Abstract
Hepatocellular carcinoma (HCC) is the main type of primary liver cancer. This study aimed to develop a basement membrane (BM) related lncRNAs risk signature to evaluate the prognosis of HCC patients. We screened differentially expressed BM-related lncRNAs (DE-BMRlncRNAs) for risk evaluation, and identified six DE-BMRlncRNAs (AC072054.1, NUP50-DT, AC026412.3, AC109322.2, POLH-AS1 and LINC00595) for prognostic risk signature. HCC patients were divided to high or low risk according to median risk score. Our prognostic model predicted that patients with higher risk score had worse prognosis. We also created a nomogram to assist clinical decision-making according to risk score and clinicopathological features. Meanwhile, we confirmed the expression of six lncRNAs in HCC tissue and cells. POLH-AS1 knockdown inhibited the migration and invasion of HCC cells. In conclusion, we established a predictive model based on BMRlncRNAs to predict the prognosis of HCC. Our findings offer a rationale to further explore BM-related biomarkers for HCC.
Keywords: Basement membrane, Hepatocellular carcinoma, lncRNAs, Biomarkers, Prediction model
1. Introduction
Liver cancer is the third leading cause of all cancer related deaths [1]. Hepatocellular carcinoma (HCC) contributes to 75–85 % of all liver cancer and its incidence has increased in recent decades [2,3]. Several risk factors are related to the development of HCC such as infection with hepatitis B virus and dietary fat intake [[4], [5], [6], [7]].
Although the prognosis of HCC patients has improved due to therapeutic strategies such as chemoradiotherapy and immune checkpoint blockade, the overall prognosis is still not satisfactory because of insidious onset, difficult early diagnosis, rapid progression, and the risk of recurrence and metastasis [8]. In addition, there are many limitations of current imaging techniques or potential biomarkers used in clinical practice such as the need for validation in larger patient cohorts and the lack of standardized cutoff values [9].
Basement membrane (BM) is located at the base of endothelial and epithelial tissues, about 50–100 nm in thickness, and is a component of extracellular matrix (ECM). ECM plays important role in tumor micro-environment (TME) [10]. ECM promotes primary tumor growth by providing a framework for a stem cell-derived cancer ecosystem, and facilitates distant metastasis by creating a milieu conducive to tumor cell proliferation and invasion [11]. Therefore, ECM dysregulation is a key factor in tumor development, and BM proteins in particular are potential biomarkers of cancer progression [12,13].
As a special type of non-coding RNAs, lncRNAs regulate tumor cell proliferation, invasion, differentiation and apoptosis [14]. Several lncRNAs have been identified that can regulate BM degradation, and therefore warrant further evaluation as diagnostic and prognostic biomarkers. For instance, a prognostic signature based on BM-associated lncRNAs has been developed for bladder cancer [15]. However, a possible connection of BM-related lncRNAs to the prognosis of HCC has not been reported so far. Therefore, in this study we developed an independent prognostic signature of HCC according to differentially expressed BM-related lncRNAs (DE-BMRlncRNAs), and verified the expression of DE-BMRlncRNAs in HCC cells and tissue.
2. Materials and methods
2.1. Data collection
The clinical information and RNA-seq of 374 HCC samples and 50 adjacent tissues were collected from TCGA datasets (https://cancergenome.nih.gov/). BM related genes were acquired from BM-BASE database.
2.2. Identification of DE-BMRlncRNAs
We first took the intersection of BM genes in BM-BASE and HCC mRNAs in TCGA. Differentially expressed BM genes (DE-BMGs) were screened as described previously [16]. Then, BM-related lncRNAs were screened through Pearson correlation analysis between DE-BMGs and lncRNAs extracted from the LIHC transcriptome data. Finally, DE-BMRlncRNAs were identified as described previously [16].
2.3. Development of the prognostic signature
Total 374 HCC samples derived from TCGA were randomly divided into test and training sets. Preliminarily screened prognosis-related DE-BMRlncRNAs were analyzed by LASSO regression to construct a DE-BMRlncRNA signature: risk score of prognostic signature = Ʃin (each DE-BMRlncRNA's expression × corresponding coefficient).
Based on risk score, all patients were separated into high or low risk group. The Kaplan-Meier (K-M) survival analysis demonstrated overall survival (OS) of the two risk groups. Receiver operating characteristic (ROC) curves of 5-year, 3-year, and 1-year risk scores were drawn as described previously [17]. The prognostic value of the risk model was analyzed by univariate and multivariate Cox regression. In addition, a nomogram was created according to risk score of DE-BMRlncRNAs signature and clinicopathological factors of 315 HCC patients for predicting the outcome in HCC patients as described previously [18].
2.4. Tumor mutational burden (TMB) evaluation
R package was utilized for the analysis of somatic mutation data [19]. According to median value of TMB, HCC patients were separated to high or low TMB group.
2.5. Immune analysis based on prognostic signature
CIBERSORT was used to analyze the abundance of 22 immune cells in HCC samples. The comparison of immune checkpoint activation in high and low risk groups were visualized by R packages ggplot2 and ggpubr [20].
2.6. Exploration of the model in clinical treatment
The anti-tumor drug data were obtained from database of Genomics of Cancer Drug Sensitivity (GDSC), and IC50 (half maximal inhibitory concentration) of the drugs in HCC patients were analyzed as described previously [21].
2.7. Tissue collection
Total 15 paired HCC cancer tissues and adjacent normal tissues were collected from HCC patients during surgery and frozen immediately. The protocol was approved by Ethics Committee of Second Affiliated Hospital of Nanjing Medical University and all patients provided informed consent.
2.8. Cell culture
Four human HCC cell lines (HepG2, Huh7,7721 and Hep3B) and one normal human liver cell lines (L02) used in this study were stored in our laboratory. All cells were cultured in DMEM (GIBCO, USA) supplemented with 10 % fetal bovine serum (FBS) (GIBCO, USA) in humid incubator with 5 % CO2 at 37 °C.
2.9. PCR
The total RNA from cells and tissues was isolated using TRIzolR (Invitrogen, USA) and PCR was performed with SYBRR PCR MasterMix (Invitrogen, USA). All reactions were performed in triplicate and repeated for three times. β-actin was utilized as control and gene expression was analyzed by 2-△△ct method.
2.10. Cell based assays
SiRNA for POLH-AS1 (si-POLH-AS1) and negative control siRNA (si-NC) were transfected into HepG2 and Huh7 cells by using Lipofectamine 2000 (Invitrogen). The viability of HCC cells was analyzed with CCK-8 kit (Krbio). The migration and invasion of HCC cells were analyzed as described in our previous study [22]. All assays were in triplicate and repeated for three times.
2.11. Statistical analysis
All data were analyzed by R version 4.1.2. ANOVA was used for multiple comparisons and adjusted by Bonferroni test if necessary. P-value <0.05 indicated significant difference.
3. Results
3.1. Acquisition of DE-BMRlncRNAs
Based on the intersection of 222 BM genes and 374 HCC genes from TCGA database, 87 DE-BMGs (79 upregulated and 8 downregulated) (Fig. 1A) were obtained. Then, we analyzed the relationship between 16,773 lncRNAs and 87 DE-BMGs. Total 2467 lncRNAs were extracted as BM-related lncRNAs (BMRlncRNAs). Ultimately, 1286 DE-BMRlncRNAs were identified (Fig. 1B).
Fig. 1.
Screening for DE-BMGs and DE-BMRlncRNAs. (A) Volcano plot of 87 DE-BMGs. (B) Volcano plot of 1286 DE-BMRlncRNAs.
3.2. Construction of prognostic risk model
Total 534 prognosis-related DE-BMRlncRNAs in the training set were extracted by univariable regression. After multivariate and LASSO Cox regression analysis, six DE-BMRlncRNAs (AC072054.1, NUP50-DT, AC026412.3, AC109322.2, POLH-AS1, LINC00595) were selected to construct risk model (Fig. 2A and B). The six DE-BMRlncRNAs showed significant differences in expression between the adjacent and tumor groups in TCGA (Fig. 2C). The six DE-BMRlncRNAs were listed in Table 1.
Fig. 2.
Development of a prognostic risk model based on BM-related lncRNAs and expression of 6 DE-BMRlncRNAs. (A) 10-fold cross-validation in LASSO Cox regression of 6 DE-BMRlncRNAs. (B) The LASSO coefficient profile of 6 DE-BMRlncRNAs. (C) Expression of 6 DE-BMRlncRNAs (AC072054.1, NUP50-DT, AC026412.3, AC109322.2, POLH-AS1, and LINC00595) in HCC and adjacent para-cancerous tissues from the TCGA database.
Table 1.
Six DE-BMRlncRNA genes.
| Gene name | Gene ID | mRNA | cor | pvalue |
|---|---|---|---|---|
| AC072054.1 | ENSG00000286874 | RPSA | 0.483663386 | 2.52E-23 |
| AC026412.3 | ENSG00000271119 | ADAMTS6 | 0.415042407 | 5.26E-17 |
| AC026412.3 | ENSG00000271119 | AGRN | 0.453954632 | 2.06E-20 |
| AC026412.3 | ENSG00000271119 | ADAM9 | 0.443887408 | 1.73E-19 |
| AC026412.3 | ENSG00000271119 | ITGA5 | 0.483914333 | 2.38E-23 |
| AC026412.3 | ENSG00000271119 | ITGAV | 0.539185402 | 1.36E-29 |
| AC109322.2 | ENSG00000288096 | ROBO3 | 0.411345043 | 1.05E-16 |
| POLH-AS1 | ENSG00000203362 | LAMC1 | 0.413507223 | 7.03E-17 |
| NUP50-DT | ENSG00000226328 | BCAN | 0.405035686 | 3.38E-16 |
| LINC00595 | ENSG00000230417 | ADAMTS9 | 0.417846937 | 3.09E-17 |
| LINC00595 | ENSG00000230417 | AGRN | 0.560999727 | 2.17E-32 |
| LINC00595 | ENSG00000230417 | ITGA5 | 0.500209020 | 4.49E-25 |
cor, correalation.
OS rate, risk score, survival and related expression criteria for six DE-BMRlncRNAs were evaluated by risk scoring formula in entire set (n = 315), training set (n = 159) and test set (n = 156). The results indicated that high-risk group had worse prognosis (Fig. 3A-L).
Fig. 3.
Relevance between 6 DE-BMRlncRNAs and prognosis in a risk model. The Kaplan–Meier curves for OS, heatmap, risk score distribution, and survival time and survival status in the entire (A–D), training (E–H), and test sets (I–L), respectively.
Univariate and multivariate regression indicated that risk score based on prognostic signature can be served as standalone prognostic risk index. The hazard ratio and 95 % confidence interval (CI) of risk score were 1.075 and 1.054–1.096 (p < 0.001) in univariate regression (Fig. 4A), and 1.059 and 1.036–1.081 (P < 0.001) in multivariate regression (Fig. 4B).
Fig. 4.
Assessment of independent prognostic performance of 6 DE-BMRlncRNAs signature. (A,B) Forest plot of univariate and multivariate Cox regression analyses of clinical variables and risk score. (C–E) Time-dependent ROC curves of predicting 1-, 3-, and 5-year total survival for HCC patients in the entire, training, and test sets. (F–H) Comparison of predicted OS of the prognostic signature with clinic factors in the entire, training and test sets.
We further verified prediction performance of six DE-BMRlncRNAs signature based on area under ROC curve (AUC). The AUC for 1-year, 3-year and 5-year was 0.772, 0.733, and 0.745, respectively, in entire set; 0.866, 0.840, and 0.859, in training set; 0.690, 0.623, and 0.630 in test set (Fig. 4C–E). The 1-year ROC curve indicated that risk score and clinical characteristics (T and stage) had predictive efficiency in 3 sets (Fig. 4F–H).
3.3. Nomogram and clinical indicators
Next, we constructed a nomogram to predict 1-year, 3-year and 5-year OS (Fig. 5A). The risk score showed the highest consistency (Fig. 5B and C). K-M survival analysis showed that high-risk group with age, male, G1-2, G3-4, T1-2 and T3-4 disease had poorer prognosis (Supplementary Figs. 1A–F).
Fig. 5.
Construction and evaluation of the nomogram of the risk model. (A) Nomogram accompanied risk scores and clinical characteristics to predict 1-, 3-, and 5-year OS. (B) Concordance index of risk score and clinical features. (C) Assessment of the nomogram by establishing the calibration curves. *p < 0.05, **p < 0.01, and ***p < 0.001.
3.4. Functional and pathway enrichment analysis
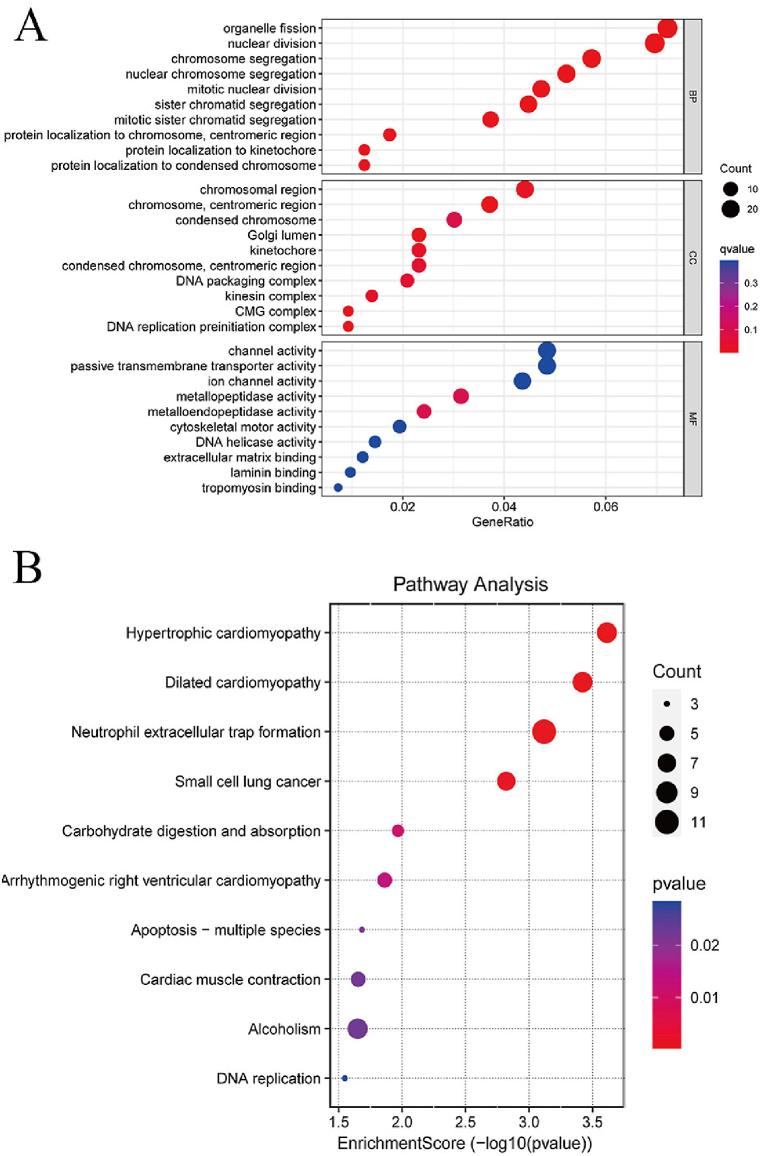
We performed GO and KEGG analysis of DEGs in high and low risk groups. Based on GO enrichment analysis, DEGs were enriched in HCC-related biological processes, including Chromosome segregation, Mitotic nuclear division and organelle fission (Fig. 6A). Based on KEGG analysis, DEGs were enriched in Hypertrophic cardiomyopathy, Dilated cardiomyopathy and Neutrophil extracellular trap formation (Fig. 6B).
Fig. 6.
Enrichment analysis of the DEGs in HCC. (A) Bubble graph for GO enrichment analysis (B) Bubble graph for KEGG pathway analysis. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; BP, Biological process; CC, Cellular component; MF, Molecular function.
3.5. TMB analysis
HCC patients were divided to low TMB group (n = 133) and high TMB group (n = 130) (Supplementary Figs. 2A and B). TP53 (35 %), TTN(22 %), CTNNB1(20 %), and MUC16(18 %) were frequently mutated genes in high-risk group, while CTNNB1(31 %), TP53(20 %), TTN(20 %) and MUC16(15 %) were frequently mutated genes in low-risk group. The overall genome mutation occurrence rate was 84.18 % and 87.25 % in high and low risk group, respectively. TMB score in two risk groups had no significant difference (Supplementary Fig. 2C), but the survival of high TMB group was shorter than that of low TMB group (Supplementary Fig. 2D). Patients with high risk score and high TMB had the worst prognosis, while patients with low risk score and low TMB had significantly better prognosis (Supplementary Fig. 2E).
3.6. Immune correlation analysis
The patients in low-risk group had higher TME scores across three categories (ESTIMATE, immune and stromal score) (Fig. 7A), and they had high number of CD8+T cells, B cells, Mast cells, Neutrophils, NK cells, pDCs, and TIL (Fig. 7B). Analysis of the expression of 29 immune checkpoints genes showed that the levels of most immune checkpoints were higher in high-risk group (Fig. 7C). Analysis of 13 immune-related pathways indicated that high-risk group had weaker immune activity compared to low-risk group (Fig. 7D). Additionally, we divided HCC patients to four immune subtypes (C1 to C4) and found that C1 had the highest risk score (Fig. 7E). IC50 analysis of four anti-tumor drugs (ABT.263, AG.014699, AICAR, and AMG.706) showed significant difference in two groups (Supplementary Figs. 3A–D).
Fig. 7.
Immunological characteristics in the high- and low-risk groups. (A) TME scores (Stromal cell score, immune cell score, and ESTIMATE score) in high-risk and low-risk groups. (B) Distribution of the enrichment scores of immune cells in high- and low-risk groups. (C) The differential expression of the 29 immune checkpoints in the high- and low-risk groups. (D) Distribution of the enrichment scores of immune-related pathways in high- and low-risk group. (E) The correlations of risk score with immune subtype. *P < 0.05, **P < 0.01, ***P < 0.001.
3.7. POLH-AS1 promoted the proliferation, migration and invasion of HCC cells
Six lncRNAs (AC072054.1, NUP50-DT, AC026412.3, AC109322.2, POLH-AS1 and LINC00595) were potential biomarkers of HCC. PCR showed that their expression was consistent in HCC tissues and cell lines (Fig. 8A and B). After knockdown of POLH-AS1 in HepG2 and Huh7 cells, cell viability decreased (Fig. 8C) and cell migration and invasion also decreased (Fig. 8D and E).
Fig. 8.
POLH-AS1 promoted HCC cell proliferation, migration and invasion. (A.B). The expression of POLH-AS1 was high in HCC tissues and cells. (C). POLH-AS1 promoted HCC cell proliferation. (D.E). POLH-AS1 promoted HCC cell migration and invasion.
4. Discussion
Hepatocytes have no characterized BM in the healthy liver parenchyma [23]. Liver fibrosis caused by viral or chronic metabolic diseases promotes the formation of hepatocyte BM, which may contribute to HCC progression [24]. A recent study identified 222 BM-related genes in HCC and other cancers [25]. Recently, different prognostic signatures were used to predict the prognosis of HCC patients [26,27]. However, there are currently no reports on prognostic relevance of BM related lncRNAs for HCC.
Therefore, this study reported novel prognostic risk model on the basis of six BMRlncRNAs for HCC. The expression of these BMRlncRNAs was significantly higher in tumor tissues than in normal tissues, suggesting their important role in HCC. HCC patients were divided to high and low risk group based on risk score, and the former had worse prognosis compared to the latter. A nomogram based on lncRNA prognostic signature showed good predictive efficacy in HCC patients. TMB can be used to discover potential oncogenic pathways [28,29]. In this study, TP53, CTNNB1 and TTN were main mutated genes in both risk groups, and patients in high risk group with high TMB had the worst prognosis.
TME plays important role in HCC development and progression, and impacts patient survival, prognosis and immunotherapy response [[30], [31], [32]]. A recent study reported the panoptosis-related gene model to characterize TME in HCC [33]. Overexpression of immune checkpoints would suppress tumor immune response in TME, and prevents the killing of tumor cells [34]. Immune checkpoint inhibitors (ICIs) have reshaped anti-cancer therapy in recent decades, and have achieved long-lasting remissions that led to improved survival in 15–20 % HCC patients [35]. In our study, patients in low risk group showed higher TME score, and immune checkpoint-related genes were mostly overexpressed in high risk group, indicative of an immunosuppressive TME. Therefore, HCC patients with high risk score may respond to ICIs, and our risk model may aid the selection of appropriate immune checkpoint target [36,37]. In addition, we validated six BMRlncRNAs in liver tissues and human HCC cells, and found that their expression levels were consistent with previous study [38].
Furthermore, we selected one of six BMRlncRNAs POLH-AS1 for further analysis. POLH-AS1 was upregulated in HCC cells, and enhanced HCC cell proliferation, migration, and invasion. However, other five BMRlncRNAs should be characterized as well. A very recent study established novel immune-associated prognostic signature for HCC [39]. This study did not explore molecular mechanisms by which BMRlncRNAs regulate HCC progression and metastasis, which is another direction for further studies.
In conclusion, we established a BMRlncRNAs related model to predict the prognosis of HCC patients. Furthermore, risk score showed significant correlation with TMB and immunotherapy response, and therefore can aid the selection of appropriate treatment approach. Our findings provide a reasonable explanation for the distinct outcomes of HCC patients, and offer a rationale to further explore BM-related biomarkers in HCC.
Data availability statement
All data are available upon request to correspondence author.
CRediT authorship contribution statement
Shenmei Xun: Methodology. Xiaocui Li: Project administration. Qingyuan Zhuang: Software, Data curation. Yefei Zhu: Writing – review & editing, Conceptualization. Lili Qu: Writing – original draft, Funding acquisition.
Declaration of competing interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests:
Lili Qu reports financial support was provided by Jiangsu Provincial Medical Association Laboratory Medicine Research Special Fund Project. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgement
This study was funded by Jiangsu Provincial Medical Association Laboratory Medicine Research Special Fund Project (No. SYH-3201160-0059) and Nanjing Health Science and Technology Development Project (No. YKK23285).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2024.e30439.
Appendix A. Supplementary data
The following is/are the supplementary data to this article.
References
- 1.Sung H., Ferlay J., Siegel R.L., Laversanne M., Soerjomataram I., Jemal A., et al. Global cancer statistics 2020: GLOBOCAN Estimates of incidence and Mortality Worldwide for 36 cancers in 185 Countries. CA Cancer J Clin. 2021;71(3):209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 2.Toh M.R., Wong E.Y.T., Wong S.H., Ng A.W.T., Loo L.H., Chow P.K., Ngeow J. Global Epidemiology and Genetics of hepatocellular carcinoma. Gastroenterology. 2023;164(5):766–782. doi: 10.1053/j.gastro.2023.01.033. [DOI] [PubMed] [Google Scholar]
- 3.Siegel R.L., Miller K.D., Wagle N.S., Jemal A. Cancer statistics. CA Cancer J. Clin. 2023. 2023;73:17–48. doi: 10.3322/caac.21763. [DOI] [PubMed] [Google Scholar]
- 4.Li J., Li H., Zeng H., Zheng R., Cao M., Sun D., He S., Ren J., Chen W., He J. Trends in high-risk rates and screening rates for the population-based cancer screening program on esophageal, stomach and liver cancer in China, 2010-2016. Journal of the National Cancer Center. 2021;1(3):101–107. doi: 10.1016/j.jncc.2021.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Liu T., Diao H. A double-edged sword: the HBV-induced non-coding RNAs alterations in hepatocellular carcinoma. Biocell. 2023;47(1):27–32. [Google Scholar]
- 6.Zhang X., Wang Y., Yang G. Research progress in hepatitis B virus covalently closed circular DNA. Cancer Biol Med. 2021;19(4):415–431. doi: 10.20892/j.issn.2095-3941.2021.0454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ji X., Wang J., Li Z., Shen Q., Tuo J., Bi J., Tan Y., Li H., Xiang Y. Dietary fat intake and liver cancer risk: a prospective cohort study in Chinese women. Cancer Biol Med. 2021;19(3):370–383. doi: 10.20892/j.issn.2095-3941.2020.0633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zeng W., Ling F., Dang K., Chi Q. SPP1 and the risk score model to improve the survival prediction of patients with hepatocellular carcinoma based on multiple algorithms and back propagation neural networks. Biocell. 2023;47(3):581–592. [Google Scholar]
- 9.Raees A., Kamran M., Özkan H., Jafri W. Updates on the diagnosis and management of hepatocellular carcinoma. Euroasian J. Hepato-Gastroenterol. 2021;11(1):32–40. doi: 10.5005/jp-journals-10018-1335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Goenka A., Khan F., Verma B., Sinha P., Dmello C.C., Jogalekar M.P., Gangadaran P., Ahn B.C. Tumor microenvironment signaling and therapeutics in cancer progression. Cancer Commun. 2023;43(5):525–561. doi: 10.1002/cac2.12416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li C., Qiu S., Liu X., Guo F., Zhai J., Li Z., Deng L., Ge L., Qian H., Yang L., Xu B. Extracellular matrix-derived mechanical force governs breast cancer cell stemness and quiescence transition through integrin-DDR signaling. Signal Transduct Target Ther. 2023;8(1):247. doi: 10.1038/s41392-023-01453-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yuan Z., Li Y., Zhang S., Wang X., Dou H., Yu X., Zhang Z., Yang S., Xiao M. Extracellular matrix remodeling in tumor progression and immune escape: from mechanisms to treatments. Mol. Cancer. 2023;22(1):48. doi: 10.1186/s12943-023-01744-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang K., Qiu X., Zhao Y., Wang H., Chen L. The Wnt/β-catenin signaling pathway in the tumor microenvironment of hepatocellular carcinoma. Cancer Biol Med. 2021;19(3):305–318. doi: 10.20892/j.issn.2095-3941.2021.0306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yu J., Mao W., Sun S., Hu Q., Wang C., Xu Z., et al. Identification of an m6A-related lncRNA signature for predicting the prognosis in patients with Kidney Renal Clear cell carcinoma. Front. Oncol. 2021;11 doi: 10.3389/fonc.2021.663263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Feng L., Yang J., Zhang W., Wang X., Li L., Peng M., et al. Prognostic significance and identification of basement membrane-associated lncRNA in bladder cancer. Front. Oncol. 2022;12 doi: 10.3389/fonc.2022.994703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chi M., Liu J., Mei C., Shi Y., Liu N., Jiang X., et al. TEAD4 functions as a prognostic biomarker and triggers EMT via PI3K/AKT pathway in bladder cancer. J. Exp. Clin. Cancer Res. 2022;41(1):175. doi: 10.1186/s13046-022-02377-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Deng Y., Zhang F., Sun Z.G., Wang S. Development and validation of a prognostic signature associated with tumor microenvironment based on Autophagy-related lncRNA analysis in hepatocellular carcinoma. Front. Med. 2021;8 doi: 10.3389/fmed.2021.762570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tang M., Wang H., Cao Y., Zeng Z., Shan X., Wang L. Nomogram for predicting occurrence and prognosis of liver metastasis in colorectal cancer: a population-based study. Int. J. Colorectal Dis. 2021;36(2):271–282. doi: 10.1007/s00384-020-03722-8. [DOI] [PubMed] [Google Scholar]
- 19.Wang Y., Tan H., Yu T., Chen X., Jing F., Shi H. Potential immune biomarker Candidates and immune subtypes of lung Adenocarcinoma for developing mRNA Vaccines. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.755401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cheng Q., Chen X., Wu H., Du Y. Three hematologic/immune system-specific expressed genes are considered as the potential biomarkers for the diagnosis of early rheumatoid arthritis through bioinformatics analysis. J. Transl. Med. 2021;19(1):18. doi: 10.1186/s12967-020-02689-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Che Y., Jiang D., Xu L., Sun Y., Wu Y., Liu Y., et al. The clinical prediction value of the Ubiquitination model Reflecting the immune Traits in LUAD. Front. Immunol. 2022;13 doi: 10.3389/fimmu.2022.846402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shen P., Qu L., Wang J., Ding Q., Zhou C., Xie R., Wang H., Ji G. LncRNA LINC00342 contributes to the growth and metastasis of colorectal cancer via targeting miR-19a-3p/NPEPL1 axis. Cancer Cell Int. 2021;21:105. doi: 10.1186/s12935-020-01705-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chang T.T., Thakar D., Weaver V.M. Force-dependent breaching of the basement membrane. Matrix Biol. 2017;57–58:178–189. doi: 10.1016/j.matbio.2016.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tatrai P., Dudas J., Batmunkh E., Mathe M., Zalatnai A., Schaff Z., et al. Agrin, a novel basement membrane component in human and rat liver, accumulates in cirrhosis and hepatocellular carcinoma. Lab. Invest. 2006;86(11):1149–1160. doi: 10.1038/labinvest.3700475. [DOI] [PubMed] [Google Scholar]
- 25.Jayadev R., Morais M., Ellingford J.M., Srinivasan S., Naylor R.W., Lawless C., et al. A basement membrane discovery pipeline uncovers network complexity, regulators, and human disease associations. Sci. Adv. 2022;8(20) doi: 10.1126/sciadv.abn2265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Xu Z., Peng B., Liang Q., Chen X., Cai Y., Zeng S., Gao K., Wang X., Yi Q., Gong Z., Yan Y. Construction of a Ferroptosis-related Nine-lncRNA signature for predicting prognosis and immune response in hepatocellular carcinoma. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.719175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang B., Tang B., Gao J., Li J., Kong L., Qin L. A hypoxia-related signature for clinically predicting diagnosis, prognosis and immune microenvironment of hepatocellular carcinoma patients. J. Transl. Med. 2020;18(1):342. doi: 10.1186/s12967-020-02492-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cai Y., Wang X., Wang N., Wu J., Ma L., Xie X., et al. Correlations between tumor mutation burden and immune infiltrates and their prognostic value in pancreatic cancer by bioinformatic analysis. Life Sci. 2021;277 doi: 10.1016/j.lfs.2021.119505. [DOI] [PubMed] [Google Scholar]
- 29.Klebanov N., Artomov M., Goggins W.B., Daly E., Daly M.J., Tsao H. Burden of unique and low prevalence somatic mutations correlates with cancer survival. Sci. Rep. 2019;9(1):4848. doi: 10.1038/s41598-019-41015-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Meng L., Zheng Y., Liu H., Fan D. The tumor microenvironment: a key player in multidrug resistance in cancer. Oncologie. 2024;26(1):41–58. [Google Scholar]
- 31.Zhang Y., Qin N., Wang X., Liang R., Liu Q., Geng R., Jiang T., Liu Y., Li J. Glycogen metabolism-mediated intercellular communication in the tumor microenvironment influences liver cancer prognosis. Oncology Research. 2024;32(3):563–576. doi: 10.32604/or.2023.029697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wang F., Yang M., Luo W., Zhou Q. Characteristics of tumor microenvironment and novel immunotherapeutic strategies for non-small cell lung cancer. Journal of the National Cancer Center. 2022;2(4):243–262. doi: 10.1016/j.jncc.2022.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shi X., Gao X., Liu W., Tang X., Liu J., Pan D., Duan X., Jin Y., Ren W., Yang L., Liu W. Construction of the panoptosis-related gene model and characterization of tumor microenvironment infiltration in hepatocellular carcinoma. Oncology Research. 2023;31(4):569–590. doi: 10.32604/or.2023.028964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pardoll D.M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer. 2012;12(4):252–264. doi: 10.1038/nrc3239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sangro B., Chan S.L., Meyer T., Reig M., El-Khoueiry A., Galle P.R. Diagnosis and management of toxicities of immune checkpoint inhibitors in hepatocellular carcinoma. J. Hepatol. 2020;72(2):320–341. doi: 10.1016/j.jhep.2019.10.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang W., Ye Y., Zhang X., Ye X., Liu C., Bao L. Construction of a Necroptosis-associated long non-coding RNA signature to predict prognosis and immune response in hepatocellular carcinoma. Front. Mol. Biosci. 2022;9 doi: 10.3389/fmolb.2022.937979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kono K., Nakajima S., Mimura K. Current status of immune checkpoint inhibitors for gastric cancer. Gastric Cancer. 2020;23(4):565–578. doi: 10.1007/s10120-020-01090-4. [DOI] [PubMed] [Google Scholar]
- 38.Qu J., Tao D., Huang W., Lu L., Fan J., Zhang S., Huang F. Assessment of prognostic role of a novel 7-lncRNA signature in HCC patients. Heliyon. 2023;9(8) doi: 10.1016/j.heliyon.2023.e18493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lin X., Tian C., Pan F., Wang R. A novel immune-associated prognostic signature based on the immune cell infiltration analysis for hepatocellular carcinoma. Oncologie. 2024;26(1):91–103. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data are available upon request to correspondence author.