Figure 2.
Electrophysiological characterization of SYNGAP1-deficient cortical neurons
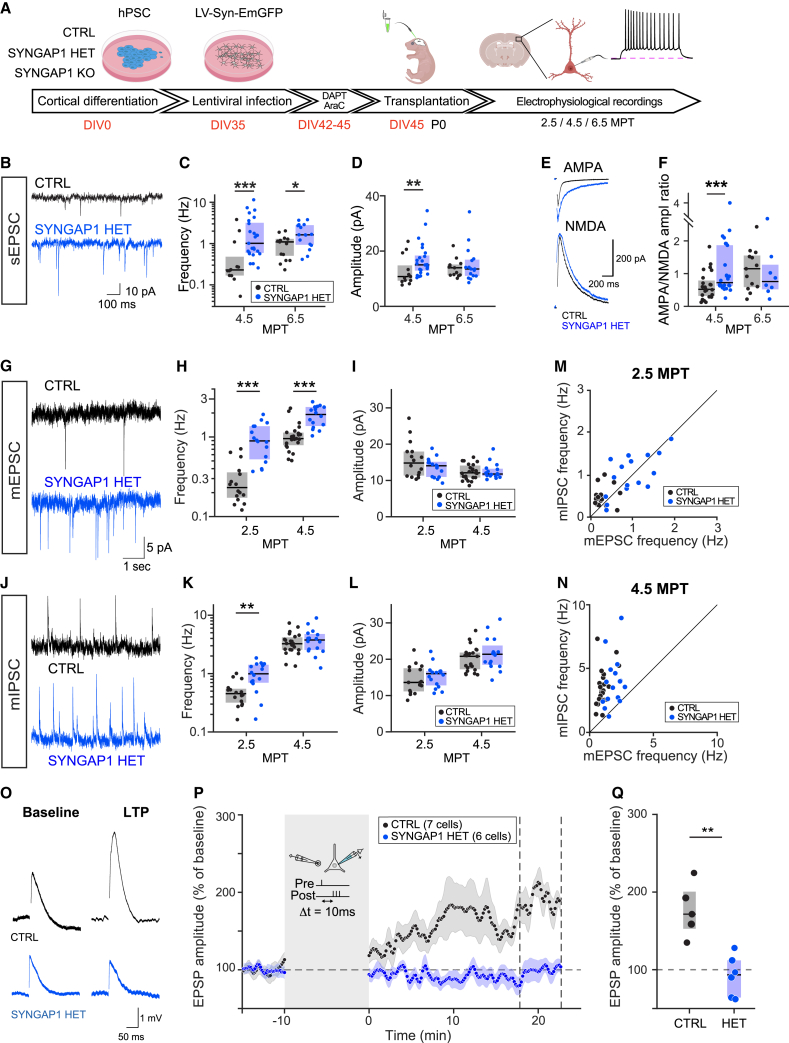
(A) Experimental timeline: differentiated mutant or control cells are infected with LV-EmGFP and transplanted. Coronal slices are prepared and EmGFP-labeled cells are targeted for patch-clamp experiments.
(B) Recording traces of SYNGAP1 CTRL and HET neurons showing example sEPSCs. Note more and larger inflections in the blue (HET) trace.
(C and D) Quantification of synaptic properties across time: (C) sEPSC frequency at 4.5 and 6.5 MPT. Medians per genotype/time point (4.5 MPT: 0.23, 1.02; 6.5 MPT: 1.10, 1.63). Data collected at 4.5 MPT from CTRL: 13 cells (4 mice); HET: 25 (4). Data collected at 6.5 MPT from CTRL: 13 cells (2 mice); HET: 16 (3). (D) sEPSC amplitude at 4.5 and 6.5 MPT. Medians per genotype/time point (4.5 MPT: 10.8, 15.00; 6.5 MPT: 14.0, 13.5). Data collected from same cells as in (D).
(E) Example AMPA (top) and NMDA (bottom) traces for both CTRL (black) and HET (blue) genotypes. AMPA currents are increased in HET (blue) neurons.
(F) AMPA/NMDA ratio at 4.5 and 6.5 MPT. Medians per genotype (4.5 MPT: 0.52, 0.72; 6.5 MPT: 1.15, 0.76). Data collected at 4.5 MPT from CTRL: 24 cells (4 mice); HET: 26 (5). Data collected at 6.5 MPT from CTRL: 13 cells (2 mice); HET: 8 (1). Boxes indicate median and IQR. Statistical comparison was done using rank-sum tests.
(G–I) Quantification of mEPSC recordings at 2.5 and 4.5 MPT. (G) Example mEPSC traces for both genotypes; TTX is applied to isolate spontaneous synaptic events. (H) mEPSC frequency at 2.5 and 4.5 MPT. Medians per genotype/time point (2.5 MPT 0.23, 0.89; 4.5 MPT: 0.95, 1.92). Data collected at 2.5 MPT from CTRL: 16 cells (2 mice); HET: 16 (3). Data collected at 4.5 MPT from CTRL: 23 cells (3 mice); HET: 16 (3). (I) mEPSC amplitude at 2.5 and 4.5 MPT. Medians per genotype/time point (2.5 MPT: 14.8, 14.00; 4.5 MPT: 12.1, 11.8).
(J–L) Quantification of miniature inhibitory post-synaptic currents (mIPSC) recordings at 2.5 and 4.5 MPT. (J) Example mIPSC traces for both genotypes, as in (G). (K) mIPSC frequency at 2.5 and 4.5 MPT. Medians per genotype/time point (2.5 MPT: 0.45, 0.99; 4.5 MPT: 3.24, 3.76). Data collected from same cells as (H). (L) mIPSC amplitude at 2.5 and 4.5 MPT. Medians per genotype/time point (2.5 MPT: 13.6, 16.00; 4.5 MPT: 20.8, 21.4). Data collected from same cells as (H).
(M and N) E/I frequency at 2.5 and 4.5 MPT. (M) Scatterplot showing miniature excitatory post-synaptic currents (mEPSC) and mIPSC frequency per cell at 2.5 MPT and (N) at 4.5 MPT. In both cases, HET (blue) neurons tend toward higher E/I ratios.
(O–Q) LTP experiments performed at 7–8 MPT. (O) Example traces during baseline (left) and after potentiation (right). (P) Average excitatory post-synaptic potential (EPSP) per genotype showing deviation from baseline after 10′ pairing protocol (see STAR Methods). CTRL neurons (black) show potentiation, whereas HET neurons (blue) show lack of potentiation. (Q) Quantification of average EPSP levels 20 min after the end of the potentiation protocol. Data collected between 7.5 and 8.5 MPT from CTRL: 5 cells (3 mice); HET: 6 (3).
See also Figures S3 and S4.