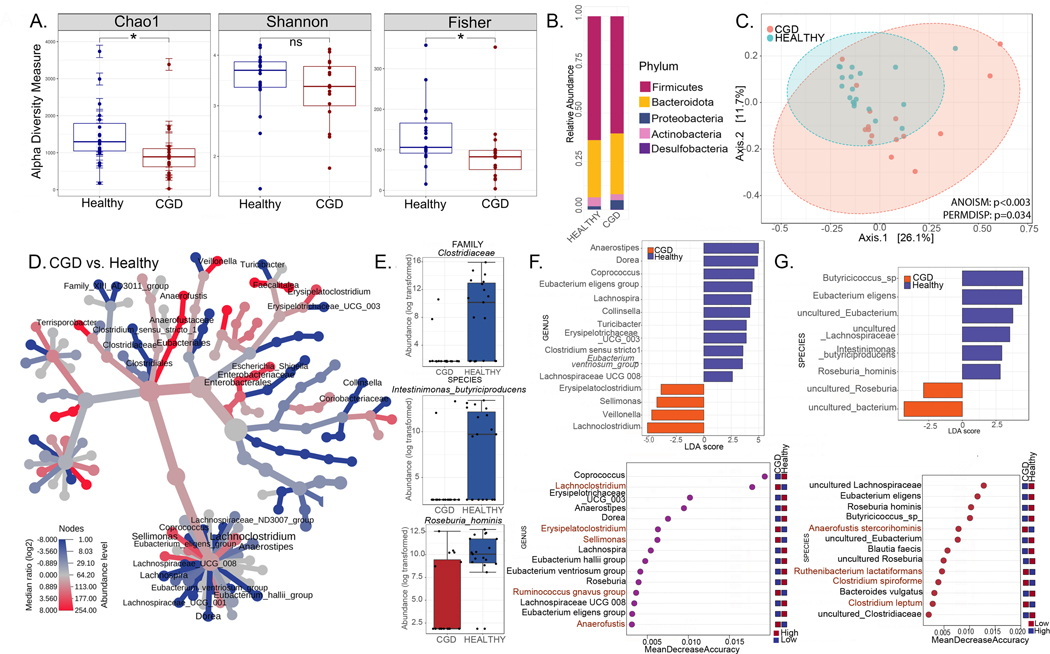
Figure 2. Intestinal microbiome signatures distinguish patients with CGD from healthy individuals.
(A-G) Comparisons for the CGD group (without a history of IBD and only on prophylactic antimicrobials, n=16) with the Healthy group (n=17). (A) Alpha diversity (within group variations) analyses (Chao1, Shannon, Fisher; * = p<0.05, **=p<0.01, ***=p<0.001, ****=p<0.0001, ns = non-significant) for CGD compared to Healthy. (B) Relative abundance of amplicon sequence variants in both groups presented at phylum level. (C) PCoA plot of beta diversity (between-group variations) based on weighted Unifrac distances for the two comparison groups with p values determined by Analysis of similarities (ANOSIM; p<0.003) and PERMDISP test for significant differences in dispersion (p =0.034). (D) Heat tree depicting the phylogenetic relationship and significant differential abundance (p<0.05) of bacterial genera between CGD and Healthy groups (red = higher abundance; blue = lower abundance). (E) Top differentially abundant family and species as identified by edgeR analysis (p values for all comparisons are <0.0001). (F) Linear discriminant analysis (LDA) score determined by the LEfSe analysis showing biomarkers at genus and species levels that are present or absent in the experimental groups shown. (G) Random forest analysis where genera and species with highest discriminatory power between CGD and Healthy groups are listed (red = high abundance in the experimental group, blue = low abundance in the experimental group).