Abstract
Natural killer (NK) cells are lymphocytes of the innate immune system that sense target cells through a panel of activating and inhibitory receptors. Together with NKG2D, the natural cytotoxicity receptors (NCRs) are major activating receptors involved in tumor cell detection. Although numerous NKG2D ligands have been identified, characterization of the molecules interacting with the NCRs is still incomplete. The identification of B7-H6 as a counter structure of the NCR NKp30 shed light on the molecular basis of NK cell immunosurveillance. We review here the current knowledge on NKp30 and B7-H6, and we discuss their potential role in anti-tumor immunity.
Keywords: NK cell, Activating receptor, NKp30, B7-H6, Tumor immunosurveillance
Introduction
NK cells were identified in the early 1970s as lymphocytes endowed with spontaneous cytotoxic activity against tumor cells without prior specific host sensitization [1, 2]. First described as “killer lymphocytes with undefined nature”, they are now recognized as very sophisticated effectors, playing a key role in innate immunity and in the orchestration of the adaptive immunity in response to allogeneic, transformed, as well as infected cells [3].
NK cells monitor their targets through a panel of activating and inhibitory receptors expressed at their surface. Integration of the opposing signals transduced by the engagement of such receptors defines the functional outcome for NK cells [4]. Inhibitory receptors include the human KIR and the rodent Ly49 molecules, and their interaction with major histocompatibility complex (MHC) class I molecules on target cells prevents activation of mature NK cells. Thus, downregulation of normally ubiquitously expressed MHC class I molecules on target cells activates NK cells, a process coined as “missing self” recognition [5]. NK cell stimulation is also induced by several activating receptors that provide the “on” signals. Identification and characterization of some of their ligands showed that NK cells are able to sense self molecules induced by cells in distress but also infectious non-self molecules produced by microbe-infected cells [6].
In humans, the major activating receptors include NKG2D and the NCRs NKp46, NKp30, and NKp44. NKG2D is a surface receptor shared by NK and T cells that recognizes several inducible self ligands that vary in structure, distribution, and regulation [7]. This field of investigation was instrumental in refining the molecular understanding of the NK cell alert system. However, the picture is still incomplete, because the nature of the NCR ligands and their characterization is in its beginnings. Although a large piece of data suggested a central role of NCRs in tumor surveillance [8–11], the ligands identified first were viral structures, in particular the influenza hemagglutinin as a ligand for NKp46 [12] as well as NKp44 [13] and the human cytomegalovirus pp65 tegument protein for NKp30 [14]. Later, the HLA-B-associated transcript 3 (BAT3) protein was shown to bind and trigger NKp30 [15]. This nuclear protein is ectopically found at the plasma membrane upon stress, but is not reported to be spontaneously expressed on tumor cell lines susceptible to NK cell lysis. Recently, we identified B7-H6, a previously unannotated gene that encodes a potent ligand for NKp30 [16]. This molecule is not detectable on normal tissues at steady state but present on a broad panel of hematopoietic and non-hematopoietic tumor cells and thus could trigger NK cells through the induced-self recognition mode. This review will provide an overview on the knowledge of the NKp30 receptor and its ligand B7-H6 in the light of already characterized NK cell receptor/ligand pairs.
NKp30: a major NK cell-activating receptor with unique function
NKp30 was identified at the end of the 1990s by Pende and colleagues as a novel 30-kDa triggering receptor expressed by all resting and activated human NK cells [10]. It is a type I transmembrane protein characterized by a single V-type immunoglobulin (Ig) extracellular domain. The intracellular tail of NKp30 has no signaling motif but its transmembrane domain associates via a charged amino acid with immunoreceptor tyrosine-based activation motif (ITAM)-bearing adapters (CD3ζ and FcRγ). NKp30 is mostly restricted to NK cells although recent studies have shown protein expression on umbilical cord blood T cells after IL-15 stimulation and on endometrial epithelial cells following progesterone exposure [17, 18]. Moreover, the recently identified IL-22-producing mucosal innate lymphoid cell population that shares properties with LTi (lymphoid tissue inducer) and NK cells also expresses NKp30 [19, 20]. NKp30 has been shown to be a major activating receptor involved in tumor cell lysis, and importantly, it plays a unique role among the activating NK cell receptors because it also targets non-transformed self cells such as dendritic cells (DC). In vitro experiments have shown that NKp30 engagement on NK cells induces the TNF-dependent maturation of monocyte-derived DC and leads to the killing of DC that did not acquire a mature phenotype [21, 22]. The nature of the ligand interacting with NKp30 during this crosstalk remains to be determined.
Few data reporting NKp30 regulation on NK cells are available. It has been shown that IL-2 induces its up-regulation [23], whereas TGFβ1 down-regulates NKp30, leading to NK cell cytotoxicity impairment towards immature DC [24]. Leukemia cells also decrease NCR expression on NK cells. Particularly, longitudinal study of AML patients showed that an NKp30dull phenotype was acquired during leukemia development. The mechanism underlying this effect is still undefined, however, it can be hypothesized that this down-regulation is a mechanism whereby a tumor would escape innate immunity [25].
NKp30 is encoded by a gene located in the class III region of the human MHC on chromosome 6 where several other genes with immune function are found (tumor necrosis factor, lymphotoxins). NKp30 has no homology with NKp44 or NKp46 encoded on chromosomes 6 and 19, respectively, but based on their expression profile and function these receptors have been grouped as NCRs [10].
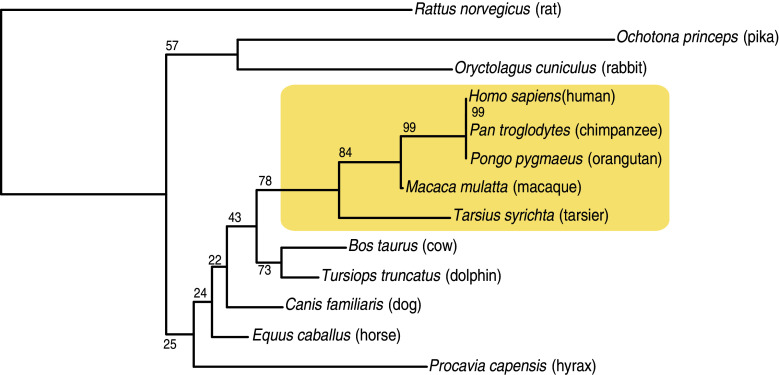
Orthologs of most known NK cell receptors including NKp30 are not present beyond the mammalian branching [26]. Phylogenic analysis of NKp30 in different mammalian species shows that gene sequences are very conserved except in exon 4 that encodes the intracellular domain. The tree constructed from amino acid alignment (Fig. 1) shows that two major clusters emerge, one including primates the other one including the rat. Notably NKp30 is a pseudogene in mice except in the wild strain Mus caroli [27]. A functional NKp30 protein is expressed on resting peripheral chimpanzee NK cells although at low level due to posttranscriptional regulation [28].
Fig. 1.
Phylogenic analysis of NKp30 protein sequences in mammals. The tree was constructed from CLUSTALW-generated amino acid alignments using the neighbor-joining method. Tree topography was evaluated by bootstrapping 1,000 times with percentage shown at nodes. Each node shows the percentage of the bootstrap replicates in which all of the sequences descended from that node were also descended from that node in the replicates. The tree is drawn to scale with branch lengths in the same units as those of the evolutionary distances used to infer phylogenetic tree. Phylogenetic analyses were conducted in Seaview. Primates are included in the shaded area
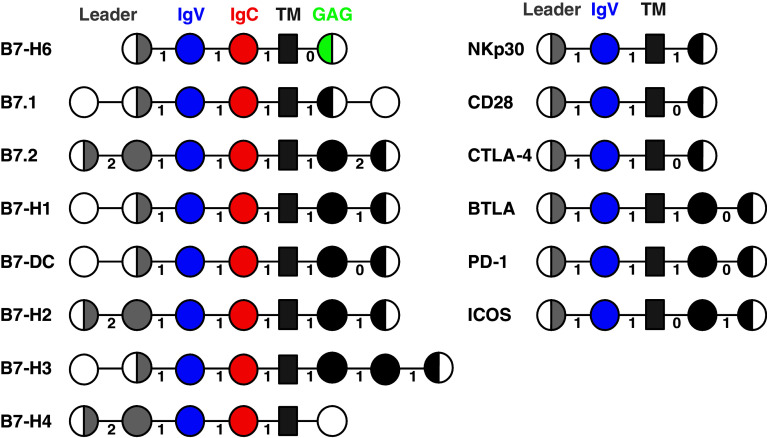
Based on BLAST homology searches, the closest homologue of NKp30 in humans is CTLA-4 (25% of identity), which is a weak but a significant hit. CTLA-4 is a member of the CD28 family whose members do not share significant sequence similarity but show a comparable gene structure. NKp30, as do the other members of this family, has a single Ig domain in its extracellular domain, while many other Ig domain-containing immunoreceptors have multiple Ig domains. Moreover, the structure and the exon patterning of the gene sequence encoding the extracellular region are identical between NKp30 and all other CD28 family genes (Fig. 2, right panel).
Fig. 2.
Gene organization of B7-H6 and NKp30 in comparison to the other members of the B7 and CD28 family. Each box represents an exon encoding the indicated domain (leader in grey, IgV in blue, IgC in red, transmembrane in dark grey, GAG domain in green, untranslated region in white, and others in black). Numbers between boxes indicate the intron phase. Canonical sequences of each gene were used to design these schemes (www.uniprot.org)
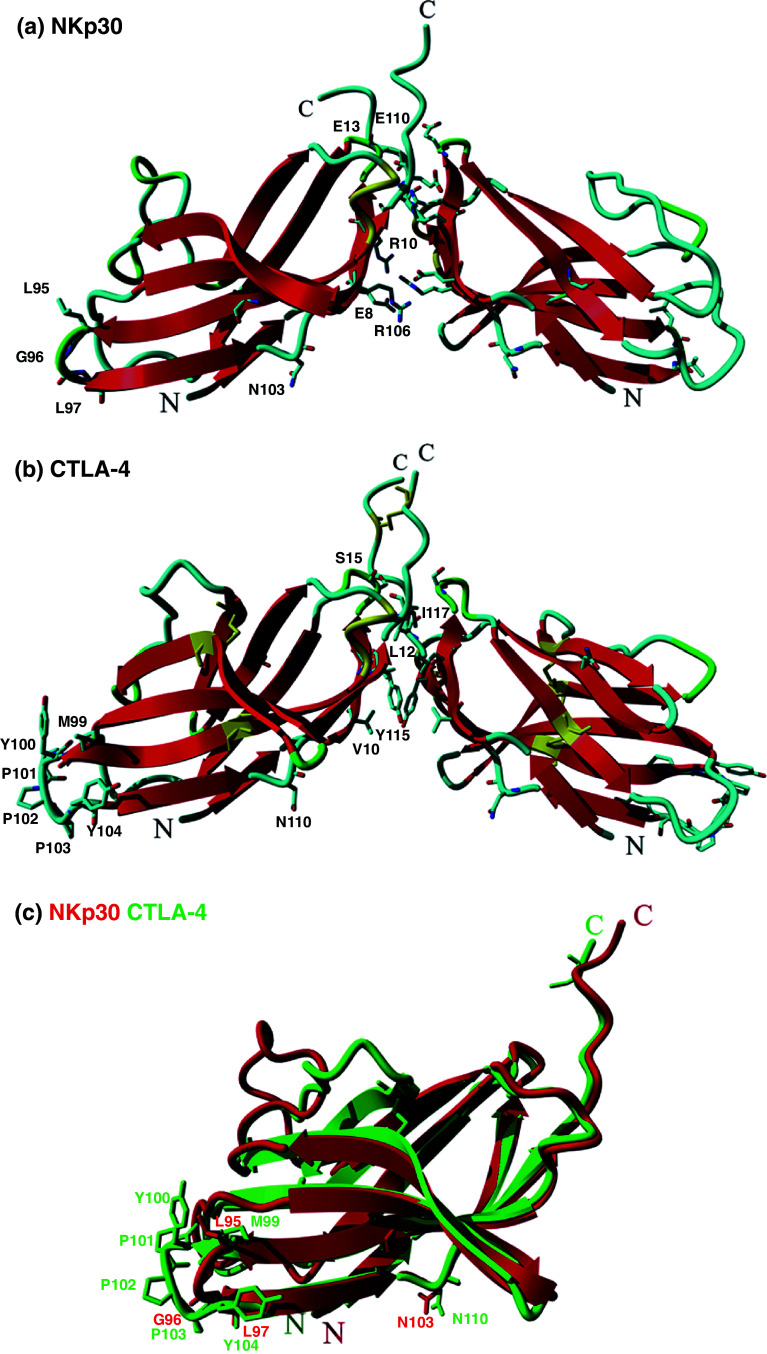
The structure of NKp30 is still not defined, however, homology modeling of the NKp30 ectodomain based on the CTLA-4 structure could be performed despite its relative weak sequence similarity. Prediction of the NKp30 ectodomain (19–126) structure was achieved using the YASARA homology modeling approach using, as templates, the structure of the CTLA-4 homodimer (3OSK) as well as the structure of the complexes B7.1/CTLA-4 (1I8L) and B7.2/CTLA-4 (1I85). After secondary structure prediction, loop construction, rotamer selection, and molecular dynamics, the best models were scored using YASARA energy force fields parameters and stereochemistry [29]. Superimposition of the NKp30 model on CTLA-4 structure reveals major differences (Fig. 3): (a) the absence of the CTLA-4 FG CDR3 loop (M99-Y104) interacting with B7-1/B7-1 GFCC′ surface that is replaced by a turn containing residues L95, G96, L97. If this short region makes contact with the B7-H6 ligand it should be associated with another NKp30-specific region in order to increase its interaction; (b) the presence of a rich network of salt-bridges/hydrogen bonds forming the putative dimer interface occurring between complementary charged residues E8, R10, E13, R106, and E110 of both monomers. In comparison, the CTLA-4 dimer interface is based on aliphatic–aromatic residues; and (c) among the two N-glycosylation sites N78 and N110 of CTLA-4, the conserved N110 site (CTLA4/CD28 family) potentially implicated in stabilizing the CTLA-4 homodimer is also conserved in the NKp30 model (N103).
Fig. 3.
Comparison of the CTLA-4 homodimer crystallographic structure and the structure of the NKp30 homodimer obtained by homology modeling. a NKp30 homodimer model and b CTLA-4 homodimer crystallographic structure (3OSK). Both structures are shown in ribbon representation, respectively, β-strands colored in red, coil in cyan, and β-turns in green. Amino acid residues found important for CTLA-4 B7.1/B7.2 interaction, for N-glycosylation, and for dimer stabilization as well as their NKp30 modeled equivalents are represented as sticks colored according to elements and residues are one letter named and numbered. Cysteines are represented as sticks with yellow color. N- and C-termini of both proteins are labeled. c Superposition of NKp30 (red) and CTLA-4 (green) monomers
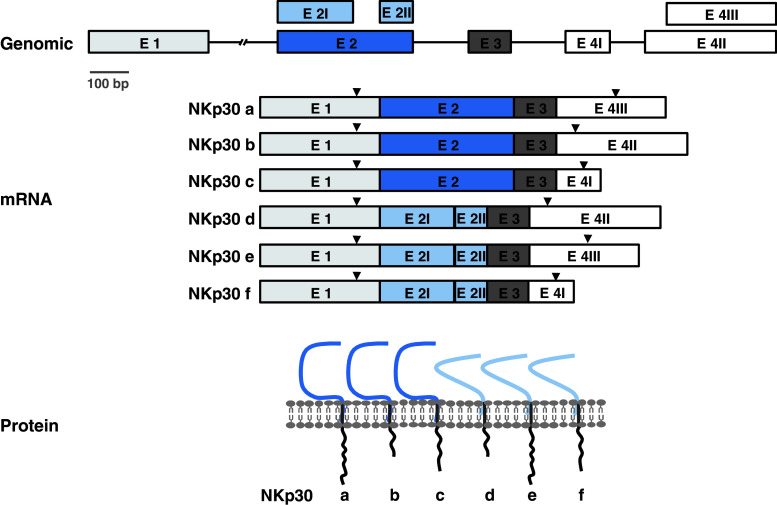
The NKp30 gene encodes six alternatively spliced transcripts (Fig. 4). Three of these (a, b, c) produce a V-type Ig extracellular domain and the three others (d, e, f) produce a C-type one. These extracellular domains are coupled with three different intracellular domains according to the exon 4 they utilize [30]. These six transcripts are not similarly distributed in tissues, and the V-type isoforms are the most abundant and ubiquitous ones. This heterogeneity in NKp30 isoforms leads to the hypothesis that engagement of each NKp30 molecule will differently trigger NK cell activation through interacting with different ligands as well as through association with distinct intracellular signaling proteins. Along this line Delahaye et al. [31] have recently shown using NKp30a, b, or c transfected NK cell lines that these three isoforms transmit distinct signals and thus mediate different cell functions; NKp30a and NKp30b are immunostimulatory isoforms that trigger TH1 cytokine production. In contrast NKp30c promotes IL-10 secretion, relaying an immunosuppressive signal through a rapid phosphorylation of p38 MAP kinase (see after for the reference). An additional layer of regulation could be provided by the existence of several glycosylation profiles of NKp30 [17]. At present, these important issues have to be unveiled.
Fig. 4.
The human NKp30 gene encodes six splice variants. The genomic organization of the exons contributing to the six transcript variants (NKp30a–f) is represented in the top panel. In transcripts a, b, and c the exon 2 is used, whereas in transcripts d, e, and f, both exon 2I and 2II are used, resulting in a 75-nt deletion. The initiation and stop codons are indicated by the arrowheads. The bottom panel represents the protein structure of each variant. NKp30a–c have an IgV domain, whereas NKp30d–f have an IgC domain
NKp30 ligands
Although the NCRs were discovered more than 10 years ago, it is still not completely understood how they are engaged to trigger NK cells in tumor surveillance. The first NCR ligands to be identified were of viral origin. In 2005, the human cytomegalovirus (HCMV) tegument protein pp65 was shown to bind NKp30, but its engagement leads to a general inhibition of the NK cell ability to kill infected and tumor cells [14]. The underlying mechanism was characterized as a decrease of the activating signal due to the CD3ζ dissociation from NKp30 upon pp65 interaction. This strategy has been hypothesized to contribute to the generalized immune suppression observed during HCMV infection. NKG2D is also the target of viral immune evasion strategy, although not directly. For example, several MCMV and HCMV genes encode for molecules called immunoevasines (such as m152/gp40 or UL16) that prevent NKG2D ligands to reach the infected cell surface thereby diverting the NK cell attack [32, 33]. In contrast, NKp46 has been shown to directly interact with hemagglutinin and its engagement with these viral structures induced NK cell activation [12]. Along this line, influenza infection has been reported to be lethal in NKp46-deficient mice [34].
The intracellular BAT3 protein was found to bind and activate NKp30 [15]. Although BAT3 has a C-terminal nuclear localization signal and lacks a putative secretory leader peptide, it can be directed to the cell surface or even secreted after heat shock. Moreover, BAT3 was shown to be expressed on the membrane of exosomes released from immature DC [35]. Because NK-dependent lysis of immature DC and DC maturation are inhibited by BAT3 antiserum, the authors of this work suggested a critical role of BAT3 in the NK-DC crosstalk.
B7-H6: an extrinsic barrier against transformed cells
Several groups had shown the constitutive expression of NKp30 ligands on tumor cells by assessing the binding of soluble NKp30; however, none of the previously described ligands fulfilled this criterion. The identification of the surface protein B7-H6 as a potent NKp30 ligand reconciles this discrepancy [16]. Because this molecule (previously named DKFZp686O24166) has a gene and a protein structure similar to other members of the B7 family, plus, two of its closest homologs are B7-H1 and B7-H3, it was designated B7-H6. Although other proteins such as TPSNR (tapasin-related protein) showed a higher level of similarity, their gene structures were different from the one of B7-H6. Like other B7 family members, B7-H6 has two extracellular Ig domains encoded by exons with adjacent phase 1 introns (Fig. 2, left panel). Inclusion of B7-H6 into the B7 family is also justified by the fact that its counter structure NKp30 shares several properties with the CD28 family members that interact with B7 proteins. In contrast to B7.1 and B7.2 that recognize both CD28 and CTLA-4, B7-H6 is not promiscuous since it does not bind any other CD28 family members nor other NCRs [16].
The B7 family members are costimulatory and co-inhibitory transmembrane proteins that contribute to the immune response regulation. The two founders B7.1 and B7.2 provide a balance of positive and negative signals required for appropriate priming of naive T cells by interacting with CD28 and CTLA-4. The five other members, B7-H1, B7-DC, B7-H2, B7-H3, and B7-H4, described as B7 homologs, have a less restricted distribution and are endowed with broader functions (Table 1). In addition to their role in adaptive immune control, they have been involved in innate immune regulation but also in non-immunoregulatory functions [36]. The B7 family members do not share a high sequence similarity but have related structures that echo the diversity of the CD28-related receptors (Fig. 5). Among B7 homologs, B7-H2 and B7-H3 induce costimulation upon engagement of their receptors, although additional data also supported an inhibitory role of B7-H3 in different experiment settings. In contrast, B7-H1 and B7-H4 dampen the immune response, and their expression on a wide range of human cancers could contribute to immune evasion by promoting T cell suppression [37].
Table 1.
Characteristics of the B7 family members

B B lymphocyte, M monocyte/macrophage, DC dendritic cells, T T lymphocytes, NK natural killer cell
* Location in the human genome
** % identity of the ectodomain
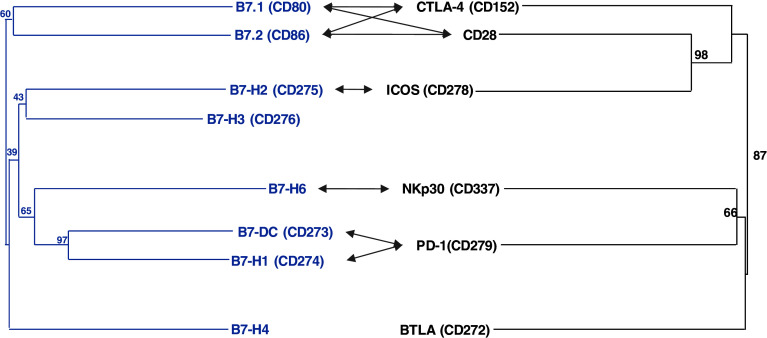
Fig. 5.
The B7 family mirrors the diversity of the CD28 members. Evolutionary relationships of human B7 (blue) and CD28 (black) family members were inferred using the neighbor-joining method. Tree topography was evaluated by bootstrapping 1,000 times with percentage shown at nodes (see legend of Fig. 1 for details). Arrows indicate the interacting receptors and ligands
B7-H6 has an intracytoplasmic domain homologous to GAG polyprotein, and contains signaling motifs such as a predicted immunoreceptor tyrosine-based inhibitory motif (ITIM) (SaYtpL), a Src homology 2 (SH2)-binding domain (YqlQ), and a Src homology 3 domain (SH3)-binding motif (PdaPilPvsP). These features suggest that upon engagement, B7-H6 could induce a response in the NK cell target through interaction with intracellular signaling proteins. This possibility has not been addressed for ligands of other activating receptors, but could have a significant impact on the target cells. Along this line, DNAM-1, an NK cell-triggering co-receptor, binds to CD155 and CD112, two transmembrane molecules involved in adherent junctions of epithelial cells and thus interacting with several intracellular proteins [38].
B7-H6 gene maps to the human chromosome region 11p15.1 and only four expressed sequence tags (EST), from testis, brain, liver, and embryonic kidney, were identified when searching with the human dbEST database. Phylogenic analysis of B7-H6 amino acid sequences in different species shows that B7-H6 is very conserved in primates but not in other mammals that miss the GAG intracellular domain (Fig. 6). In the mouse genome, the B7-H6-related sequence corresponds only to the first exon of the human gene and there are no related entries in mouse EST libraries. Hence, similarly to NKp30, but in contrast to other B7 members, a functional B7-H6 gene is missing in Mus musculus.
Fig. 6.
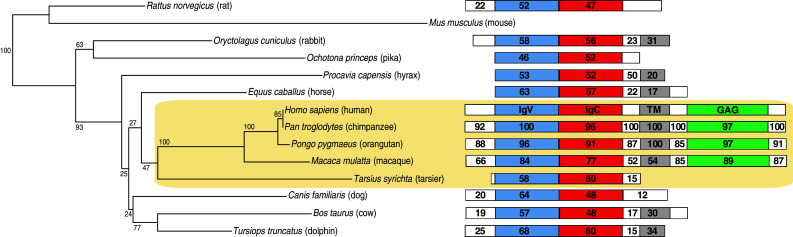
Phylogenic analysis of the B7-H6 protein sequences in mammals. Left part: the tree was constructed from CLUSTALW generated amino acid alignments using the neighbor-joining method. Tree topography was evaluated by bootstrapping 1,000 times with percentage shown at nodes (see legend of Fig. 1 for details). Phylogenetic analyses were conducted in Seaview. Primates are included in the shaded area. Right part: schematic representation of the predicted protein for each species. Percentage of similarity with the human B7-H6 protein is indicated for each domain (white leader or interdomain, blue IgV domain, red IgC domain, green GAG domain)
Strikingly, B7-H6 transcripts have not been detected in most normal adult tissues, consistent with the absence of the protein on circulating cells isolated from healthy individuals. In contrast, B7-H6 cell surface expression is observed in tumor cell lines from various origins including lymphoma, leukemia, melanoma, and carcinoma as well as on primary tumor blood cells. The pattern of B7-H6 expression, which appears so far to be limited to tumor cells, is another example of stress-induced self-recognition by NK cells. In the case of NKG2D ligands, several mechanisms have been shown to regulate their expression. For example, genotoxic stress was shown to up-regulate NKG2D ligands on non-tumor cell lines, whereas interfering with the DNA damage pathway in tumor cells inhibited their constitutive expression [39]. In pilot experiments, treatment of some NKp30 ligand-negative tumor cells with a panel of DNA-damaging agents had no major effect on B7-H6 expression. Several studies have also demonstrated that upon exposure to Toll-like receptor (TLR) ligands, human macrophages expressed NKG2D ligands that are sensed by NK cells, inducing a reciprocal regulation of the two partners [40, 41]. This TLR pathway has to be explored for B7-H6 to define whether non-tumor cells could be the target of NK cells through B7-H6/NKp30 interaction. Diverse cytokines such as TGF-β and particularly interferons also impact the regulation of NKG2D ligands, but depending on the cell types, they inhibit or increase expression of these ligands [42, 43]. Recently, different mechanisms have been described to control NKG2D ligand translation and protein turnover. For instance, the transcripts of the human NKG2D ligands MICA and MICB are the target of miRNAs that repress the protein expression [44]. Nothing is known yet concerning regulation of B7-H6 by miRNAs, however, two other members of the B7 family, B7-H1, and B7-H3 are also susceptible to RNA interference [45, 46]. Interestingly, the miRNA that targets B7-H1 is down-regulated in human cholangiocytes when exposed to IFN-γ, allowing expression of B7-H1 on their surface. Moreover, protein lysosomal degradation after ubiquitination has been shown to control the turnover of MULT1, another mouse NKG2D ligand. This process is altered in response to heat shock or UV radiation, consequently increasing MULT1 stability [47]. The proteasome machinery also impacts NKG2D ligand cell surface expression since MICA, MICB, as well as ULBP2 have been shown to be up-regulated after treatment with the proteasome inhibitor bortezomib [48, 49]. The implication of all these pathways (genotoxic agents, TLR ligands, cytokines, miRNA, proteasome inhibitors) in the control of B7-H6 expression remains to be explored. Finally, secreted forms of NKG2D ligands have been described in various human cancers. Protein shedding results from different processes that include ligand cleavage by proteases [50] and by RNA splicing [51]. Splice variant transcripts of B7-H6 have not been detected in normal tissue but analysis of cells from cancer patients is under study. This issue is crucial because soluble forms of activating ligands could provide an efficient way for tumor cells to evade NK cells immunosurveillance.
Concluding remarks
The identification of B7-H6 as a ligand for NKp30 unveiled a novel molecular mechanism of NK cell triggering. This finding prompts the characterization of B7-H6 expression, distribution, and regulation. The absence of B7-H6 transcripts in normal tissues and presence in tumor cells define B7-H6, as a novel example of a stress-induced self molecule. Longitudinal studies in tumor-bearing patients are now required to determine whether B7-H6 expression is not altered by tumor editing. If B7-H6 is specific to tumor cells and stable over time, immunotherapeutic strategies based on the development of cytotoxic antibody could be of major clinical interest.
Footnotes
Two papers describing NKp30 structures free and complexed with B7-H6 have been published during the preparation of this manuscript.
References
- 1.Herberman RB, Nunn ME, Lavrin DH. Natural cytotoxic reactivity of mouse lymphoid cells against syngeneic acid allogeneic tumors. I. distribution of reactivity and specificity. Int J Cancer. 1975;16:216–229. doi: 10.1002/ijc.2910160204. [DOI] [PubMed] [Google Scholar]
- 2.Kiessling R, Klein E, Wigzell H. “Natural” killer cells in the mouse. I. cytotoxic cells with specificity for mouse Moloney leukemia cells. Specificity and distribution according to genotype. Eur J Immunol. 1975;5:112–117. doi: 10.1002/eji.1830050208. [DOI] [PubMed] [Google Scholar]
- 3.Vivier E, Tomasello E, Baratin M, Walzer T, Ugolini S. Functions of natural killer cells. Nat Immunol. 2008;9:503–510. doi: 10.1038/ni1582. [DOI] [PubMed] [Google Scholar]
- 4.Vivier E, Nunes JA, Vely F. Natural killer cell signaling pathways. Science. 2004;306:1517–1519. doi: 10.1126/science.1103478. [DOI] [PubMed] [Google Scholar]
- 5.Karre K, Ljunggren HG, Piontek G, Kiessling R. Selective rejection of H-2-deficient lymphoma variants suggests alternative immune defence strategy. Nature. 1986;319:675–678. doi: 10.1038/319675a0. [DOI] [PubMed] [Google Scholar]
- 6.Lanier LL. NK cell recognition. Annu Rev Immunol. 2005;23:225–274. doi: 10.1146/annurev.immunol.23.021704.115526. [DOI] [PubMed] [Google Scholar]
- 7.Champsaur M, Lanier LL. Effect of NKG2D ligand expression on host immune responses. Immunol Rev. 2010;235:267–285. doi: 10.1111/j.0105-2896.2010.00893.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Byrd A, Hoffmann SC, Jarahian M, Momburg F, Watzl C. Expression analysis of the ligands for the natural killer cell receptors NKp30 and NKp44. PLoS One. 2007;2:e1339. doi: 10.1371/journal.pone.0001339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Castriconi R, Dondero A, Augugliaro R, Cantoni C, Carnemolla B, Sementa AR, Negri F, Conte R, Corrias MV, Moretta L, Moretta A, Bottino C. Identification of 4Ig-B7–H3 as a neuroblastoma-associated molecule that exerts a protective role from an NK cell-mediated lysis. Proc Natl Acad Sci USA. 2004;101:12640–12645. doi: 10.1073/pnas.0405025101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pende D, Parolini S, Pessino A, Sivori S, Augugliaro R, Morelli L, Marcenaro E, Accame L, Malaspina A, Biassoni R, Bottino C, Moretta L, Moretta A. Identification and molecular characterization of NKp30, a novel triggering receptor involved in natural cytotoxicity mediated by human natural killer cells. J Exp Med. 1999;190:1505–1516. doi: 10.1084/jem.190.10.1505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pende D, Rivera P, Marcenaro S, Chang CC, Biassoni R, Conte R, Kubin M, Cosman D, Ferrone S, Moretta L, Moretta A. Major histocompatibility complex class I-related chain A and UL16-binding protein expression on tumor cell lines of different histotypes: analysis of tumor susceptibility to NKG2D-dependent natural killer cell cytotoxicity. Cancer Res. 2002;62:6178–6186. [PubMed] [Google Scholar]
- 12.Mandelboim O, Lieberman N, Lev M, Paul L, Arnon TI, Bushkin Y, Davis DM, Strominger JL, Yewdell JW, Porgador A. Recognition of haemagglutinins on virus-infected cells by NKp46 activates lysis by human NK cells. Nature. 2001;409:1055–1060. doi: 10.1038/35059110. [DOI] [PubMed] [Google Scholar]
- 13.Arnon TI, Lev M, Katz G, Chernobrov Y, Porgador A, Mandelboim O. Recognition of viral hemagglutinins by NKp44 but not by NKp30. Eur J Immunol. 2001;31:2680–2689. doi: 10.1002/1521-4141(200109)31:9<2680::AID-IMMU2680>3.0.CO;2-A. [DOI] [PubMed] [Google Scholar]
- 14.Arnon TI, Achdout H, Levi O, Markel G, Saleh N, Katz G, Gazit R, Gonen-Gross T, Hanna J, Nahari E, Porgador A, Honigman A, Plachter B, Mevorach D, Wolf DG, Mandelboim O. Inhibition of the NKp30 activating receptor by pp65 of human cytomegalovirus. Nat Immunol. 2005;6:515–523. doi: 10.1038/ni1190. [DOI] [PubMed] [Google Scholar]
- 15.Pogge von Strandmann E, Simhadri VR, von Tresckow B, Sasse S, Reiners KS, Hansen HP, Rothe A, Boll B, Simhadri VL, Borchmann P, McKinnon PJ, Hallek M, Engert A. Human leukocyte antigen-B-associated transcript 3 is released from tumor cells and engages the NKp30 receptor on natural killer cells. Immunity. 2007;27:965–974. doi: 10.1016/j.immuni.2007.10.010. [DOI] [PubMed] [Google Scholar]
- 16.Brandt CS, Baratin M, Yi EC, Kennedy J, Gao Z, Fox B, Haldeman B, Ostrander CD, Kaifu T, Chabannon C, Moretta A, West R, Xu W, Vivier E, Levin SD. The B7 family member B7-H6 is a tumor cell ligand for the activating natural killer cell receptor NKp30 in humans. J Exp Med. 2009;206:1495–1503. doi: 10.1084/jem.20090681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ponnampalam AP, Gargett CE, Rogers PA. Identification and hormonal regulation of a novel form of NKp30 in human endometrial epithelium. Eur J Immunol. 2008;38:216–226. doi: 10.1002/eji.200636429. [DOI] [PubMed] [Google Scholar]
- 18.Tang Q, Grzywacz B, Wang H, Kataria N, Cao Q, Wagner JE, Blazar BR, Miller JS, Verneris MR. Umbilical cord blood T cells express multiple natural cytotoxicity receptors after IL-15 stimulation, but only NKp30 is functional. J Immunol. 2008;181:4507–4515. doi: 10.4049/jimmunol.181.7.4507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cupedo T, Crellin NK, Papazian N, Rombouts EJ, Weijer K, Grogan JL, Fibbe WE, Cornelissen JJ, Spits H. Human fetal lymphoid tissue-inducer cells are interleukin 17-producing precursors to RORC+ CD127+ natural killer-like cells. Nat Immunol. 2009;10:66–74. doi: 10.1038/ni.1668. [DOI] [PubMed] [Google Scholar]
- 20.Vivier E, Spits H, Cupedo T. Interleukin-22-producing innate immune cells: new players in mucosal immunity and tissue repair? Nat Rev Immunol. 2009;9:229–234. doi: 10.1038/nri2522. [DOI] [PubMed] [Google Scholar]
- 21.Ferlazzo G, Tsang ML, Moretta L, Melioli G, Steinman RM, Munz C. Human dendritic cells activate resting natural killer (NK) cells and are recognized via the NKp30 receptor by activated NK cells. J Exp Med. 2002;195:343–351. doi: 10.1084/jem.20011149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vitale M, Della Chiesa M, Carlomagno S, Pende D, Arico M, Moretta L, Moretta A. NK-dependent DC maturation is mediated by TNFalpha and IFNgamma released upon engagement of the NKp30 triggering receptor. Blood. 2005;106:566–571. doi: 10.1182/blood-2004-10-4035. [DOI] [PubMed] [Google Scholar]
- 23.Della Chiesa M, Carlomagno S, Frumento G, Balsamo M, Cantoni C, Conte R, Moretta L, Moretta A, Vitale M. The tryptophan catabolite L-kynurenine inhibits the surface expression of NKp46- and NKG2D-activating receptors and regulates NK-cell function. Blood. 2006;108:4118–4125. doi: 10.1182/blood-2006-03-006700. [DOI] [PubMed] [Google Scholar]
- 24.Castriconi R, Cantoni C, Della Chiesa M, Vitale M, Marcenaro E, Conte R, Biassoni R, Bottino C, Moretta L, Moretta A. Transforming growth factor beta 1 inhibits expression of NKp30 and NKG2D receptors: consequences for the NK-mediated killing of dendritic cells. Proc Natl Acad Sci USA. 2003;100:4120–4125. doi: 10.1073/pnas.0730640100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Fauriat C, Just-Landi S, Mallet F, Arnoulet C, Sainty D, Olive D, Costello RT. Deficient expression of NCR in NK cells from acute myeloid leukemia: evolution during leukemia treatment and impact of leukemia cells in NCRdull phenotype induction. Blood. 2007;109:323–330. doi: 10.1182/blood-2005-08-027979. [DOI] [PubMed] [Google Scholar]
- 26.Walzer T, Jaeger S, Chaix J, Vivier E. Natural killer cells: from CD3(−) NKp46(+) to post-genomics meta-analyses. Curr Opin Immunol. 2007;19:365–372. doi: 10.1016/j.coi.2007.04.004. [DOI] [PubMed] [Google Scholar]
- 27.Hollyoake M, Campbell RD, Aguado B. NKp30 (NCR3) is a pseudogene in 12 inbred and wild mouse strains, but an expressed gene in Mus caroli. Mol Biol Evol. 2005;22:1661–1672. doi: 10.1093/molbev/msi162. [DOI] [PubMed] [Google Scholar]
- 28.Rutjens E, Mazza S, Biassoni R, Koopman G, Radic L, Fogli M, Costa P, Mingari MC, Moretta L, Heeney J, De Maria A. Differential NKp30 inducibility in chimpanzee NK cells and conserved NK cell phenotype and function in long-term HIV-1-infected animals. J Immunol. 2007;178:1702–1712. doi: 10.4049/jimmunol.178.3.1702. [DOI] [PubMed] [Google Scholar]
- 29.Krieger E, Joo K, Lee J, Raman S, Thompson J, Tyka M, Baker D, Karplus K. Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: four approaches that performed well in CASP8. Proteins. 2009;77(Suppl 9):114–122. doi: 10.1002/prot.22570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Neville MJ, Campbell RD. A new member of the Ig superfamily and a V-ATPase G subunit are among the predicted products of novel genes close to the TNF locus in the human MHC. J Immunol. 1999;162:4745–4754. [PubMed] [Google Scholar]
- 31.Delahaye NF, Rusakiewicz S, Martins I, Ménard C, Roux S, Lyonnet L, Paul P, Sarabi M, Chaput N, Semeraro M, Minard-Colin V, Poirier-Colame V, Chaba K, Flament C, Baud V, Authier H, Kerdine-Römer S, Pallardy M, Cremer I, Peaudecerf L, Rocha B, Valteau-Couanet D, Celis Gutierrez J, Nunès JA, Commo F, Bonvalot S, Ibrahim N, Terrier P, Opolon P, Bottino C, Moretta A, Tavernier J, Rihet P, Coindre JM, Blay JY, Isambert N, Emile JF, Vivier E, Lecesne A, Kroemer G, Zitvogel L. Alternatively spliced NKp30 isoforms affect the prognosis of gastrointestinal stromal tumors. Nat Med. 2011;17:700–707. doi: 10.1038/nm.2366. [DOI] [PubMed] [Google Scholar]
- 32.Dunn C, Chalupny NJ, Sutherland CL, Dosch S, Sivakumar PV, Johnson DC, Cosman D. Human cytomegalovirus glycoprotein UL16 causes intracellular sequestration of NKG2D ligands, protecting against natural killer cell cytotoxicity. J Exp Med. 2003;197:1427–1439. doi: 10.1084/jem.20022059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhi L, Mans J, Paskow MJ, Brown PH, Schuck P, Jonjic S, Natarajan K, Margulies DH. Direct interaction of the mouse cytomegalovirus m152/gp40 immunoevasin with RAE-1 isoforms. Biochemistry. 2010;49:2443–2453. doi: 10.1021/bi902130j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gazit R, Gruda R, Elboim M, Arnon TI, Katz G, Achdout H, Hanna J, Qimron U, Landau G, Greenbaum E, Zakay-Rones Z, Porgador A, Mandelboim O. Lethal influenza infection in the absence of the natural killer cell receptor gene Ncr1. Nat Immunol. 2006;7:517–523. doi: 10.1038/ni1322. [DOI] [PubMed] [Google Scholar]
- 35.Simhadri VR, Reiners KS, Hansen HP, Topolar D, Simhadri VL, Nohroudi K, Kufer TA, Engert A, Pogge von Strandmann E. Dendritic cells release HLA-B-associated transcript-3 positive exosomes to regulate natural killer function. PLoS One. 2008;3:e3377. doi: 10.1371/journal.pone.0003377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Greenwald RJ, Freeman GJ, Sharpe AH. The B7 family revisited. Annu Rev Immunol. 2005;23:515–548. doi: 10.1146/annurev.immunol.23.021704.115611. [DOI] [PubMed] [Google Scholar]
- 37.Zou W, Chen L. Inhibitory B7-family molecules in the tumour microenvironment. Nat Rev Immunol. 2008;8:467–477. doi: 10.1038/nri2326. [DOI] [PubMed] [Google Scholar]
- 38.Bottino C, Castriconi R, Pende D, Rivera P, Nanni M, Carnemolla B, Cantoni C, Grassi J, Marcenaro S, Reymond N, Vitale M, Moretta L, Lopez M, Moretta A. Identification of PVR (CD155) and Nectin-2 (CD112) as cell surface ligands for the human DNAM-1 (CD226) activating molecule. J Exp Med. 2003;198:557–567. doi: 10.1084/jem.20030788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gasser S, Orsulic S, Brown EJ, Raulet DH. The DNA damage pathway regulates innate immune system ligands of the NKG2D receptor. Nature. 2005;436:1186–1190. doi: 10.1038/nature03884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kloss M, Decker P, Baltz KM, Baessler T, Jung G, Rammensee HG, Steinle A, Krusch M, Salih HR. Interaction of monocytes with NK cells upon toll-like receptor-induced expression of the NKG2D ligand MICA. J Immunol. 2008;181:6711–6719. doi: 10.4049/jimmunol.181.10.6711. [DOI] [PubMed] [Google Scholar]
- 41.Nedvetzki S, Sowinski S, Eagle RA, Harris J, Vely F, Pende D, Trowsdale J, Vivier E, Gordon S, Davis DM. Reciprocal regulation of human natural killer cells and macrophages associated with distinct immune synapses. Blood. 2007;109:3776–3785. doi: 10.1182/blood-2006-10-052977. [DOI] [PubMed] [Google Scholar]
- 42.Bui JD, Carayannopoulos LN, Lanier LL, Yokoyama WM, Schreiber RD. IFN-dependent down-regulation of the NKG2D ligand H60 on tumors. J Immunol. 2006;176:905–913. doi: 10.4049/jimmunol.176.2.905. [DOI] [PubMed] [Google Scholar]
- 43.Jinushi M, Takehara T, Kanto T, Tatsumi T, Groh V, Spies T, Miyagi T, Suzuki T, Sasaki Y, Hayashi N. Critical role of MHC class I-related chain A and B expression on IFN-alpha-stimulated dendritic cells in NK cell activation: impairment in chronic hepatitis C virus infection. J Immunol. 2003;170:1249–1256. doi: 10.4049/jimmunol.170.3.1249. [DOI] [PubMed] [Google Scholar]
- 44.Stern-Ginossar N, Gur C, Biton M, Horwitz E, Elboim M, Stanietsky N, Mandelboim M, Mandelboim O. Human microRNAs regulate stress-induced immune responses mediated by the receptor NKG2D. Nat Immunol. 2008;9:1065–1073. doi: 10.1038/ni.1642. [DOI] [PubMed] [Google Scholar]
- 45.Gong AY, Zhou R, Hu G, Li X, Splinter PL, O’Hara SP, LaRusso NF, Soukup GA, Dong H, Chen XM. MicroRNA-513 regulates B7-H1 translation and is involved in IFN-gamma-induced B7-H1 expression in cholangiocytes. J Immunol. 2009;182:1325–1333. doi: 10.4049/jimmunol.0804370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Xu H, Cheung IY, Guo HF, Cheung NK. MicroRNA miR-29 modulates expression of immunoinhibitory molecule B7-H3: potential implications for immune based therapy of human solid tumors. Cancer Res. 2009;69:6275–6281. doi: 10.1158/0008-5472.CAN-08-4517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nice TJ, Coscoy L, Raulet DH. Posttranslational regulation of the NKG2D ligand Mult1 in response to cell stress. J Exp Med. 2009;206:287–298. doi: 10.1084/jem.20081335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Armeanu S, Krusch M, Baltz KM, Weiss TS, Smirnow I, Steinle A, Lauer UM, Bitzer M, Salih HR. Direct and natural killer cell-mediated antitumor effects of low-dose bortezomib in hepatocellular carcinoma. Clin Cancer Res. 2008;14:3520–3528. doi: 10.1158/1078-0432.CCR-07-4744. [DOI] [PubMed] [Google Scholar]
- 49.Vales-Gomez M, Chisholm SE, Cassady-Cain RL, Roda-Navarro P, Reyburn HT. Selective induction of expression of a ligand for the NKG2D receptor by proteasome inhibitors. Cancer Res. 2008;68:1546–1554. doi: 10.1158/0008-5472.CAN-07-2973. [DOI] [PubMed] [Google Scholar]
- 50.Salih HR, Rammensee HG, Steinle A. Cutting edge: down-regulation of MICA on human tumors by proteolytic shedding. J Immunol. 2002;169:4098–4102. doi: 10.4049/jimmunol.169.8.4098. [DOI] [PubMed] [Google Scholar]
- 51.Cao W, Xi X, Hao Z, Li W, Kong Y, Cui L, Ma C, Ba D, He W. RAET1E2, a soluble isoform of the UL16-binding protein RAET1E produced by tumor cells, inhibits NKG2D-mediated NK cytotoxicity. J Biol Chem. 2007;282:18922–18928. doi: 10.1074/jbc.M702504200. [DOI] [PubMed] [Google Scholar]