Extended Data Fig. 1 |. Frequency of Major Breast Cell Types Across Women and Sample Types.
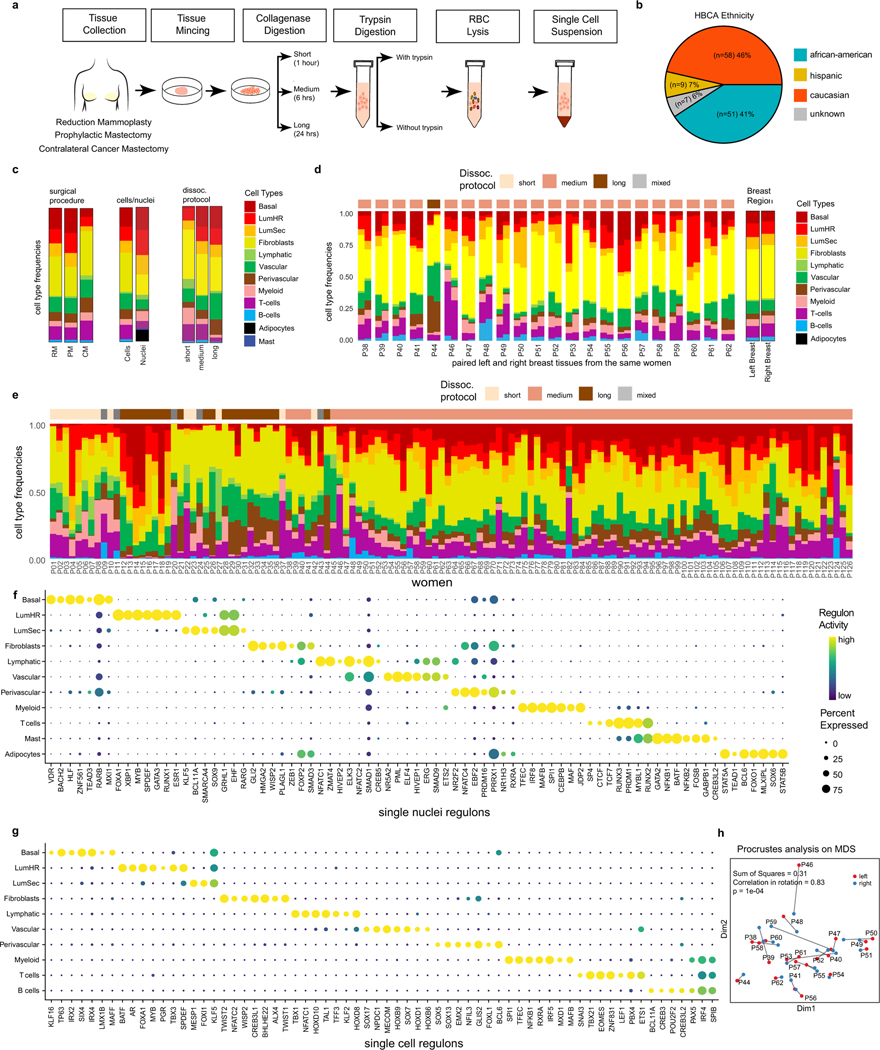
a, Experimental workflow for breast tissue processing for scRNA-seq, showing different conditions used for digestion times and trypsin treatments. b, Pie chart showing ethnic backgrounds of women who provided tissue samples for the breast atlas. c, Cell type frequencies for different tissue sources (reduction mammoplasties - RM, prophylactic mastectomies - PM and contralateral mastectomies - CM), cells vs nuclei single cell RNA-seq protocols, and experimental dissociation protocol. d, Major cell type frequencies of matched left and right breasts from 22 women (left) and averages across all left and all right breast tissues (right). e, Stacked barplot showing the variation of cell type frequencies across the 126 women in scRNA-seq data. The top annotation bar shows the experimental workflow used (short/medium/long). f, Top regulons identified with SCENIC for each cell type cluster from the snRNA-seq data. g, Top regulons identified with SCENIC for each cell type cluster from the scRNA-seq data. h, Multi-dimensional scaling and Procrustes analysis to determine the concordance of left and right breast cell type frequencies and Pearson correlations for 22 women with matched breast tissue samples. P-value was calculated based on a two-sided test.