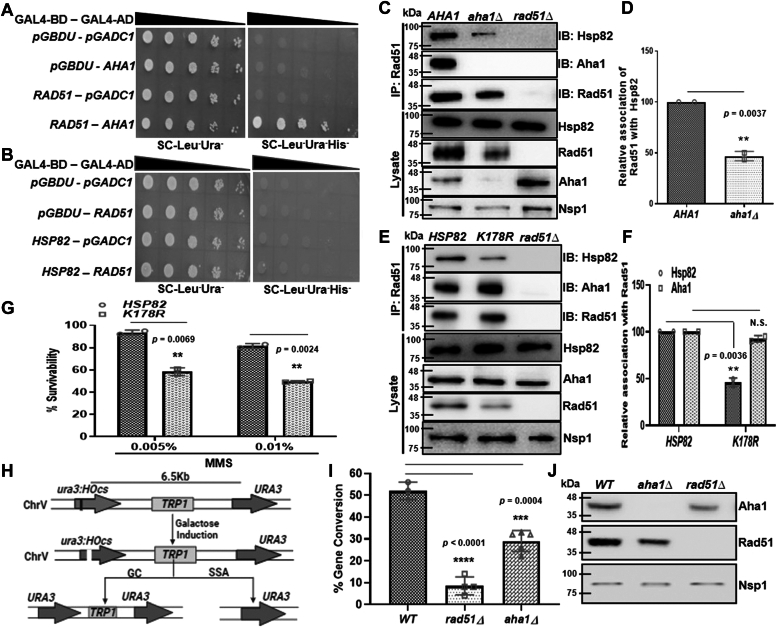
Figure 4.
Intracellular complex formation between Rad51 and Hsp82 is facilitated by Aha1.A, yeast two-hybrid assay depicting the interaction of Rad51 with Aha1. Strains harboring varied bait and prey vectors were grown till 0.5 absorbance and serially diluted. The diluted cells were spotted on SC-Leu⁻Ura⁻ (left panel), and SC-Leu⁻Ura⁻His⁻ to assess the interaction (right panel). B, yeast two-hybrid analysis showed no physical interaction of Rad51 with Hsp82. C, Western blots showing CoIP of Rad51 with Hsp82 and Aha1 in WT and aha1Δ strains. D, the above experiment was repeated two times and quantification of Western blots depicted a two-fold reduction in the interaction of Rad51 with Hsp82 in the absence of Aha1; the mean value (±SD) was plotted; p values were calculated using the two-tailed Student’s t test (∗∗p < 0.01). E, Western blots showing CoIP of Rad51 with Hsp82 and Aha1 in WT and K178Rhsp82 strains. F, the quantification of Western blot images showed significant reduced association between Rad51 and Hsp82 in K178Rhsp82 strain, while Rad51 interaction with Aha1 remained unchanged; the mean value (±SD) was plotted; p values were calculated using the two-tailed Student’s t test (∗∗p < 0.01). G, return to growth assay depicting the cell survivability of WT and K178Rhsp82 strain upon 0.005%- and 0.01%-MMS treatment. The above experiment was conducted in two independent sets, and the mean value (±SD) was plotted; p values were calculated using the two-tailed Student’s t test (∗∗p < 0.01). H, schematic presentation behind the principle of gene conversion assay in NA3 strain. The cell harbors two copies of URA3 in chromosome V, one of which has a HO endonuclease cut site incorporated within it. Upon induction of HO, the single DSB will be created and it can be repaired either by the Rad51 dependent gene conversion (GC), utilizing URA3 template that is 6 kb apart or by the Rad51 independent single strand annealing (SSA). In GC-mediated DNA repair, the intermediate TRP cassette is retained, however; the TRP cassette is deleted in SSA mediated DNA repair. I, percent gene conversion efficiency was plotted for the individual strain. The experiment was repeated more than three times and the mean value (±SD) was plotted; p values were calculated using the two-tailed Student’s t test (∗∗∗∗p < 0.0001, ∗∗∗p < 0.001). J, Western blot confirms the deletion of Aha1 in NA3 aha1Δ strain. Rad51 levels were compared between the NA3 and NA3 aha1Δ strain. CoIP, coimmunoprecipitationl; DSB, double-strand break; MMS, methyl methane sulfonate.