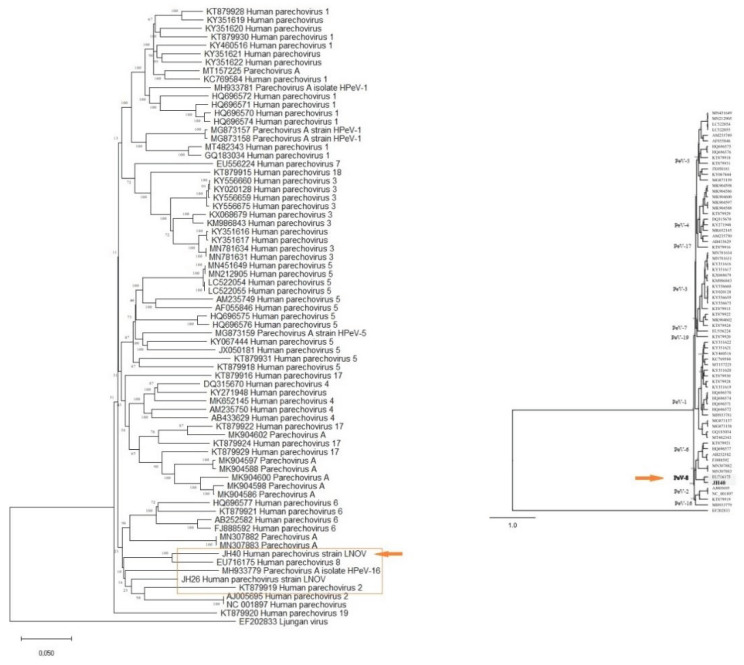
Figure 3.
(A) Phylogenomic tree of parechovirus. The evolutionary distances were computed using the Maximum Composite Likelihood method and are in the units of the number of base substitutions per site. This analysis involved 83 nucleotide sequences in MEGA X. (B) A phylogenetic tree of the complete genome of parechovirus created using Bayesian inference in the Geneious software with MrBayes. The program produced consensus trees and partition tables summarizing the trees and branch lengths. The consensus trees were written with both branch lengths and posterior clade probabilities.