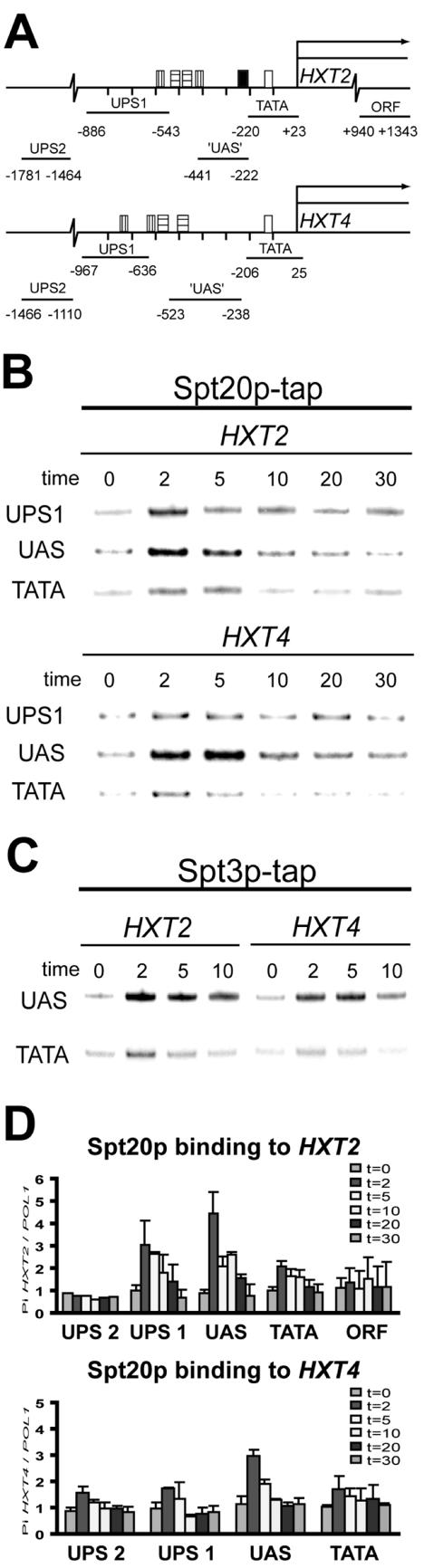
FIG. 2.
SAGA specifically binds to the upstream region of both HXT2 and HXT4 genes. (A) HXT promoter structure and location of primers used to analyze Spt20p and Spt3 binding. Boxes indicate bind-ing sites for Mig1p (vertical stripes), Rgt1p (horizontal stripes), putative UAS (filled), and TBP (open) (39). Lines represent the DNA fragment, which was amplified in a multiplex PCR analysis. Numbers represent the exact location of HXT2 and HXT4 primers relative to the start site of the coding region. (B and C) Cells expressing a TAP-tagged version of Spt20p (B) or Spt3p (C) were subjected to a glucose shift, and cross-linking was initiated by addition of formaldehyde at the indicated time points. Input and immunoprecipitated DNA was analyzed by multiplex PCR with primers spanning the TATA box (TATA), putative UAS, or further upstream (UPS1) regions of the HXT genes as indicated. (D) Quantification of ChIP signals over the HXT2 and HXT4 genes. Radioactive signals of the indicated PCR fragments were quantified with a PhosphorImager. The signals are expressed relative to the POL1 signal, which was included as a negative control in the multiplex PCR analysis.