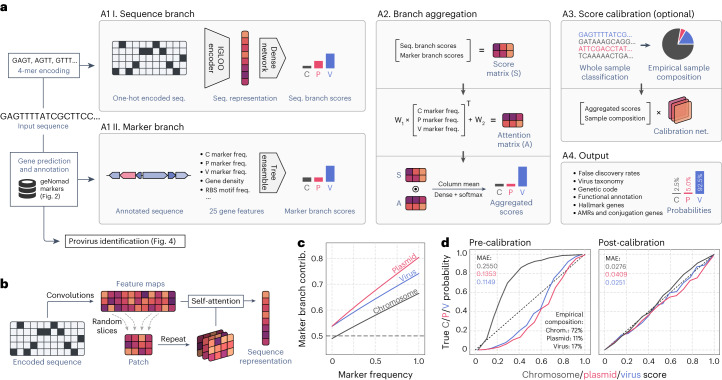
Fig. 1. A hybrid framework for identifying and annotating plasmids and viruses.
a, geNomad processes user-provided nucleotide sequences through two branches. In the sequence branch, the inputs are one-hot encoded fed to an IGLOO neural network, which scores inputs based on the detection of non-local sequence motifs (A1 I). In the marker branch, proteins encoded by the input sequences are annotated using markers that are specific to chromosomes, plasmids or viruses (A1 II). A set of numerical features is then extracted from the annotated proteins and fed to a tree ensemble model, which scores the inputs based on their marker content. Next, the scores provided by both branches are aggregated by weighing the contribution of each branch based on the frequency of markers in the sequence (A2). Aggregated scores can then be calibrated to approximate probabilities in a process that leverages the sample composition inferred from the classification of sequences from the same batch (A3). Lastly, classification results are summarized and presented together with additional data, such as virus taxonomy, gene function and the inferred genetic code (A4). b, The sequence branch is based on the IGLOO architecture, which uses convolutions to produce a feature map from a one-hot encoded input. Patches encoding non-local relationships within the sequence are then generated by slicing the feature map. Lastly, these patches are used as an attention matrix to produce a sequence representation from the feature map. c, The relative contribution of the marker branch (y axis, quantified using SHAP) increases as the marker frequency (fraction of genes assigned to a marker) in the sequence increases. d, Calibration curves of pre-calibration (left) and post-calibration (right) scores, showing that sample composition can be used to map classification scores to actual probabilities. The x axis represents scores averaged across multiple bins; the y axis represents the fraction of positives in each bin; the 45° dashed line represents a perfect calibration scenario. freq., frequency; MAE, mean absolute error of the scores relative to the true probabilities.