Figure 3.
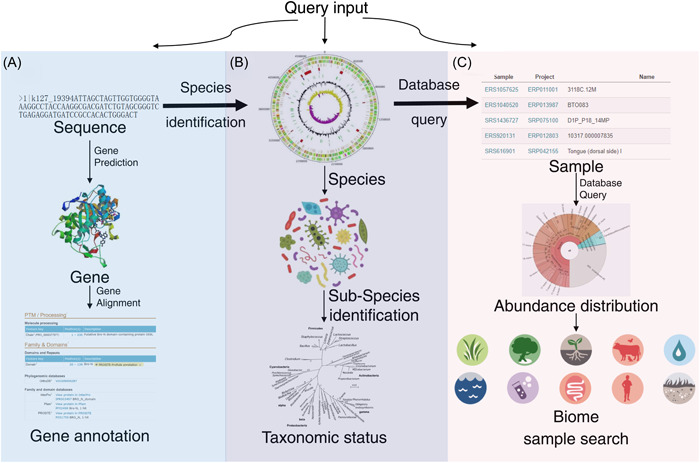
Investigation the gene‐species‐sample relationship with one sequence (https://microexpert.aimicrobiome.cn/search). (A) Gene annotation workflow. For the query nucleotide sequences, the MicroEXPERT platform searches the genes in the NCBI nucleotide database using the tool BLAST+ (version 2.5.1). The detailed matched information based on the NCBI nucleotide database are also illustrated. (B) Species identification workflow. To identify the possible source species of the query sequence, Kraken2 (version 2.0.7) and MetaPhlAn4 (version 4.0.2) are used. The pedigree‐based and phylogenetic relationship of the matched species is deduced using the NCBI taxonomy database. (C) Sample search workflow. To explore the distribution of samples and the biome of the query species in the database, all analysis results and sample information are stored in a separate table to return the abundance information of the query species in different environments and samples. The samples containing the queried species are illustrated according to their relative abundance in the related samples.
