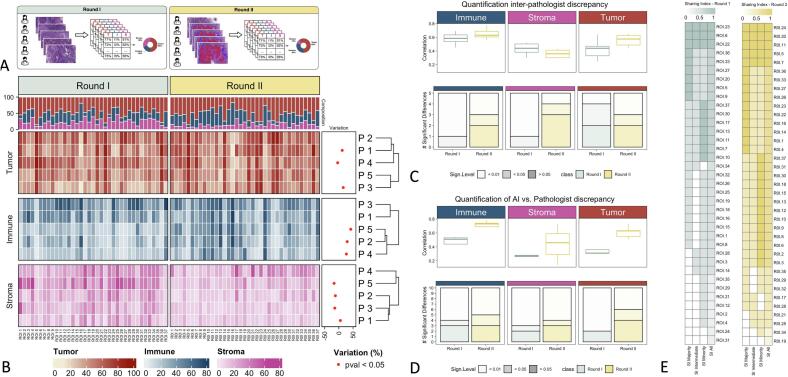
Fig. 2.
(A) Illustration of the pathologist's estimation rounds. Round I and Round II have been performed on the same ROIs with and without the visual support of AI-based cell classification. (B) Pathologist's estimations heatmap. The heatmap shows the percentage of the three cell types assigned by each of the pathologists (P1–P5) for all ROIs. For each ROI, the top annotation describes the average cell composition. Statistically significant percentual variation (across ROIs) in estimates provided by pathologists in Round I and Round II is described and shown in the right annotation. (C) Quantification of inter-pathologist discrepancy in Round I vs Round II. The boxplots represent Kendall correlation coefficients, calculated on the estimates given by all couples of pathologists (across ROIs). The bar plots represent the number of significant Wilcoxon tests performed on the estimates given by all couples of pathologists (across ROIs), at two different p-value thresholds (0.05 and 0.01). (D) Quantification of AI vs. Pathologist discrepancy. The boxplots represent Kendall correlation coefficients, calculated on the estimates given by the AI vs. each of the pathologists (across ROIs). The bar plots represent the number of significant Wilcoxon tests performed on the estimates given by the AI vs. each of the pathologists (across ROIs), at two different p-value thresholds (0.05 and 0.01). (E) Sharing Index (SI) heatmap in Round I and Round II for all three hierarchical positions as well as majority, minority, and intermediate hierarchical position individually. ROIs on the y-axis are sorted from the highest to the lowest global SI. A SI of 1 indicates complete agreement, whereas a SI of 0 indicates the maximum level of discrepancy across pathologists.