs (cell background), infected RBCs (control) and treated parasitized RBCs as described above. The thin blood smears were drawn following the incubation of the parasite treated with drugs and observed under the microscope. 25 μL of SYBR Green-I (10X) was added to each well and incubated further for 30 min at 37°C. The final read-outs was recorded at Ex485/Em530 nm using microplate reader (Biotek Synergy).84 The parasites were treated alone and in combination with different concentrations of ART and OLP (40, 80, 160, and 320 nM) for 72 h. The drug treated parasites were harvested and processed for further experimentation.
Immunofluorescence assay with P. falciparum
Synchronized P. falciparum 3D7 and D10-Atg18 strains at 4–5% parasitemia were cultured till the late-trophozoite stage (36 h), and the cultures were divided into four equal parts. One part was cultured in complete culture medium, the 2nd part was grown in albumax-deficient culture medium (starvation), the 3rd part cultured in complete culture medium containing 320 nM OLP, and the 4th part was cultured in complete culture medium with 3 mM 3-MA. All the cultures were grown for 3 additional hours and then processed for microscopy. Briefly, a 100 μL aliquot of each culture from was washed with PBS, diluted in PBS to 10% hematocrit, immobilized on poly-L-lysine coated slides, washed with PBS again to remove non-attached cells, the cells were fixed (3% paraformaldehyde and 0.01% glutaraldehyde in PBS), and then quenched with 0.1 M glycine (v/v in PBS). The cells were permeabilized using 0.1% Triton X-100 (v/v in PBS), blocked with 3% bovine serum albumin (in PBS) for 1 h at RT. The slides were incubated with mouse anti-Atg8 antibodies for 1.5 h at RT, washed with PBS, and incubated with donkey Alexa Fluor 488-conjugated anti-mouse antibodies with DAPI (10 μg/mL) for 1 h at RT. The slides were washed with PBS, air dried, mounted with prolong diamond antifade, covered with glass coverslips, and edges were sealed. The slides were observed under 100× objective of ZEISS Axioimager, images were captured with Zeiss AxioCam and analyzed using Axiovision software.
Animal assays
P. berghei maintenance
The parasitized (P. berghei; ANKA strain) whole blood was maintained at the Institute of Science, Nirma University by cyclical transmission in 8–10-week-old male Balb/C mice and Anopheles stephensi mosquitoes. The infection was confirmed by the microscopic examination of thin blood smears drawn (minimum of 1000 RBCs) from tail vein and Giemsa staining each time before the infected blood samples were cryopreserved.
Preparation of drug formulations
OLP was dissolved in 10% DMSO, and the stock solution of 50 mg/mL was prepared using endotoxin free PBS (Himedia: ML164-100ML). The three different dosages (30, 100, & 500 mg/kg were prepared and administered in the experimental animals via intraperitoneal route (IP) of administration. CQ was dissolved in PBS at the final concentration of 50 mg/mL, and two dosages (30 & 100 mg/kg) were administered intraperitoneally. Similarly, ART was dissolved in the ethanol and stock solution of 25 mg/mL was prepared. 30 mg/kg was injected through the IP route. The working solution of all dosages were prepared using endotoxin free PBS, and 100 μL injectable volume was administered to the experimental mice.
Challenge model of P. berghei infection
Mice were first infected with wild-type P. berghei ANKA strain. Following 1–2 passages to achieve higher parasitemia, 106 infected/parasitized RBCs were injected into the intraperitoneal cavity of mice (n = 6 mice/group), and parasitemia was monitored daily by counting minimum 1,000 RBCs of Giemsa-stained thin blood smears. Once the parasitemia reached nearly 5%, animals were treated with the test drugs as well as experimental (CQ) and positive control (ART) (Scheme 1).
Determination of parasite load in vitro
Mice were treated with different dosages of OLP, CQ & ART at 5% parasitemia (Scheme 1). The drugs were administered consecutively for 7 days and trend in the parasite growth and development was determined in terms of % parasitemia calculated through reading the thin blood smears (under the light microscope; Olympus BX53, USA) drawn from the experimental and control mice. The % parasitemia was determined by counting a minimum of 1,000 RBCs.
Gene transcription profiling by RT-PCR
The blood samples were collected on day 5 following the treatment (3 mice/group) to determine fold change regulation in the transcriptional expression of drugs (OLP, ART& CQ) treated mice infected with P. berghei. The RNA was isolated from the blood samples using GeneJET RNA purification kit (Cat K0731, ThermoFisher Scientific). Purity and quantification of extracted RNA was done by the NanoDrop Lite Spectrophotometer (ThermoFisher Scientific: ND-LITE). Further, cDNA was synthesized using RevertAid First Strand cDNA Synthesis Kit (Cat K1622, ThermoFisher Scientific), and relative gene expression profiling was carried out by the RT-PCR/qPCR.
Relative mRNA expression was measured by the real-time PCR system Analyticjena qTOWER3 G and using GENES2ME Green Eye-Ab universal qPCR Master Mix (Cat no SMM03, Genes2me, Imperial Life Sciences, India). Briefly, 5 μL of 2X SYBER green, 0.5 μL forward and reverse primers (SI information; Table S3), 2 μL of diluted cDNA and nuclease free water were added to make the reaction volume up-to 10μL to set RT-PCR using the following thermal conditions; 95°C for activating the Taq Polymerase and rest 45 cycles were performed at 95°C for 15 s, annealing was carried out at 52°C for 1 min for the gene expression of the parasite. The gene amplification of mice was done for 40 cycles at 95°C for 15 s, the annealing was done at 52°C for 30 s followed by the extension at 72°C for 30 s. The melt curve at 95°C for 15 s and 65°C was also included.85 mRNA expression of mice was quantified by the changes quantified in the threshold method (ΔΔCT) and normalized to the house keeping mRNA (encoding β actin) expression. Further, mRNA expression of the parasite was presented in terms alterations seen in the CT values.
Histopathology
Three mice from each group and uninfected and untreated control mice were euthanized by the cervical dislocation on day 7 following the drug treatment. Then, the deep-seated organs (spleen, liver & brain) were collected & fixed in 4% paraformaldehyde. The paraffin embedded section of these organs were stained with Haematoxylin & Eosin (H & E), cell infiltration and other histological changes were observed under the microscope.
Quantification and statistical analysis
The statistical analysis was performed using Graphpad prism (version 8, www.graphpad.com). The statistical significance was calculated using unpaired Student’s t test. p value of ≤0.05 was considered as statistically significant.
Acknowledgments
Nikunj Tandel would like to thank the Indian Council of Medical Research (ICMR) for providing the fellowship to carry out his research (ICMR award letter No.: 2020-7623/CMB-BMS). R.K.T. would like to express his thanks to the central MIL facility of CSIR-IMTECH, Chandigarh. N.T. would like to thank Nirma University for providing the facility to carry out the animal experiments. R.K.T. expresses his thanks to Digna Patel, Mansi Thakker, Shruti Malani, Tejaswini Vaidya, Nimi Kapopara, Mihir Makwana, Manan Katariya, and Harshil Patel from Institute of Science, Nirma University for their help to carry out animal experiments. Dr. Pradip Sen, Ravi Mishra, and Rashmi Kumar are duly acknowledged for allowing to use their instruments for in vitro experiments. Dr. Pawan Gupta’s help is duly solicited in providing us with the plasticware. Dr. Amit Kumar is greatly solicited for all his scientific inputs.
Author contributions
Conceptualization, R.K.T., Praveen Sharma., N.T.; design and execution of the experiments, R.K.T., Praveen Sharma, N.T., Rajinder Kumar, S.N., Prakriti Sharma, S.D., K.S., N.R.C., S.S., Reetesh Kumar; materials supply, B.S.C., P.S.S.; writing – original draft preparation, R.K.T., P.S., N.T.; writing – review and editing, R.K.T. All authors have read and agreed to the published version of the manuscript.
Declaration of interests
The authors declare no competing interests.
Published: March 11, 2024
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2024.109463.
Supplemental information
References
- 1.Weiland A.S. Recent Advances in Imported Malaria Pathogenesis, Diagnosis, and Management. Curr. Emerg. Hosp. Med. Rep. 2023;11:49–57. doi: 10.1007/s40138-023-00264-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Yu L., Chen Y., Tooze S.A. Autophagy pathway: Cellular and molecular mechanisms. Autophagy. 2018;14:207–215. doi: 10.1080/15548627.2017.1378838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ghartey-Kwansah G., Adu-Nti F., Aboagye B., Ankobil A., Essuman E.E., Opoku Y.K., Abokyi S., Abu E.K., Boampong J.N. Autophagy in the control and pathogenesis of parasitic infections. Cell Biosci. 2020;10:101. doi: 10.1186/s13578-020-00464-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ghosh D., Walton J.L., Roepe P.D., Sinai A.P. Autophagy is a cell death mechanism in Toxoplasma gondii. Cell Microbiol. 2012;14:589–607. doi: 10.1111/j.1462-5822.2011.01745.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cirmi S., Maugeri A., Russo C., Musumeci L., Navarra M., Lombardo G.E. Oleacein Attenuates Lipopolysaccharide-Induced Inflammation in THP-1-Derived Macrophages by the Inhibition of TLR4/MyD88/NF-kappaB Pathway. Int. J. Mol. Sci. 2022;23 doi: 10.3390/ijms23031206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Leleu I., Alloo J., Cazenave P.A., Roland J., Pied S. Autophagy Pathways in the Genesis of Plasmodium-Derived Microvesicles: A Double-Edged Sword? Life. 2022;12 doi: 10.3390/life12030415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lins P.G., Marina Piccoli Pugine S., Scatolini A.M., de Melo M.P. In vitro antioxidant activity of olive leaf extract (Olea europaea L.) and its protective effect on oxidative damage in human erythrocytes. Heliyon. 2018;4 doi: 10.1016/j.heliyon.2018.e00805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Porcu C., Sideri S., Martini M., Cocomazzi A., Galli A., Tarantino G., Balsano C. Oleuropein Induces AMPK-Dependent Autophagy in NAFLD Mice, Regardless of the Gender. Int. J. Mol. Sci. 2018;19 doi: 10.3390/ijms19123948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Asgharzade S., Sheikhshabani S.H., Ghasempour E., Heidari R., Rahmati S., Mohammadi M., Jazaeri A., Amini-Farsani Z. The effect of oleuropein on apoptotic pathway regulators in breast cancer cells. Eur. J. Pharmacol. 2020;886 doi: 10.1016/j.ejphar.2020.173509. [DOI] [PubMed] [Google Scholar]
- 10.Yan C.M., Chai E.Q., Cai H.Y., Miao G.Y., Ma W. Oleuropein induces apoptosis via activation of caspases and suppression of phosphatidylinositol 3-kinase/protein kinase B pathway in HepG2 human hepatoma cell line. Mol. Med. Rep. 2015;11:4617–4624. doi: 10.3892/mmr.2015.3266. [DOI] [PubMed] [Google Scholar]
- 11.Liu L., Ahn K.S., Shanmugam M.K., Wang H., Shen H., Arfuso F., Chinnathambi A., Alharbi S.A., Chang Y., Sethi G., Tang F.R. Oleuropein induces apoptosis via abrogating NF-kappaB activation cascade in estrogen receptor-negative breast cancer cells. J. Cell. Biochem. 2019;120:4504–4513. doi: 10.1002/jcb.27738. [DOI] [PubMed] [Google Scholar]
- 12.Yamamoto Y., Gaynor R.B. Therapeutic potential of inhibition of the NF-kappaB pathway in the treatment of inflammation and cancer. J. Clin. Invest. 2001;107:135–142. doi: 10.1172/JCI11914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Misganaw D., Engidawork E., Nedi T. Evaluation of the anti-malarial activity of crude extract and solvent fractions of the leaves of Olea europaea (Oleaceae) in mice. BMC Compl. Alternative Med. 2019;19:171. doi: 10.1186/s12906-019-2567-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.White N.J. Emergence of Artemisinin-Resistant Plasmodium falciparum in East Africa. N. Engl. J. Med. 2021;385:1231–1232. doi: 10.1056/NEJMe2110659. [DOI] [PubMed] [Google Scholar]
- 15.Tyagi R.K., Gleeson P.J., Arnold L., Tahar R., Prieur E., Decosterd L., Pérignon J.L., Olliaro P., Druilhe P. High-level artemisinin-resistance with quinine co-resistance emerges in P. falciparum malaria under in vivo artesunate pressure. BMC Med. 2018;16:181. doi: 10.1186/s12916-018-1156-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Stokes B.H., Ward K.E., Fidock D.A. Evidence of Artemisinin-Resistant Malaria in Africa. N. Engl. J. Med. 2022;386:1385–1386. doi: 10.1056/NEJMc2117480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Joy S., Thirunavukkarasu L., Agrawal P., Singh A., Sagar B.K.C., Manjithaya R., Surolia N. Basal and starvation-induced autophagy mediates parasite survival during intraerythrocytic stages of Plasmodium falciparum. Cell Death Dis. 2018;4:43. doi: 10.1038/s41420-018-0107-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ray A., Mathur M., Choubey D., Karmodiya K., Surolia N. Autophagy Underlies the Proteostasis Mechanisms of Artemisinin Resistance in P. falciparum Malaria. mBio. 2022;13 doi: 10.1128/mbio.00630-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Fayez N., Khalil W., Abdel-Sattar E., Abdel-Fattah A.F.M. In vitro and in vivo assessment of the anti-inflammatory activity of olive leaf extract in rats. Inflammopharmacology. 2023;31:1529–1538. doi: 10.1007/s10787-023-01208-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mikami T., Kim J., Park J., Lee H., Yaicharoen P., Suidasari S., Yokozawa M., Yamauchi K. Olive leaf extract prevents obesity, cognitive decline, and depression and improves exercise capacity in mice. Sci. Rep. 2021;11 doi: 10.1038/s41598-021-90589-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Blanchett S., Boal-Carvalho I., Layzell S., Seddon B. NF-kappaB and Extrinsic Cell Death Pathways - Entwined Do-or-Die Decisions for T cells. Trends Immunol. 2021;42:76–88. doi: 10.1016/j.it.2020.10.013. [DOI] [PubMed] [Google Scholar]
- 22.Geismann C., Hauser C., Grohmann F., Schneeweis C., Bölter N., Gundlach J.P., Schneider G., Röcken C., Meinhardt C., Schäfer H., et al. NF-kappaB/RelA controlled A20 limits TRAIL-induced apoptosis in pancreatic cancer. Cell Death Dis. 2023;14:3. doi: 10.1038/s41419-022-05535-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ponder K.G., Boise L.H. The prodomain of caspase-3 regulates its own removal and caspase activation. Cell Death Dis. 2019;5:56. doi: 10.1038/s41420-019-0142-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ji R., Ma L., Chen X., Sun R., Zhang L., Saiyin H., Wei W. Characterizing the distributions of IDO-1 expressing macrophages/microglia in human and murine brains and evaluating the immunological and physiological roles of IDO-1 in RAW264.7/BV-2 cells. PLoS One. 2021;16 doi: 10.1371/journal.pone.0258204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Noguchi M., Hirata N., Tanaka T., Suizu F., Nakajima H., Chiorini J.A. Autophagy as a modulator of cell death machinery. Cell Death Dis. 2020;11:517. doi: 10.1038/s41419-020-2724-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wu Y., Lin X., Song F., Xue D., Wang Y. Vitamin D3 promotes autophagy in THP-1 cells infected with Mycobacterium tuberculosis. Exp. Ther. Med. 2022;23:240. doi: 10.3892/etm.2022.11165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lee J.P.W., Foote A., Fan H., Peral de Castro C., Lang T., Jones S.A., Gavrilescu N., Mills K.H.G., Leech M., Morand E.F., Harris J. Loss of autophagy enhances MIF/macrophage migration inhibitory factor release by macrophages. Autophagy. 2016;12:907–916. doi: 10.1080/15548627.2016.1164358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tanida I., Ueno T., Kominami E. LC3 and Autophagy. Methods Mol. Biol. 2008;445:77–88. doi: 10.1007/978-1-59745-157-4_4. [DOI] [PubMed] [Google Scholar]
- 29.Leleu I., Genete D., Desnoulez S.S., Saidi N., Brodin P., Lafont F., Tomavo S., Pied S. A noncanonical autophagy is involved in the transfer of Plasmodium-microvesicles to astrocytes. Autophagy. 2022;18:1583–1598. doi: 10.1080/15548627.2021.1993704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Christian F., Smith E.L., Carmody R.J. The Regulation of NF-kappaB Subunits by Phosphorylation. Cells. 2016;5:12. doi: 10.3390/cells5010012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Riol-Blanco L., Delgado-Martín C., Sánchez-Sánchez N., Alonso-C L.M., Gutiérrez-López M.D., Del Hoyo G.M., Navarro J., Sánchez-Madrid F., Cabañas C., Sánchez-Mateos P., Rodríguez-Fernández J.L. Immunological synapse formation inhibits, via NF-kappaB and FOXO1, the apoptosis of dendritic cells. Nat. Immunol. 2009;10:753–760. doi: 10.1038/ni.1750. [DOI] [PubMed] [Google Scholar]
- 32.Tyagi R.K., Miles B., Parmar R., Garg N.K., Dalai S.K., Baban B., Cutler C.W. Human IDO-competent, long-lived immunoregulatory dendritic cells induced by intracellular pathogen, and their fate in humanized mice. Sci. Rep. 2017;7 doi: 10.1038/srep41083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Scicchitano S., Vecchio E., Battaglia A.M., Oliverio M., Nardi M., Procopio A., Costanzo F., Biamonte F., Faniello M.C. The Double-Edged Sword of Oleuropein in Ovarian Cancer Cells: From Antioxidant Functions to Cytotoxic Effects. Int. J. Mol. Sci. 2023;24 doi: 10.3390/ijms24010842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Mathieu L.C., Singh P., Monteiro W.M., Magris M., Cox H., Lazrek Y., Melo G.C., Marchesini P., Alexandre J.S.F., Alvarez A.M., et al. Kelch13 mutations in Plasmodium falciparum and risk of spreading in Amazon basin countries. J. Antimicrob. Chemother. 2021;76:2854–2862. doi: 10.1093/jac/dkab264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nair U., Cao Y., Xie Z., Klionsky D.J. Roles of the lipid-binding motifs of Atg18 and Atg21 in the cytoplasm to vacuole targeting pathway and autophagy. J. Biol. Chem. 2010;285:11476–11488. doi: 10.1074/jbc.M109.080374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kannan D., Joshi N., Gupta S., Pati S., Bhattacharjee S., Langsley G., Singh S. Cytoprotective autophagy as a pro-survival strategy in ART-resistant malaria parasites. Cell Death Dis. 2023;9:160. doi: 10.1038/s41420-023-01401-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.van Schalkwyk D.A., Burrow R., Henriques G., Gadalla N.B., Beshir K.B., Hasford C., Wright S.G., Ding X.C., Chiodini P.L., Sutherland C.J. Culture-adapted Plasmodium falciparum isolates from UK travellers: in vitro drug sensitivity, clonality and drug resistance markers. Malar. J. 2013;12:320. doi: 10.1186/1475-2875-12-320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Klonis N., Crespo-Ortiz M.P., Bottova I., Abu-Bakar N., Kenny S., Rosenthal P.J., Tilley L. Artemisinin activity against Plasmodium falciparum requires hemoglobin uptake and digestion. Proc. Natl. Acad. Sci. USA. 2011;108:11405–11410. doi: 10.1073/pnas.1104063108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dhorda M., Amaratunga C., Dondorp A.M. Artemisinin and multidrug-resistant Plasmodium falciparum - a threat for malaria control and elimination. Curr. Opin. Infect. Dis. 2021;34:432–439. doi: 10.1097/QCO.0000000000000766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cheruiyot J., Ingasia L.A., Omondi A.A., Juma D.W., Opot B.H., Ndegwa J.M., Mativo J., Cheruiyot A.C., Yeda R., Okudo C., et al. Polymorphisms in Pfmdr1, Pfcrt, and Pfnhe1 genes are associated with reduced in vitro activities of quinine in Plasmodium falciparum isolates from western Kenya. Antimicrob. Agents Chemother. 2014;58:3737–3743. doi: 10.1128/AAC.02472-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Andriantsoanirina V., Khim N., Ratsimbasoa A., Witkowski B., Benedet C., Canier L., Bouchier C., Tichit M., Durand R., Ménard D. Plasmodium falciparum Na+/H+ exchanger (pfnhe-1) genetic polymorphism in Indian Ocean malaria-endemic areas. Am. J. Trop. Med. Hyg. 2013;88:37–42. doi: 10.4269/ajtmh.2012.12-0359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Sudhakar R., Das D., Thanumalayan S., Gorde S., Sijwali P.S. Plasmodium falciparum Atg18 localizes to the food vacuole via interaction with the multi-drug resistance protein 1 and phosphatidylinositol 3-phosphate. Biochem. J. 2021;478:1705–1732. doi: 10.1042/BCJ20210001. [DOI] [PubMed] [Google Scholar]
- 43.Luckhart S., Giulivi C., Drexler A.L., Antonova-Koch Y., Sakaguchi D., Napoli E., Wong S., Price M.S., Eigenheer R., Phinney B.S., et al. Sustained activation of Akt elicits mitochondrial dysfunction to block Plasmodium falciparum infection in the mosquito host. PLoS Pathog. 2013;9 doi: 10.1371/journal.ppat.1003180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Day J., Passecker A., Beck H.P., Vakonakis I. The Plasmodium falciparum Hsp70-x chaperone assists the heat stress response of the malaria parasite. Faseb. J. 2019;33:14611–14624. doi: 10.1096/fj.201901741R. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Huet D., Rajendran E., van Dooren G.G., Lourido S. Identification of cryptic subunits from an apicomplexan ATP synthase. Elife. 2018;7 doi: 10.7554/eLife.38097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Benot-Dominguez R., Tupone M.G., Castelli V., d'Angelo M., Benedetti E., Quintiliani M., Cinque B., Forte I.M., Cifone M.G., Ippoliti R., et al. Olive leaf extract impairs mitochondria by pro-oxidant activity in MDA-MB-231 and OVCAR-3 cancer cells. Biomed. Pharmacother. 2021;134 doi: 10.1016/j.biopha.2020.111139. [DOI] [PubMed] [Google Scholar]
- 47.Moita D., Maia T.G., Duarte M., Andrade C.M., Albuquerque I.S., Dwivedi A., Silva J.C., González-Céron L., Janse C.J., Mendes A.M., Prudêncio M. A genetically modified Plasmodium berghei parasite as a surrogate for whole-sporozoite vaccination against P. vivax malaria. NPJ Vaccines. 2022;7:163. doi: 10.1038/s41541-022-00585-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hsu M.L., Huang W.C., Zhou Y.R., Hu S., Huang C.H., Wu S.J. Oleuropein Protects Human Retinal Pigment Epithelium Cells from IL-1beta-Induced Inflammation by Blocking MAPK/NF-kappaB Signaling Pathways. Inflammation. 2022;45:297–307. doi: 10.1007/s10753-021-01546-4. [DOI] [PubMed] [Google Scholar]
- 49.Pojero F., Aiello A., Gervasi F., Caruso C., Ligotti M.E., Calabrò A., Procopio A., Candore G., Accardi G., Allegra M. Effects of Oleuropein and Hydroxytyrosol on Inflammatory Mediators: Consequences on Inflammaging. Int. J. Mol. Sci. 2022;24 doi: 10.3390/ijms24010380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Serreli G., Deiana M. Extra Virgin Olive Oil Polyphenols: Modulation of Cellular Pathways Related to Oxidant Species and Inflammation in Aging. Cells. 2020;9:478. doi: 10.3390/cells9020478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pennisi R., Ben Amor I., Gargouri B., Attia H., Zaabi R., Chira A.B., Saoudi M., Piperno A., Trischitta P., Tamburello M.P., Sciortino M.T. Analysis of Antioxidant and Antiviral Effects of Olive (Olea europaea L.) Leaf Extracts and Pure Compound Using Cancer Cell Model. Biomolecules. 2023;13 doi: 10.3390/biom13020238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Burja B., Kuret T., Janko T., Topalović D., Živković L., Mrak-Poljšak K., Spremo-Potparević B., Žigon P., Distler O., Čučnik S., et al. Olive Leaf Extract Attenuates Inflammatory Activation and DNA Damage in Human Arterial Endothelial Cells. Front. Cardiovasc. Med. 2019;6:56. doi: 10.3389/fcvm.2019.00056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Filipek A., Mikołajczyk T.P., Guzik T.J., Naruszewicz M. Oleacein and Foam Cell Formation in Human Monocyte-Derived Macrophages: A Potential Strategy Against Early and Advanced Atherosclerotic Lesions. Pharmaceuticals. 2020;13 doi: 10.3390/ph13040064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Filipek A., Czerwińska M.E., Kiss A.K., Polański J.A., Naruszewicz M. Oleacein may inhibit destabilization of carotid plaques from hypertensive patients. Impact on high mobility group protein-1. Phytomedicine. 2017;32:68–73. doi: 10.1016/j.phymed.2017.06.004. [DOI] [PubMed] [Google Scholar]
- 55.Deregibus M.C., Buttiglieri S., Russo S., Bussolati B., Camussi G. CD40-dependent activation of phosphatidylinositol 3-kinase/Akt pathway mediates endothelial cell survival and in vitro angiogenesis. J. Biol. Chem. 2003;278:18008–18014. doi: 10.1074/jbc.M300711200. [DOI] [PubMed] [Google Scholar]
- 56.Zhang Y., Morgan M.J., Chen K., Choksi S., Liu Z.G. Induction of autophagy is essential for monocyte-macrophage differentiation. Blood. 2012;119:2895–2905. doi: 10.1182/blood-2011-08-372383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Jiang G.-M., Tan Y., Wang H., Peng L., Chen H.-T., Meng X.-J., Li L.-L., Liu Y., Li W.-F., Shan H. The relationship between autophagy and the immune system and its applications for tumor immunotherapy. Mol. Cancer. 2019;18:17. doi: 10.1186/s12943-019-0944-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Tougan T., Edula J.R., Morita M., Takashima E., Honma H., Tsuboi T., Horii T. The malaria parasite Plasmodium falciparum in red blood cells selectively takes up serum proteins that affect host pathogenicity. Malar. J. 2020;19:155. doi: 10.1186/s12936-020-03229-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tomlins A.M., Ben-Rached F., Williams R.A., Proto W.R., Coppens I., Ruch U., Gilberger T.W., Coombs G.H., Mottram J.C., Müller S., Langsley G. Plasmodium falciparum ATG8 implicated in both autophagy and apicoplast formation. Autophagy. 2013;9:1540–1552. doi: 10.4161/auto.25832. [DOI] [PubMed] [Google Scholar]
- 60.Setua S., Enguita F.J., Chora Â.F., Ranga-Prasad H., Lahree A., Marques S., Sundaramurthy V., Mota M.M. Disrupting Plasmodium UIS3-host LC3 interaction with a small molecule causes parasite elimination from host cells. Commun. Biol. 2020;3:688. doi: 10.1038/s42003-020-01422-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Agrawal P., Manjithaya R., Surolia N. Autophagy-related protein PfATG18 participates in food vacuole dynamics and autophagy-like pathway in Plasmodium falciparum. Mol. Microbiol. 2020;113:766–782. doi: 10.1111/mmi.14441. [DOI] [PubMed] [Google Scholar]
- 62.de Bock M., Thorstensen E.B., Derraik J.G.B., Henderson H.V., Hofman P.L., Cutfield W.S. Human absorption and metabolism of oleuropein and hydroxytyrosol ingested as olive (Olea europaea L.) leaf extract. Mol. Nutr. Food Res. 2013;57:2079–2085. doi: 10.1002/mnfr.201200795. [DOI] [PubMed] [Google Scholar]
- 63.Moss S., Mańko E., Vasileva H., Da Silva E.T., Goncalves A., Osborne A., Phelan J., Rodrigues A., Djata P., D'Alessandro U., et al. Population dynamics and drug resistance mutations in Plasmodium falciparum on the Bijagos Archipelago, Guinea-Bissau. Sci. Rep. 2023;13:6311. doi: 10.1038/s41598-023-33176-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Ward K.E., Christensen P., Racklyeft A., Dhingra S.K., Chua A.C.Y., Remmert C., Suwanarusk R., Matheson J., Blackman M.J., Kaneko O., et al. Integrative Genetic Manipulation of Plasmodium cynomolgi Reveals Multidrug Resistance-1 Y976F Associated With Increased In Vitro Susceptibility to Mefloquine. J. Infect. Dis. 2023;227:1121–1126. doi: 10.1093/infdis/jiac469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Amambua-Ngwa A., Button-Simons K.A., Li X., Kumar S., Brenneman K.V., Ferrari M., Checkley L.A., Haile M.T., Shoue D.A., McDew-White M., et al. Chloroquine resistance evolution in Plasmodium falciparum is mediated by the putative amino acid transporter AAT1. Nat. Microbiol. 2023;8:1213–1226. doi: 10.1038/s41564-023-01377-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ward K.E., Fidock D.A., Bridgford J.L. Plasmodium falciparum resistance to artemisinin-based combination therapies. Curr. Opin. Microbiol. 2022;69 doi: 10.1016/j.mib.2022.102193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Li F.J., Shen Q., Wang C., Sun Y., Yuan A.Y., He C.Y. A role of autophagy in Trypanosoma brucei cell death. Cell Microbiol. 2012;14:1242–1256. doi: 10.1111/j.1462-5822.2012.01795.x. [DOI] [PubMed] [Google Scholar]
- 68.Ji S.T., Kim Y.J., Jung S.Y., Kim D.Y., Kang S., Park J.H., Jang W.B., Ha J., Yun J., Kwon S.M. Oleuropein attenuates hydrogen peroxide-induced autophagic cell death in human adipose-derived stem cells. Biochem. Biophys. Res. Commun. 2018;499:675–680. doi: 10.1016/j.bbrc.2018.03.211. [DOI] [PubMed] [Google Scholar]
- 69.de Oca M.M., Engwerda C., Haque A. Plasmodium berghei ANKA (PbA) infection of C57BL/6J mice: a model of severe malaria. Methods Mol. Biol. 2013;1031:203–213. doi: 10.1007/978-1-62703-481-4_23. [DOI] [PubMed] [Google Scholar]
- 70.Andrews K.A., Wesche D., McCarthy J., Möhrle J.J., Tarning J., Phillips L., Kern S., Grasela T. Model-Informed Drug Development for Malaria Therapeutics. Annu. Rev. Pharmacol. Toxicol. 2018;58:567–582. doi: 10.1146/annurev-pharmtox-010715-103429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Coppens I. Metamorphoses of malaria: the role of autophagy in parasite differentiation. Essays Biochem. 2011;51:127–136. doi: 10.1042/bse0510127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Reiss A.B., Carsons S.E., Anwar K., Rao S., Edelman S.D., Zhang H., Fernandez P., Cronstein B.N., Chan E.S.L. Atheroprotective effects of methotrexate on reverse cholesterol transport proteins and foam cell transformation in human THP-1 monocyte/macrophages. Arthritis Rheum. 2008;58:3675–3683. doi: 10.1002/art.24040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Sharma P., Sharma V., Ahluwalia T.S., Dogra N., Kumar S., Singh S. Let-7a induces metabolic reprogramming in breast cancer cells via targeting mitochondrial encoded ND4. Cancer Cell Int. 2021;21:629. doi: 10.1186/s12935-021-02339-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Pan C., Kang J., Hwang J.S., Li J., Boese A.C., Wang X., Yang L., Boggon T.J., Chen G.Z., Saba N.F., et al. Cisplatin-mediated activation of glucocorticoid receptor induces platinum resistance via MAST1. Nat. Commun. 2021;12:4960. doi: 10.1038/s41467-021-24845-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Sharma P., Singh S. Combinatorial Effect of DCA and Let-7a on Triple-Negative MDA-MB-231 Cells: A Metabolic Approach of Treatment. Integr. Cancer Ther. 2020;19 doi: 10.1177/1534735420911437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Abdelbary G., Haider M. In vitro characterization and growth inhibition effect of nanostructured lipid carriers for controlled delivery of methotrexate. Pharmaceut. Dev. Technol. 2013;18:1159–1168. doi: 10.3109/10837450.2011.614251. [DOI] [PubMed] [Google Scholar]
- 77.Ramachandran S., Kota P., Ding F., Dokholyan N.V. Automated minimization of steric clashes in protein structures. Proteins. 2011;79:261–270. doi: 10.1002/prot.22879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Trott O., Olson A.J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010;31:455–461. doi: 10.1002/jcc.21334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Liu Y., Grimm M., Dai W.T., Hou M.C., Xiao Z.X., Cao Y. CB-Dock: a web server for cavity detection-guided protein-ligand blind docking. Acta Pharmacol. Sin. 2020;41:138–144. doi: 10.1038/s41401-019-0228-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Sudhakar R., Adhikari N., Pamnani S., Panda A., Bhattacharjee M., Rizvi Z., Shehzad S., Gupta D., Sijwali P.S. Bazedoxifene, a Postmenopausal Drug, Acts as an Antimalarial and Inhibits Hemozoin Formation. Microbiol. Spectr. 2022;10 doi: 10.1128/spectrum.02781-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Trager W., Jensen J.B. Human malaria parasites in continuous culture. Science (New York, N.Y.) 1976;193:673–675. doi: 10.1126/science.781840. [DOI] [PubMed] [Google Scholar]
- 82.Lambros C., Vanderberg J.P. Synchronization of Plasmodium falciparum erythrocytic stages in culture. J. Parasitol. 1979;65:418–420. [PubMed] [Google Scholar]
- 83.Prasad R., Atul K.,V.K., Kolla V.K., Legac J., Singhal N., Navale R., Rosenthal P.J., Sijwali P.S. Blocking Plasmodium falciparum development via dual inhibition of hemoglobin degradation and the ubiquitin proteasome system by MG132. PLoS One. 2013;8 doi: 10.1371/journal.pone.0073530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Vossen M.G., Pferschy S., Chiba P., Noedl H. The SYBR Green I malaria drug sensitivity assay: performance in low parasitemia samples. Am. J. Trop. Med. Hyg. 2010;82:398–401. doi: 10.4269/ajtmh.2010.09-0417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Varga A., James D. Real-time RT-PCR and SYBR Green I melting curve analysis for the identification of Plum pox virus strains C, EA, and W: effect of amplicon size, melt rate, and dye translocation. J. Virol. Methods. 2006;132:146–153. doi: 10.1016/j.jviromet.2005.10.004. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
-
•
This study does not report any original code.
-
•
All unique/stable reagents generated in this study are available from the lead contact without restriction.
-
•
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.