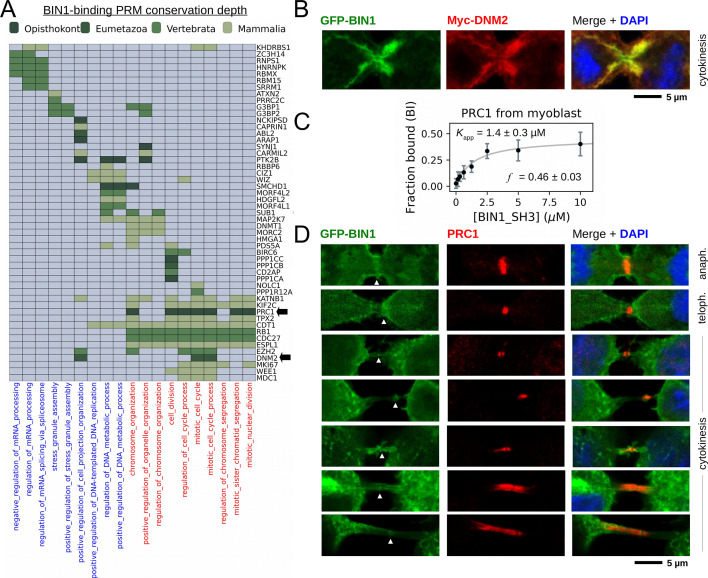
Figure 6. BIN1 interacts with multiple proteins involved in the mitotic phase and is localized at the membrane bridge formed between the daughter cells.
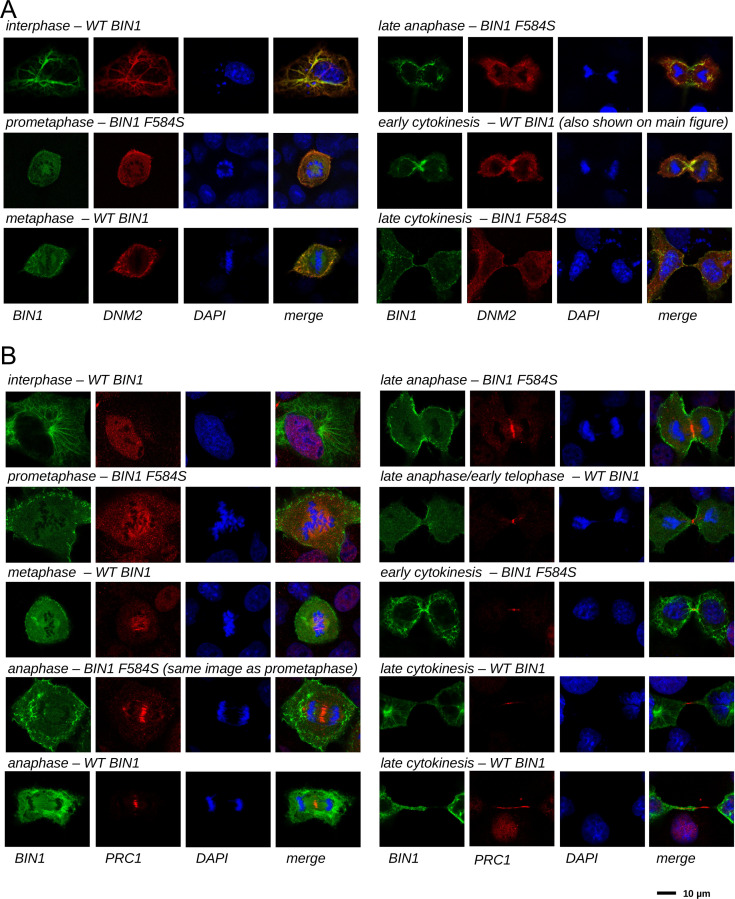
(A) Functional clustering of the identified BIN1 partners that contains BIN1-binding PxxP motifs. At the bottom of the panel, nuclear, or nucleic-acid-related processes are colored in blue and mitotic processes are colored in red. Heatmap color coding is according to the conservation depth of the highest affinity BIN1-interacting motif. See Supplementary file 3 for data. (B) During cytokinesis, BIN1 and DNM2 were found to localize at the cleavage furrow. A representative image of dividing Cos-1 cells, that were transfected with GFP-BIN1 and Myc-DNM2. (C) Titration nHU to further characterize the interactions of BIN1 with PRC1. PRC1 binding is measured from the same binding experiment using myoblast extract, that was used to characterize DNM2 binding (Figure 3A). (The average and standard deviation of three nHU western blot experiments are shown.) (D) Representative confocal images of the membrane bridge between daughter cells. Cos-1 cells were transfected with GFP-BIN1 and stained for endogenous PRC1. White arrows indicate the cleavage furrow or the midbody. For additional supporting confocal images see Figure 6—figure supplement 1.