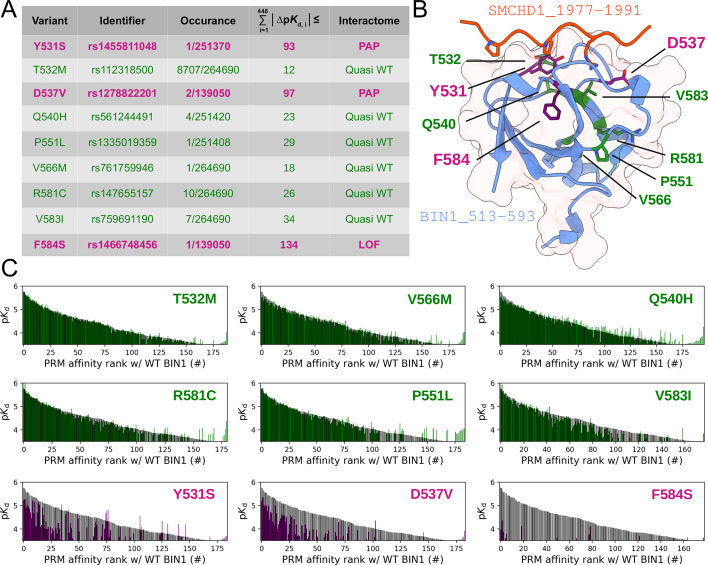
Figure 7. Natural variants of BIN1 can cause affinity perturbation at an interactomic scale.
(A) A summary of our affinity interactomic tests performed with 9 natural variants of the BIN1 SH3 domain. The cumulative Euclidean affinity distances to the WT BIN1, calculated based on the explored 448 dimensional affinity space, are indicated for each variant. For affinities where no detectable binding was observed the detection threshold was used for calculation, hence only the lower limit of the Euclidean distance could be estimated. Variants colored green have minor effect on affinity interactomes, while variants colored in purple displaying either perturbed affinity profiles (PAP) or general loss of functions (LOF). (B) Location of the assayed variants on the structure of WT BIN1. D537 and Y531 are positioned near the PRM binding interface, F584 is buried in the hydrophobic core of the SH3 domain. The structure of the BIN1 SH3 domain bound to a high affinity peptide taken from SMCHD1 was generated using AlphaFold2 (Tunyasuvunakool et al., 2021). (C) Affinity interactome profiles of the BIN1 variants (colored in green or purple) compared with the affinity profile of WT BIN1 (colored in black). Motifs in the affinity profiles were ranked according to their affinities measured with WT BIN1. Only the motifs displaying detectable binding out of the 449 assayed PRMs are shown. See Supplementary file 2 for further details.