Extended Data Fig. 5 |. Proteins with similar functions are correlated across the dataset.
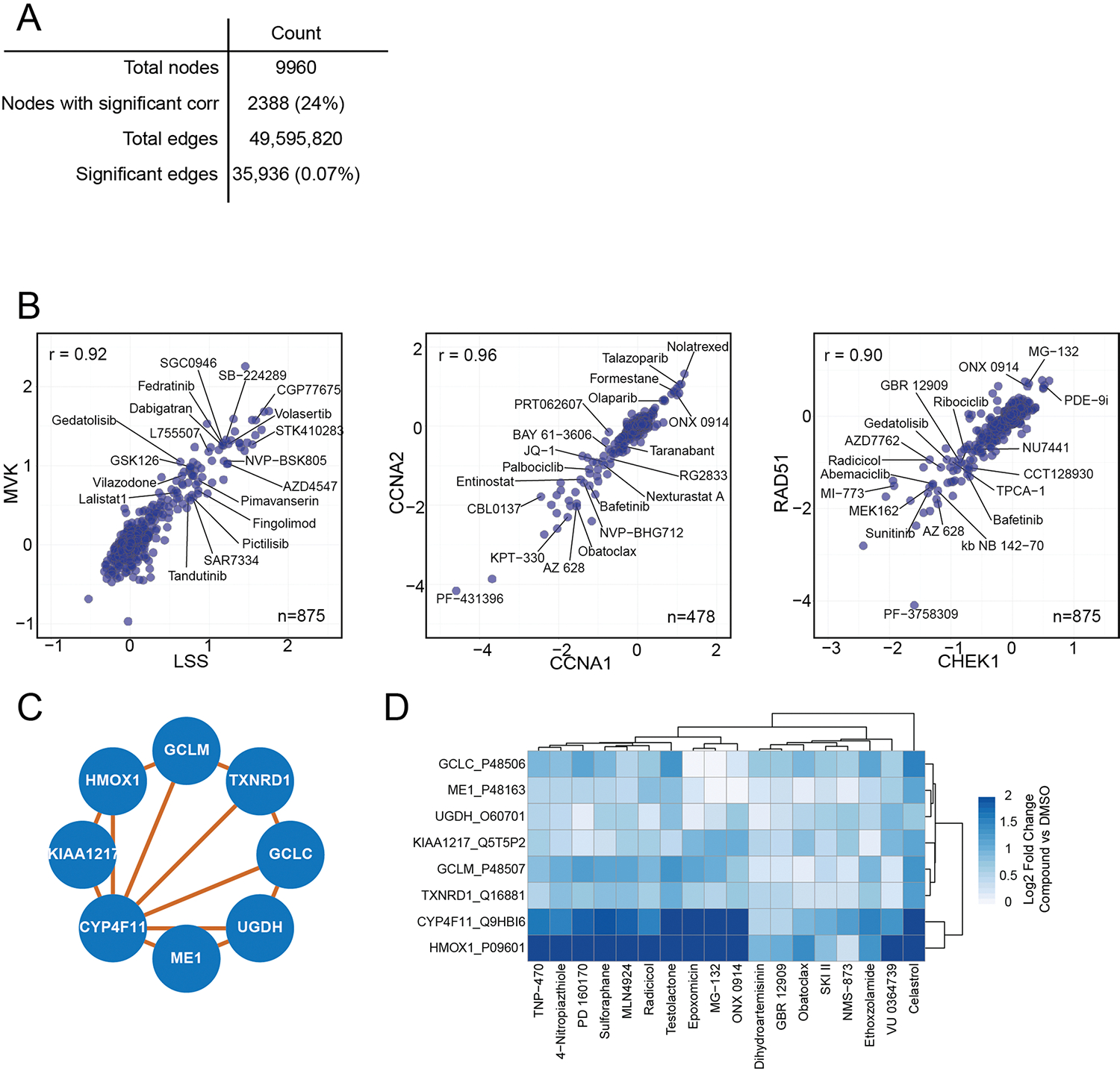
(a) A table highlighting the number of nodes and edges in the overall network from Fig. 5a, after filtering to a FDR less than 5%. The percentage of nodes and edges included in the network out of the total possible is shown in parentheses. (b) Correlation plots for select pairs of strongly correlated proteins. All x- and y-axes are represented as Log2 fold change versus DMSO. Each point represents one compound. Plots include: 1) mevalonate kinase (MVK) versus lanosterol synthase (LSS), two proteins essential for cholesterol biosynthesis; 2) cyclin A1 (CCNA1) and cyclin A2 (CCNA2); 3) DNA damage proteins RAD51 and CHEK1. (c) A subcommunity of proteins (from Fig. 5a) correlated to cytochrome P450 monooxygenase, including redox sensors HMOX and TXNRD1. (d) A heatmap of protein expression for each of the proteins from (c) for all compounds that increase their expression by a median Log2 fold change of >0.6.
