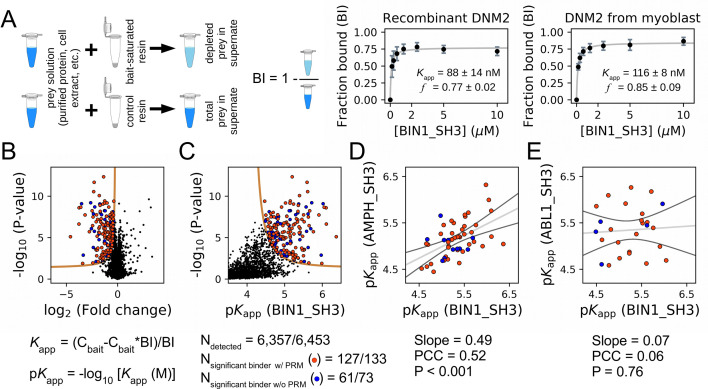
Figure 3. Affinity measurements between the SH3 domain of BIN1 and full-length proteins from cell extracts using nHU-MS.
(A) Outline of the holdup assay and benchmarking of nHU. Holdup is a simple tool to measure the fraction bound of prey molecules. This prey solution can be either a single purified protein, or a complex mixture of molecules and the prey depletion can be monitored with a multitude of analytical approches, such as western blot, or mass spectrometry. Titration holdup experiments were used to further characterize the interactions of BIN1_SH3 and full-length DNM2 using either recombinant, purified DNM2, or total myoblast extract containing endogenous DNM2 as a prey. The two experiments show nearly identical binding affinity and partial activity. (The average and standard deviation of three nHU western blot experiments are shown.) (B) Results of single point nHU-MS experiments carried out with the SH3 domain of BIN1 and total Jurkat extracts. Interaction partners above the significance threshold (tan line) are colored in orange if putative PRMs were identified in their sequence and blue if not. (C) Measured depletion values were converted to affinities using the functions indicated below panel B, assuming a simple binding mechanism and 10 μM estimated bait concentration. The number of unique affinity measurements performed and the identified BIN1 interaction partners found in a single experiment/in all measurements are indicated below panel C. (D, E) We also performed nHU-MS experiments with a set of closely or distantly related SH3 domains and compared their affinity profiles with BIN1. This way, we could quantify that related SH3-domains, for example the one found in AMPH, show similarities in their affinity interactomes, displaying statistically significant correlation between the measured affinity constants. In contrast, unrelated SH3 domains, such as the one found in ABL1, bind targets with dissimilar affinities. A linear fit (grey line) and a 95% confidence band (black line) is shown on all affinity comparisons. The statistical significance of correlation was determined by two-tailed, unpaired T-test. See Figure 3—figure supplements 1–3 and Supplementary file 1 for further details. Mass spectrometry experiments were performed with three injection replicates.