Abstract
Natural killer (NK) cells are major innate lymphocytes. NK cells do not require prior antigen exposure to mediate antitumor cytotoxicity or proinflammatory cytokine production. Since they use only nonclonotypic receptors, they possess high clinical value in treatment against a broad spectrum of malignancies. Irrespective of this potential, however, the transcriptional regulation that governs human NK cell development remains far from fully defined. Various environmental cues initiate a complex network of transcription factors (TFs) during their early development, one of which is GATA2, a master regulator that drives the commitment of common lymphoid progenitors (CLPs) into immature NK progenitors (NKPs). GATA2 forms a core heptad complex with six other TFs (TAL1, FLI1, RUNX1, LYL1, LMO2, and ERG) to mediate its transcriptional regulation in various cell types. Patients with GATA2 haploinsufficiency specifically lose CD56bright NK cells, with or without a reduced number of CD56dim NK cells. Here, we review the recent progress in understanding GATA2 and its role in human NK cell development and functions.
Keywords: GATA2, NK cell, NK cell development
I. INTRODUCTION
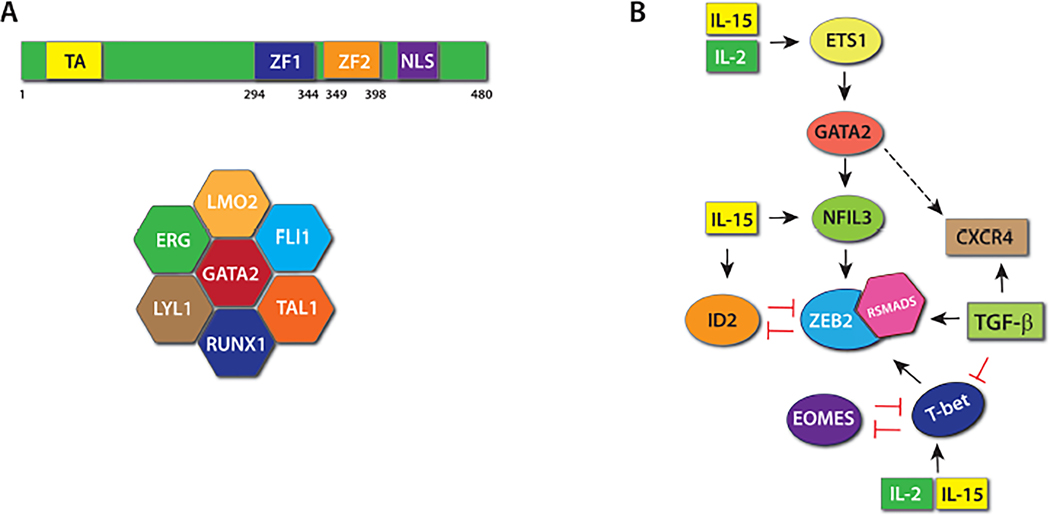
GATA2 is an essential transcription factor (TF) of the generation, survival, proliferation, and differentiation of hematopoietic stem cells (HSCs),1–3 as well as of the formation of blood and lymphatic vessels.4,5 This pioneer factor comprises two zinc finger (ZF) domains, two transactivation domains, one nuclear localization signal, and one negative regulatory domain6,7 (Fig. 1A). The two ZF domains (N-terminal and C-terminal) are highly conserved, interacting with a network of TFs, including SPI1 (PU.1), RUNX1, TAL1, FLI1, and LMO2.8,9 These TFs are known to be involved in specifying early hematopoietic lineage commitment.9–12 In human HSCs, GATA2 forms a core heptad complex with six other TFs (TAL1, FLI1, RUNX1, LYL1, LMO2, and ERG) to directly impact the survival and differentiation of HSCs by regulating more than 1,000 target genes9,13 (Fig. 1A).
FIG. 1:
GATA2, its transcriptional partners, and their transcriptional role in human NK cells. (A) GATA2 comprises two zinc finger (ZF) domains, two transactivation domains, one nuclear localization signal, and one negative regulatory domain (Top). GATA2 forms a core heptad complex with six other TFs (TAL1, FLI1, RUNX1, LYL1, LMO2, and ERG) to directly impact the survival differentiation of HSCs by regulating more than 1,000 target genes (Bottom). (B) Multiple potential upstream and downstream factors of GATA2 and cytokines. This predicted pathway was assembled using findings from NK and other cell types.
GATA2 deficiency can result in monoMAC syndrome (monocytopenia with atypical mycobacterial infection),14 DCML [dendritic cell (DC), monocyte, B cell, and NK lymphocyte deficiency],15,16 MDS/AML (familial myelodysplastic syndrome/acute myeloid leukemia),17 and Emberger syndrome (consisting of MDS, lymphedema, and warts from human papillomavirus infection).4 Although GATA2 deficiency results in various clinical manifestations, the major symptoms in most patients are chronic infections, including HPV, EBV, mycobacterium, fungal and other bacterial infections, MDS/AML, cytopenia (B cell, NK cell, monocyte, DC, and CD4+ T cell deficiency), and lymphedema.18 Available patient data suggest that GATA2 is a master regulator of human NK cell development whose haploinsufficiency leads to NK cell dysfunction.19
In this review, we summarize the recent progress related to the overall molecular mechanisms employed by GATA2 and its role in human NK cell development and functions.
II. HUMAN NK CELL DEVELOPMENT
NK cells are the major subset of innate lymphocytes, which mediate both proinflammatory cytokine production and cytotoxicity in response to viral infections or malignant transformation.20,21 The release of lytic granules containing perforin and granzymes results in the lysis of target cells.22,23 Antibody-dependent cell–mediated cytotoxicity (ADCC) is another essential mechanism in recognizing infected or transformed target cells.24,25 Since NK cell–mediated functions are not limited by clonotypic receptors,26,27 cellular therapies can be used against a broad spectrum of cancers.28,29
In spite of their clinical potential, NK cells have not been utilized to their fullest in the clinic. For this reason, understanding the transcriptional regulation of the development and functions of human NK cells is paramount in formulating effective cellular therapies. Customized in vitro generation of mature functional human NK cells from inducible pluripotent stem cells (iPSCs) provides an exceptional opportunity for individualized front-line cancer therapy. However, lacking essential knowledge of master transcriptional regulators we are limited in our ability to fully realize the clinical potential of human NK cells. Thus, defining the role of crucial TFs, such as GATA2, in the early commitment and development of human NK cells is of high clinical relevance.
Recent studies show that human NK cells develop and mature in bone marrow and secondary lymphoid tissues (SLTs), such as lymph nodes (LNs), spleen, and tonsils.30 Traditionally, the differential expression of surface markers defines distinct developmental stages of human NK cells. We recently summarized these markers.31,32 Human NK cells primarily arise from self-renewing pluripotent HSCs in bone marrow.33,34 Lineage-negative CD34+ HSCs give rise to lymphoid-primed multipotential progenitors (LMPPs) and CD34+CD244+ common lymphoid progenitors (CLPs) in sequential order. CLPs commit to the NK cell lineage, becoming NKPs (characterized by CD117+CD127+CD122+IL-1R1+).
Immature NK cells (iNKs) express a higher level of CD56 (CD56bright) and IL-1R1 along with CD161 (NK1.1), CD314 (NKG2D), CD335 (NKp46), and CD337 (NKp30).35 The maximal expression of NK1.1, NKG2D, NKp46, and NKp30 marks the commitment of iNKs to the transitional stage (TransNKs), which is separated into two substages by the expression of NKp80 (relatively immature substage a: NKG2D+CD337+CD161+NKG2A+CD56bright NKp80–; and relatively mature substage b: NKG2D+CD337+CD161+NKG2A+CD56bright NKp80+). TransNKs develop into the mature stage, becoming mNKs, with decreased CD56, increased CD16, and distinct expressions of CD158 (KIR) subtypes. The mNKs are NKp80+CD56dimCD16+KIR−/+. Expression of CD57 and increased KIR expression indicate terminally mature NK cells (TermNKs).35,36
Multiple cytokines in the bone marrow are responsible for the commitment, development, and maturation of NK cells (Fig. 2). Among these, stem cell factor (SCF), FMS-like tyrosine kinase 3 ligand (Flt3L), c-Kit, and IL-7 promote the early commitment of HSCs to CLPs.37–39 IL-15 is essential for early NK cell development and survival,40,41 and IL-2 is critical for survival, activation, and expansion.42–44 Synergizing with IL-15 and IL-2, IL-21 enhances NK cell cytotoxicity.45 IL-12 and IL-18 augment IFN-γ production and promote the cytotoxicity of NK cells.46 TGF-β sustains the stemness of CD34+ HSCs by blocking their commitment to the NK cell lineage.47 However, at a later stage, it functions as a checkpoint to maintain NK cell immaturity.48 While these and other soluble mediators play obligatory roles in the development of NK cells, the activation and functions of downstream transcriptional regulators have been only partially defined.
FIG. 2:
Role of major transcription factors, including GATA2, in human NK cell development. Transcription factors and the specific developmental stages of human NK cells are indicated. GATA2 plays an essential role in the transition of NKPs into immature NKs. The role of GATA2 in the early commitment of CLPs into NKPs is yet to be established.
III. TRANSCRIPTIONAL CONTROL OF HUMAN NK CELL DEVELOPMENT
Distinct TFs drive NK cells to transition from one stage to another in sequential order.31 The upstream and downstream regulators of GATA2 during NK cell development need to be fully determined, but, limited information is available about the select few TFs that link it with human NK cell development (Fig. 2).
Notch proteins drive HSCs into CLPs; ID2 is a member of the inhibitor of DNA-binding proteins; and RUNX3 drives CLPs into the NK cell lineage. GATA2, NFIL3, and ETS1 promote NKP transition into immature NK cells.1,16,49–51 These TFs either positively or negatively regulate transition to maintain balanced development. Cytokine signaling pathways also play critical roles, either upstream or downstream of TFs, during human NK cell development. The constitutive expression of ID2 enhances commitment to the NK cell lineage from CD34+ CLPs. High ID2 expression synergizes with IL-15 and results in an increased NKPs.52 RUNX3 expression starts at the NKP stage, reaching its highest level in iNKs and mNKs,53 and is essential for the commitment of NKPs.53 RUNX3 binds the promoters of KIR and NKp46 and initiates their transcription,54 which demonstrates the role of this TF in the terminal maturation of human NK cells.
ETS1 is predominantly a lymphocyte-specific TF.55,56 It is first expressed in the progenitor stage of NK cells development and reaches its peak level in the NKP late stage.57 The absolute number of NK cells is significantly reduced among ETS1-deficient human cord blood cells, although the number of NKPs is unaltered.50 The remaining NK cells from ETS1 loss-of-function HSCs have reduced cytotoxicity and IFN-γ production.50 ETS1 contributes to NK cell development and function by regulating the expression of several critical TFs, including GATA3, NFIL3, T-bet, BLIMP1, and HOBIT.50 Earlier studies reported that ETS1 binds to the promoter region of GATA2 and initiates its transcription.58
Evidence from murine models identifies distinct TFs upstream of Ets1. Notch upregulates the transcription of Ets1,59 while PU.1 directly inhibits it.60,61 Ets1, in turn, promotes transcription of IDB2 (the Id2 gene) and TBX21 (the T-bet gene).57 The activation of human NK cells with IL-2 and IL-15 upregulates the expression of ETS1.62 NK and NKT cells, but not T or B cells, highly express NFIL3 (E4BP4).63 NFIL3 plays an essential role in NK cell commitment from NKPs to iNKs.63 It is involved in NK cell development as a downstream target of the IL-15–mediated signaling pathway during the commitment to iNKs.64 Also, it directly binds to the promoter region of IBD2 and EOMES and activates their transcription.65 GATA2 binds to the promoter region of NFIL3,66 which may promote iNK commitment and maintain cell survival.67
Eomesodermin (EOMES) and T-bet belong to the T-box family and play obligatory roles during NK cell development.68–70 Without it, NK cell development is blocked at an immature stage.71 However, the lack of T-bet blocks EOMES before the terminally mature stage.72 EOMES and T-bet antagonize the each other’s expression during NK cell maturation.68,69,71 IL-12 and IL-15 stimulation upregulates the expression of T-bet. Both T-bet and Eomes are the downstream targets of IL-15R–mediated signaling.73
ZEB2 (zinc finger E-box-binding protein) plays a critical role in the TGF-β–mediated signaling involved in epithelial-to-mesenchymal transition through the activation of R-Smads.74,75 Zeb2 regulates the terminal maturation of NK cells, indicating a potential role for TGF-β at the later stages of NK cell development.76 T-bet is required to induce the expression of Zeb2, which is critical to maintaining the transcriptional activity of T-bet, implicating a mutual regulation between these two TFs.76 Overexpression of Zeb2 partially restores NK cell defects that result from T-bet deficiency.76 Zeb2 is downstream of NFIL3 during dendritic cell specification, implying that IL-15R–mediated activation leads to initiation of the NFIL3-Zeb2 axis during early NK cell development.77 Mutual transcriptional repression between Id2 and Zeb2 has been found during DC lineage development which may also be operative in NKPs.77 Thus, the transcriptional control of early NK cell development by the NFIL3-Zeb2-Id2 axis needs to be investigated.
IV. IMPACTS OF GATA2 DEFICIENCY ON IMMUNE CELLS
Patients with GATA2 haploinsufficiency possess a reduced number of CD34+ cells in bone marrow (BM) but maintain maturation capability of hematopoietic stem cells.78 However, induced pluripotent stem cells (iPSCs) derived from GATA2-deficient patients do not replicate the defects in committing to hematopoietic progenitors.79 Patients with GATA2 deficiency also show near complete loss of B cell precursors78 and reduced transitional and naïve B cells, while memory B cells are enriched and appeared skewed to the mature state.80
Specific loss of CD56bright NK cells is a significant and consistent feature of GATA2 deficiency.19,80,81 T cell deficiency varies in patients. One cohort study reported that 50% of 57 patients with the GATA2 mutation displayed reduced CD4+ T cell numbers.18 Another result from a cohort study, including 30 patients with GATA2 mutation, showed reduced naïve and central memory CD8+ T cells revealed by decreased CD27, CD62L, CD38, and HLA-DR.80
Interestingly, several groups have reported an increased CD3+CD56+ T cell population in GATA2-deficient patients.19,80,82 These NK-like T cells play an essential role in eliminating cytomegalovirus (CMV) infections.83 However, because the role of GATA2 in their development and functions has not been determined, and while the type of NK or B cell deficiency is well-correlated with GATA2 mutation, the developmental defects associated with T cell subtypes have yet to be fully characterized.
DC deficiency is highlighted in patients with GATA2 deficiency and may relate to elevated FLT3L in patients with GATA2 haploinsufficiency. Indeed, studies have shown the number of DCs to be inversely correlated with the level of FLT3L.37,84 The high elevation of FLT3L is a unique serological feature of patients with GATA2 mutation, and its progressive elevation relates to the clinical advancement of associated disorders.15,80
Both CD14+ and CD16+ monocytes are significantly reduced in patients with GATA2 haploinsufficiency.80 In addition to dysfunctional immune cells, around 75% of patients who carry the GATA2 mutation develop a malignant blood disease, including MDS, AML, and chronic myelomonocytic leukemia (CMML).85 The acquisition of additional genetic abnormalities in GATA2 deficiency usually results in the rapid onset of hematological abnormalities with a poor prognosis for survival.86,87 These abnormalities include monosomy 7, trisomy 8, trisomy 21, and mutations in ASXL1, CEBPA, and other genes, alone or concomitant.86,88,89
To date, over 350 GATA2 genomic variants have been reported on the ClinVar website (https://www.ncbi.nlm.nih.gov/clinvar/; January 2021). Around 85% of them are single nucleotide substitutions. There are three major categories of mutations present in GATA2-deficient patients: (1) the N-terminal ZF domain (amino acid 259–319), (2) the C-terminal ZF domain (amino acid 349–373), and (3) other regions (such as +9.5 intronic enhancer).90 Somatic mutations predominately occur in the N-terminal ZF domain. GATA2sLeu321Phe is the most prevalent of these.91
All germline mutations are observed in the C-terminal ZF domain. The most common missense mutations, GATA2gThr354Met, GATA2gArg396Gln, and GATA2gArg398Trp, result in loss of function.17,92 Uniquely, GATA2Leu359Val is a gain-of-function mutation.93,94 The C-terminal ZF domain of GATA2 interacts with PU.1,95 which directly binds and drives the expression of FLT3 and granulocyte-macrophage colony–stimulating factor (GM-CSF)—essential growth factors in the development of DCs.96 Mutated GATA2Thr354Met and GATA2Cys373Arg physically bind to PU.1 with higher affinity due to the altered C-terminal ZF structure.97 Interaction between GATA2 and PU.1 results in reciprocal functional antagonism through regulation of their transcription and DNA binding.98,99
GATA1, another GATA family member, is a crucial TF that drives the differentiation of HSCs into megakaryocytes.100 While GATA2 is essential for maintaining stemness, it is also critical to early activation of GATA1 erythroid/megakaryocyte lineage commitment from HSCs. This phenomenon, known as the “GATA switch,” facilitates the displacement of GATA2 from the chromatin by GATA1.101–103 GATA2 can bind its own upstream promoter region and upregulate its transcription. Increased GATA2 activates GATA1, which in turn represses GATA2 expression. Both GATA1 and GATA2 are capable of autoactivating their own expression.104 New findings have challenged this paradigm, where high DNA methylation by DNMT1, not GATA2, is responsible for an inactive GATA1 locus.102,105 Detailed study is required to determine the role of the GATA switch in the commitment of HSCs to the NK lineage.
One of the critical TFs in lymphocyte development, Ets1, can bind to the GATA2 promoter and positively regulate its expression on erythroid differentiation.58 GATA2 protein directly binds to the NFIL3 promoter region, a TF that is essential in developing T, B, NK, and dendritic cells.106,107
GATA2−/− mice are embryonically lethal and die at day 10.5 (E10.5) due to loss of vascular integrity and anemia.1 Following tamoxifen-induced deletion, GATA2fl/flERCre adult mice display depleted splenic B cells, T cells, NK cells, monocytes, and DCs, along with impaired DC differentiation.108 Notably, CD-49b+NK1.1+ NK cell numbers are significantly reduced in these mice.108 However, the extent of the reduction in defective NK cell development and the impaired signaling pathways is not known. Mice with GATA2 haploinsufficiency display a reduced number of HSCs and granulocyte-macrophage progenitor (GMP) cells as well as an impaired ability to differentiate to monocytes109; however, they do not exhibit defects in DC numbers or DC differentiation,108 the development of MDS, or leukemia.109
Further studies are required to determine if NK cells display impaired development in GATA2+/− mice. Overexpression of GATA2 inhibits hematopoiesis through defective cell cycle pathways.110 Elevated GATA2 correlates with an adverse prognosis for patients with AML.111,112 Thus, a fine-tuned balance of functional GATA2 protein is critical to the self-renewal and differentiation of HSCs into lineage-committed progenitors. Although GATA2fl/fl ERCre and GATA2+/− mice can mimic some GATA2-deficiency features, a better disease model is necessary to define the molecular mechanisms of GATA2-mediated transcriptional regulations.
V. ROLE OF GATA2 IN HUMAN NK CELL DEVELOPMENT
GATA2 is an essential TF in lineage commitment and early NK cell development in humans.16 The specific loss of the CD56bright NK cell population is a striking feature of GATA2-deficient patients19 with or without reduced total CD56dim NK cells.19,81,82 Earlier, we reported that upregulated apoptosis is potentially the mechanism behind reduced NK cell numbers, supported by the augmented expression of the proapoptotic genes GIMAP4 and GIMAP7.35 In an in vitro culture, purified CD34+ hematopoietic precursors from GATA2-deficient patients failed to differentiate into CD56bright NK cells but gave rise to CD56dim NK cells, albeit in significantly lower numbers than expected.19,81
iPSCs derived from GATA2-deficient patients do not show significant defects in differentiating to NK cells.79 The reasons for these differences are not understood. GATA2 is predominately expressed in CD56bright cells, which may indicate its vital role in maintaining this early immature subset.19 However, it has been shown that CD56+ NK cells only express GATA3 and that GATA2 is detected uniquely on HSCs.81 Thus, the expression of GATA2 on the CD56bright NK subset requires detailed analyses to substantiate its role in early developmental stages.
NK cells in patients with GATA2 haploinsufficiency exhibit severe functional defects, including cytotoxicity of remaining CD56dim cells.19 Severe HPV or EBV infection in these patients further indicates dysfunctional mature CD56dim NK cells at multiple levels, although precise molecular defects are so far unknown. A reduced expression of effector molecules, such as perforin and different granzymes, could be due to a reduction in responsible TFs, including T-bet and EOMES.81 NK cells in these patients also express less PLZF, FceRg, and SYK, which define adaptive NK cells.81 Adaptive NK cells persist longer term, but it is not clear whether increased adaptive NK cells can protect patients from viral infections. NK cell development is not significantly altered in PLZF-null mice, suggesting other transcription factors downstream of GATA2.113 NK cells from these patients have lower levels of CXCR4 and dysfunctional CXCL12/CXCR4-mediated chemotaxis.35,82 Proper functioning of the CXCL12/CXCR4 axis is essential for the bone marrow homing of NK cells.114
It is not clear whether impairment of the CXCL12/CXCR4 axis results in reduced cell numbers in the bone marrow of patients with GATA2 haploinsufficiency. As a master regulator, GATA2 may cooperate with a network of TFs to govern human NK cell development. ETS1, one GATA2 upstream regulator,58 NFIL3, a GATA2 downstream target,106,107 and several other TFs interacting with GATA2, such as PLZF115 and PU.1,95 all play critical roles in NK cell development and functions. Future studies should focus on whether an ETS1-GATA2-NFIL3 transcriptional axis governs human NK cell development (Fig. 2).
VI. SUMMARY AND FUTURE DIRECTIONS
Human NK cells account for around 10%–20% of circulating lymphocytes in the peripheral blood. Their ability to kill malignant cells without prior sensitization indicates a promising role in immunotherapeutic applications. Driving the full functional potential of NK cells in patients or ex vivo is essential for successful clinical application. Achieving success in immunotherapy requires a thorough understanding of the TF network and its potential mechanisms of action in human NK cell development. Most known TFs of NK cells are based on murine models, which have not been fully replicated on human NK cells. The human disease conditions that specifically lack either a CD56bright or a CD56dim population provide a unique opportunity to understand human NK cell development.
ACKNOWLEDGMENTS
We dedicate this work to heroic patients with GATA2 deficiency who motivate us in our therapeutic endeavors, and to our inspiring colleague Dr. Mathew Riese MD, PhD, who passed away young in December 2020. This work was supported in part by NIH R01 AI102893 and NCI R01 CA179363 (S.M.); HRHM Program of MACC Fund (S.M.), Nicholas Family Foundation (S.M.); Gardetto Family (S.M.); MCW-Cancer Center-Large Seed Grant (S.M. and M.S.T.); MACC Fund (S.M.); Ann’s Hope Melanoma Foundation (S.M.); and Advancing Healthier Wisconsin (S.M.).
ABBREVIATIONS:
- ADCC
antibody-dependent cell–mediated cytotoxicity
- AML
acute myeloid leukemia
- CLP
common lymphoid progenitor
- DC
dendritic cell
- HSC
hematopoietic stem cell
- iNK
immature NK cell
- iPSC
inducible pluripotent stem cell
- MDS
myeloid dysplastic syndrome
- mNK
mature NK cell
- NK
natural killer
- NKP
NK cell progenitor
- TF
transcription factor
- ZF
zinc finger
REFERENCES
- 1.Tsai FY, Keller G, Kuo FC, Weiss M, Chen J, Rosenblatt M, Alt FW, Orkin SH. An early haematopoietic defect in mice lacking the transcription factor GATA-2. Nature. 1994. Sep 15;371(6494):221–6. PubMed PMID: 8078582. Epub 1994/09/15. [DOI] [PubMed] [Google Scholar]
- 2.Tsai FY, Orkin SH. Transcription factor GATA-2 is required for proliferation/survival of early hematopoietic cells and mast cell formation, but not for erythroid and myeloid terminal differentiation. Blood. 1997. May 15;89(10):3636–43. PubMed PMID: 9160668. Epub 1997/05/15. [PubMed] [Google Scholar]
- 3.Ling KW, Ottersbach K, van Hamburg JP, Oziemlak A, Tsai FY, Orkin SH, Ploemacher R, Hendriks RW, Dzierzak E. GATA-2 plays two functionally distinct roles during the ontogeny of hematopoietic stem cells. J Exp Med. 2004. Oct 4;200(7):871–82. PubMed PMID: 15466621. PMCID: PMC2213282. Epub 2004/10/07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ostergaard P, Simpson MA, Connell FC, Steward CG, Brice G, Woollard WJ, Dafou D, Kilo T, Smithson S, Lunt P, Murday VA, Hodgson S, Keenan R, Pilz DT, Martinez-Corral I, Makinen T, Mortimer PS, Jeffery S, Trembath RC, Mansour S. Mutations in GATA2 cause primary lymphedema associated with a predisposition to acute myeloid leukemia (Emberger syndrome). Nat Genet. 2011. Sep 4;43(10):929–31. PubMed PMID: 21892158. Epub 2011/09/06. [DOI] [PubMed] [Google Scholar]
- 5.Kazenwadel J, Secker GA, Liu YJ, Rosenfeld JA, Wildin RS, Cuellar-Rodriguez J, Hsu AP, Dyack S, Fernandez CV, Chong CE, Babic M, Bardy PG, Shimamura A, Zhang MY, Walsh T, Holland SM, Hickstein DD, Horwitz MS, Hahn CN, Scott HS, Harvey NL. Loss-of-function germline GATA2 mutations in patients with MDS/AML or MonoMAC syndrome and primary lymphedema reveal a key role for GATA2 in the lymphatic vasculature. Blood. 2012. Feb 2;119(5):1283–91. PubMed PMID: 22147895. PMCID: PMC3277359. Epub 2011/12/08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Minegishi N, Suzuki N, Yokomizo T, Pan X, Fujimoto T, Takahashi S, Hara T, Miyajima A, Nishikawa S, Yamamoto M. Expression and domain-specific function of GATA-2 during differentiation of the hematopoietic precursor cells in midgestation mouse embryos. Blood. 2003. Aug 1;102(3):896–905. PubMed PMID: 12689939. Epub 2003/04/12. [DOI] [PubMed] [Google Scholar]
- 7.Viger RS, Guittot SM, Anttonen M, Wilson DB, Heikinheimo M. Role of the GATA family of transcription factors in endocrine development, function, and disease. Mol Endocrinol. 2008. Apr;22(4):781–98. PubMed PMID: 18174356. PMCID: PMC2276466. Epub 2008/01/05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Orkin SH, Zon LI. Hematopoiesis: An evolving paradigm for stem cell biology. Cell. 2008. Feb 22;132(4):631–44. PubMed PMID: 18295580. PMCID: PMC2628169. Epub 2008/02/26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Beck D, Thoms JA, Perera D, Schutte J, Unnikrishnan A, Knezevic K, Kinston SJ, Wilson NK, O’Brien TA, Gottgens B, Wong JW, Pimanda JE. Genome-wide analysis of transcriptional regulators in human HSPCs reveals a densely interconnected network of coding and noncoding genes. Blood. 2013. Oct 3;122(14):e12–22. PubMed PMID: 23974199. Epub 2013/08/27. [DOI] [PubMed] [Google Scholar]
- 10.Scott EW, Simon MC, Anastasi J, Singh H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science. 1994. Sep 9;265(5178):1573–7. PubMed PMID: 8079170. Epub 1994/09/09. [DOI] [PubMed] [Google Scholar]
- 11.McKercher SR, Torbett BE, Anderson KL, Henkel GW, Vestal DJ, Baribault H, Klemsz M, Feeney AJ, Wu GE, Paige CJ, Maki RA. Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J. 1996. Oct 15;15(20):5647–58. PubMed PMID: 8896458. PMCID: PMC452309. Epub 1996/10/15. [PMC free article] [PubMed] [Google Scholar]
- 12.Ran D, Shia WJ, Lo MC, Fan JB, Knorr DA, Ferrell PI, Ye Z, Yan M, Cheng L, Kaufman DS, Zhang DE. RUNX1a enhances hematopoietic lineage commitment from human embryonic stem cells and inducible pluripotent stem cells. Blood. 2013. Apr 11;121(15):2882–90. PubMed PMID: 23372166. PMCID: PMC3624936. Epub 2013/02/02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wilson NK, Foster SD, Wang X, Knezevic K, Schutte J, Kaimakis P, Chilarska PM, Kinston S, Ouwehand WH, Dzierzak E, Pimanda JE, de Bruijn MF, Gottgens B. Combinatorial transcriptional control in blood stem/progenitor cells: Genome-wide analysis of ten major transcriptional regulators. Cell Stem Cell. 2010. Oct 8;7(4):532–44. PubMed PMID: 20887958. Epub 2010/10/05. [DOI] [PubMed] [Google Scholar]
- 14.Vinh DC, Patel SY, Uzel G, Anderson VL, Freeman AF, Olivier KN, Spalding C, Hughes S, Pittaluga S, Raffeld M, Sorbara LR, Elloumi HZ, Kuhns DB, Turner ML, Cowen EW, Fink D, Long-Priel D, Hsu AP, Ding L, Paulson ML, Whitney AR, Sampaio EP, Frucht DM, DeLeo FR, Holland SM. Autosomal dominant and sporadic monocytopenia with susceptibility to mycobacteria, fungi, papillomaviruses, and myelodysplasia. Blood. 2010. Feb 25;115(8):1519–29. PubMed PMID: 20040766. PMCID: PMC2830758. Epub 2009/12/31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bigley V, Haniffa M, Doulatov S, Wang XN, Dickinson R, McGovern N, Jardine L, Pagan S, Dimmick I, Chua I, Wallis J, Lordan J, Morgan C, Kumararatne DS, Doffinger R, van der Burg M, van Dongen J, Cant A, Dick JE, Hambleton S, Collin M. The human syndrome of dendritic cell, monocyte, B and NK lymphoid deficiency. J Exp Med. 2011. Feb 14;208(2):227–34. PubMed PMID: 21242295. PMCID: PMC3039861. Epub 2011/01/19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dickinson RE, Griffin H, Bigley V, Reynard LN, Hussain R, Haniffa M, Lakey JH, Rahman T, Wang XN, McGovern N, Pagan S, Cookson S, McDonald D, Chua I, Wallis J, Cant A, Wright M, Keavney B, Chinnery PF, Loughlin J, Hambleton S, Santibanez-Koref M, Collin M. Exome sequencing identifies GATA-2 mutation as the cause of dendritic cell, monocyte, B and NK lymphoid deficiency. Blood. 2011. Sep 8;118(10):2656–8. PubMed PMID: 21765025. PMCID: PMC5137783. Epub 2011/07/19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hahn CN, Chong CE, Carmichael CL, Wilkins EJ, Brautigan PJ, Li XC, Babic M, Lin M, Carmagnac A, Lee YK, Kok CH, Gagliardi L, Friend KL, Ekert PG, Butcher CM, Brown AL, Lewis ID, To LB, Timms AE, Storek J, Moore S, Altree M, Escher R, Bardy PG, Suthers GK, D’Andrea RJ, Horwitz MS, Scott HS. Heritable GATA2 mutations associated with familial myelodysplastic syndrome and acute myeloid leukemia. Nat Genet. 2011. Sep 4;43(10):1012–7. PubMed PMID: 21892162. PMCID: PMC3184204. Epub 2011/09/06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Spinner MA, Sanchez LA, Hsu AP, Shaw PA, Zerbe CS, Calvo KR, Arthur DC, Gu W, Gould CM, Brewer CC, Cowen EW, Freeman AF, Olivier KN, Uzel G, Zelazny AM, Daub JR, Spalding CD, Claypool RJ, Giri NK, Alter BP, Mace EM, Orange JS, Cuellar-Rodriguez J, Hickstein DD, Holland SM. GATA2 deficiency: A protean disorder of hematopoiesis, lymphatics, and immunity. Blood. 2014. Feb 6;123(6):809–21. PubMed PMID: 24227816. PMCID: PMC3916876. Epub 2013/11/15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mace EM, Hsu AP, Monaco-Shawver L, Makedonas G, Rosen JB, Dropulic L, Cohen JI, Frenkel EP, Bagwell JC, Sullivan JL, Biron CA, Spalding C, Zerbe CS, Uzel G, Holland SM, Orange JS. Mutations in GATA2 cause human NK cell deficiency with specific loss of the CD56(bright) subset. Blood. 2013. Apr 4;121(14):2669–77. PubMed PMID: 23365458. PMCID: PMC3617632. Epub 2013/02/01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vivier E, Tomasello E, Baratin M, Walzer T, Ugolini S. Functions of natural killer cells. Nat Immunol. 2008. May;9(5):503–10. PubMed PMID: 18425107. Epub 2008/04/22. [DOI] [PubMed] [Google Scholar]
- 21.Dokun AO, Kim S, Smith HR, Kang HS, Chu DT, Yokoyama WM. Specific and nonspecific NK cell activation during virus infection. Nat Immunol. 2001. Oct;2(10):951–6. PubMed PMID: 11550009. Epub 2001/09/11. [DOI] [PubMed] [Google Scholar]
- 22.Blanchard DK, Michelini-Norris MB, Pearson CA, Freitag CS, Djeu JY. Mycobacterium avium-intracellulare induces interleukin-6 from human monocytes and large granular lymphocytes. Blood. 1991. May 15;77(10):2218–24. PubMed PMID: 2029580. Epub 1991/05/15. [PubMed] [Google Scholar]
- 23.van den Bosch G, Preijers F, Vreugdenhil A, Hendriks J, Maas F, De Witte T. Granulocyte-macrophage colony-stimulating factor (GM-CSF) counteracts the inhibiting effect of monocytes on natural killer (NK) cells. Clin Exp Immunol. 1995. Sep;101(3):515–20. PubMed PMID: 7664499. PMCID: PMC1553231. Epub 1995/09/01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ojo E, Wigzell H. Natural killer cells may be the only cells in normal mouse lymphoid cell populations endowed with cytolytic ability for antibody-coated tumour target cells. Scand J Immunol. 1978. Apr;7(4):297–306. PubMed PMID: 418499. Epub 1978/04/01. [DOI] [PubMed] [Google Scholar]
- 25.Jewett A, Man YG, Cacalano N, Kos J, Tseng HC. Natural killer cells as effectors of selection and differentiation of stem cells: Role in resolution of inflammation. J Immunotoxicol. 2014. Oct;11(4):297–307. PubMed PMID: 24575813. Epub 2014/03/01. [DOI] [PubMed] [Google Scholar]
- 26.Kiessling R, Klein E, Wigzell H. “Natural” killer cells in the mouse. I. Cytotoxic cells with specificity for mouse Moloney leukemia cells. Specificity and distribution according to genotype. Eur J Immunol. 1975. Feb;5(2):112–7. PubMed PMID: 1234049. Epub 1975/02/01. [DOI] [PubMed] [Google Scholar]
- 27.Kiessling R, Klein E, Pross H, Wigzell H. “Natural” killer cells in the mouse. II. Cytotoxic cells with specificity for mouse Moloney leukemia cells. Characteristics of the killer cell. Eur J Immunol. 1975. Feb;5(2):117–21. PubMed PMID: 1086218. Epub 1975/02/01. [DOI] [PubMed] [Google Scholar]
- 28.Boissel L, Betancur-Boissel M, Lu W, Krause DS, Van Etten RA, Wels WS, Klingemann H. Retargeting NK-92 cells by means of CD19- and CD20-specific chimeric antigen receptors compares favorably with antibody-dependent cellular cytotoxicity. Oncoimmunology. 2013. Oct 1;2(10):e26527. PubMed PMID: 24404423. PMCID: PMC3881109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu E, Marin D, Banerjee P, Macapinlac HA, Thompson P, Basar R, Nassif Kerbauy L, Overman B, Thall P, Kaplan M, Nandivada V, Kaur I, Nunez Cortes A, Cao K, Daher M, Hosing C, Cohen EN, Kebriaei P, Mehta R, Neelapu S, Nieto Y, Wang M, Wierda W, Keating M, Champlin R, Shpall EJ, Rezvani K. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N Engl J Med. 2020. Feb 6;382(6):545–53. PubMed PMID: 32023374. PMCID: PMC7101242. Epub 2020/02/06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Scoville SD, Freud AG, Caligiuri MA. Modeling human natural killer cell development in the era of innate lymphoid cells. Front Immunol. 2017;8:360. PubMed PMID: 28396671. PMCID: PMC5366880. Epub 2017/04/12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang D, Malarkannan S. Transcriptional regulation of natural killer cell development and functions. Cancers. 2020. Jun 16;12(6):1591. PubMed PMID: 32560225. PMCID: PMC7352776. Epub 2020/06/21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Abel AM, Yang C, Thakar MS, Malarkannan S. Natural killer cells: Development, maturation, and clinical utilization. Front Immunol. 2018;9:1869. PubMed PMID: 30150991. PMCID: PMC6099181. Epub 2018/08/29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lanier LL, Spits H, Phillips JH. The developmental relationship between NK cells and T cells. Immunol Today. 1992. Oct;13(10):392–5. [DOI] [PubMed] [Google Scholar]
- 34.Phillips JH, Hori T, Nagler A, Bhat N, Spits H, Lanier LL. Ontogeny of human natural killer (NK) cells: Fetal NK cells mediate cytolytic function and express cytoplasmic CD3 epsilon, delta proteins. J Exp Med. 1992;175(4):1055–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yang C, Siebert JR, Burns R, Gerbec ZJ, Bonacci B, Rymaszewski A, Rau M, Riese MJ, Rao S, Carlson KS, Routes JM, Verbsky JW, Thakar MS, Malarkannan S. Heterogeneity of human bone marrow and blood natural killer cells defined by single-cell transcriptome. Nat Commun. 2019. Sep 2;10(1):3931. PubMed PMID: 31477722. PMCID: PMC6718415. Epub 2019/09/04. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Crinier A, Milpied P, Escaliere B, Piperoglou C, Galluso J, Balsamo A, Spinelli L, Cervera-Marzal I, Ebbo M, Girard-Madoux M, Jaeger S, Bollon E, Hamed S, Hardwigsen J, Ugolini S, Vely F, Narni-Mancinelli E, Vivier E. High-dimensional single-cell analysis identifies organ-specific signatures and conserved NK cell subsets in humans and mice. Immunity. 2018. Nov 1;49(5):971–86. e5. PubMed PMID: 30413361. Epub 2018/11/11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.McKenna HJ, Stocking KL, Miller RE, Brasel K, De Smedt T, Maraskovsky E, Maliszewski CR, Lynch DH, Smith J, Pulendran B, Roux ER, Teepe M, Lyman SD, Peschon JJ. Mice lacking flt3 ligand have deficient hematopoiesis affecting hematopoietic progenitor cells, dendritic cells, and natural killer cells. Blood. 2000. Jun 1;95(11):3489–97. PubMed PMID: 10828034. Epub 2000/05/29. [PubMed] [Google Scholar]
- 38.Mackarehtschian K, Hardin JD, Moore KA, Boast S, Goff SP, Lemischka IR. Targeted disruption of the flk2/flt3 gene leads to deficiencies in primitive hematopoietic progenitors. Immunity. 1995. Jul;3(1):147–61. PubMed PMID: 7621074. Epub 1995/07/01. [DOI] [PubMed] [Google Scholar]
- 39.Reilly JT. Receptor tyrosine kinases in normal and malignant haematopoiesis. Blood Rev. 2003. Dec;17(4):241–8. PubMed PMID: 14556779. Epub 2003/10/15. [DOI] [PubMed] [Google Scholar]
- 40.Mrozek E, Anderson P, Caligiuri MA. Role of interleukin-15 in the development of human CD56+ natural killer cells from CD34+ hematopoietic progenitor cells. Blood. 1996. Apr 1;87(7):2632–40. PubMed PMID: 8639878. Epub 1996/04/01. [PubMed] [Google Scholar]
- 41.Vosshenrich CA, Ranson T, Samson SI, Corcuff E, Colucci F, Rosmaraki EE, Di Santo JP. Roles for common cytokine receptor gamma-chain-dependent cytokines in the generation, differentiation, and maturation of NK cell precursors and peripheral NK cells in vivo. J Immunol. 2005. Feb 1;174(3):1213–21. PubMed PMID: 15661875. Epub 2005/01/22. [DOI] [PubMed] [Google Scholar]
- 42.Kundig TM, Schorle H, Bachmann MF, Hengartner H, Zinkernagel RM, Horak I. Immune responses in interleukin-2-deficient mice. Science. 1993. Nov 12;262(5136):1059–61. PubMed PMID: 8235625. Epub 1993/11/12. [DOI] [PubMed] [Google Scholar]
- 43.Gasteiger G, Hemmers S, Firth MA, Le Floc’h A, Huse M, Sun JC, Rudensky AY. IL-2-dependent tuning of NK cell sensitivity for target cells is controlled by regulatory T cells. J Exp Med. 2013. Jun 3;210(6):1167–78. PubMed PMID: 23650441. PMCID: PMC3674692. Epub 2013/05/08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cho D, Campana D. Expansion and activation of natural killer cells for cancer immunotherapy. Korean J Lab Med. 2009. Apr;29(2):89–96. PubMed PMID: 19411773. PMCID: PMC2771620. Epub 2009/05/05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wendt K, Wilk E, Buyny S, Schmidt RE, Jacobs R. Interleukin-21 differentially affects human natural killer cell subsets. Immunology. 2007. Dec;122(4):486–95. PubMed PMID: 17635612. PMCID: PMC2266033. Epub 2007/07/20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lusty E, Poznanski SM, Kwofie K, Mandur TS, Lee DA, Richards CD, Ashkar AA. IL-18/IL-15/IL-12 synergy induces elevated and prolonged IFN-gamma production by ex vivo expanded NK cells which is not due to enhanced STAT4 activation. Mol Immunol. 2017. Aug;88:138–47. PubMed PMID: 28644973. Epub 2017/06/24. [DOI] [PubMed] [Google Scholar]
- 47.Allan DS, Rybalov B, Awong G, Zuniga-Pflucker JC, Kopcow HD, Carlyle JR, Strominger JL. TGF-beta affects development and differentiation of human natural killer cell subsets. Eur J Immunol. 2010. Aug;40(8):2289–95. PubMed PMID: 20540115. PMCID: PMC3066635. Epub 2010/06/12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Marcoe JP, Lim JR, Schaubert KL, Fodil-Cornu N, Matka M, McCubbrey AL, Farr AR, Vidal SM, Laouar Y. TGF-beta is responsible for NK cell immaturity during ontogeny and increased susceptibility to infection during mouse infancy. Nat Immunol. 2012. Sep;13(9):843–50. PubMed PMID: 22863752. PMCID: PMC3426626. Epub 2012/08/07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gascoyne DM, Long E, Veiga-Fernandes H, de Boer J, Williams O, Seddon B, Coles M, Kioussis D, Brady HJ. The basic leucine zipper transcription factor E4BP4 is essential for natural killer cell development. Nat Immunol. 2009;10(10):1118–24. [DOI] [PubMed] [Google Scholar]
- 50.Taveirne S, Wahlen S, Van Loocke W, Kiekens L, Persyn E, Van Ammel E, De Mulder K, Roels J, Tilleman L, Aumercier M, Matthys P, Van Nieuwerburgh F, Kerre TCC, Taghon T, Van Vlierberghe P, Vandekerckhove B, Leclercq G. The transcription factor ETS1 is an important regulator of human NK cell development and terminal differentiation. Blood. 2020. Jul 16;136(3):288–98. PubMed PMID: 32350509. Epub 2020/05/01. [DOI] [PubMed] [Google Scholar]
- 51.Barton K, Muthusamy N, Fischer C, Ting CN, Walunas TL, Lanier LL, Leiden JM. The Ets-1 transcription factor is required for the development of natural killer cells in mice. Immunity. 1998;9(4):555–63. PMID: 9806641. doi: 10.1016/s1074-7613(00)80638-x. [DOI] [PubMed] [Google Scholar]
- 52.Schotte R, Dontje W, Nagasawa M, Yasuda Y, Bakker AQ, Spits H, Blom B. Synergy between IL-15 and Id2 promotes the expansion of human NK progenitor cells, which can be counteracted by the E protein HEB required to drive T cell development. J Immunol. 2010. Jun 15;184(12):6670–9. PubMed PMID: 20483740. Epub 2010/05/21. [DOI] [PubMed] [Google Scholar]
- 53.Ohno S, Sato T, Kohu K, Takeda K, Okumura K, Satake M, Habu S. Runx proteins are involved in regulation of CD122, Ly49 family and IFN-gamma expression during NK cell differentiation. Int Immunol. 2008. Jan;20(1):71–9. PubMed PMID: 18003603. Epub 2007/11/16. [DOI] [PubMed] [Google Scholar]
- 54.Lai CB, Mager DL. Role of runt-related transcription factor 3 (RUNX3) in transcription regulation of natural cytotoxicity receptor 1 (NCR1/NKp46), an activating natural killer (NK) cell receptor. J Biol Chem. 2012. Mar 2;287(10):7324–34. PubMed PMID: 22253448. PMCID: PMC3293567. Epub 2012/01/19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Barton K, Muthusamy N, Fischer C, Ting CN, Walunas TL, Lanier LL, Leiden JM. The Ets-1 transcription factor is required for the development of natural killer cells in mice. Immunity. 1998. Oct;9(4):555–63. PubMed PMID: 9806641. Epub 1998/11/07.eng. [DOI] [PubMed] [Google Scholar]
- 56.Bories JC, Willerford DM, Grevin D, Davidson L, Camus A, Martin P, Stehelin D, Alt FW. Increased T-cell apoptosis and terminal B-cell differentiation induced by inactivation of the Ets-1 proto-oncogene. Nature. 1995. Oct 19;377(6550):635–8. PubMed PMID: 7566176. Epub 1995/10/19. [DOI] [PubMed] [Google Scholar]
- 57.Ramirez K, Chandler KJ, Spaulding C, Zandi S, Sigvardsson M, Graves BJ, Kee BL. Gene deregulation and chronic activation in natural killer cells deficient in the transcription factor ETS1. Immunity. 2012. Jun 29;36(6):921–32. PubMed PMID: 22608498. PMCID: PMC3389314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Lulli V, Romania P, Morsilli O, Gabbianelli M, Pagliuca A, Mazzeo S, Testa U, Peschle C, Marziali G. Overexpression of Ets-1 in human hematopoietic progenitor cells blocks erythroid and promotes megakaryocytic differentiation. Cell Death Differ. 2006. Jul;13(7):1064–74. PubMed PMID: 16294212. Epub 2005/11/19. [DOI] [PubMed] [Google Scholar]
- 59.Chaves P, Zriwil A, Wittmann L, Boukarabila H, Peitzsch C, Jacobsen SEW, Sitnicka E. Loss of canonical notch signaling affects multiple steps in NK cell development in mice. J Immunol. 2018. Dec 1;201(11):3307–19. PubMed PMID: 30366956. Epub 2018/10/28. [DOI] [PubMed] [Google Scholar]
- 60.Colucci F, Samson SI, DeKoter RP, Lantz O, Singh H, Di Santo JP. Differential requirement for the transcription factor PU.1 in the generation of natural killer cells versus B and T cells. Blood. 2001. May 1;97(9):2625–32. PubMed PMID: 11313251. Epub 2001/04/21. [DOI] [PubMed] [Google Scholar]
- 61.Hosokawa H, Ungerback J, Wang X, Matsumoto M, Nakayama KI, Cohen SM, Tanaka T, Rothenberg EV. Transcription factor PU.1 represses and activates gene expression in early T cells by redirecting partner transcription factor binding. Immunity. 2018. Jun 19;48(6):1119–34 e7. PubMed PMID: 29924977. PMCID: PMC6063530. Epub 2018/06/21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Grund EM, Spyropoulos DD, Watson DK, Muise-Helmericks RC. Interleukins 2 and 15 regulate Ets1 expression via ERK1/2 and MNK1 in human natural killer cells. J Biol Chem. 2005. Feb 11;280(6):4772–8. PubMed PMID: 15563472. Epub 2004/11/26. [DOI] [PubMed] [Google Scholar]
- 63.Gascoyne DM, Long E, Veiga-Fernandes H, de Boer J, Williams O, Seddon B, Coles M, Kioussis D, Brady HJ. The basic leucine zipper transcription factor E4BP4 is essential for natural killer cell development. Nat Immunol. 2009. Oct;10(10):1118–24. PubMed PMID: 19749763. Epub 2009/09/15. [DOI] [PubMed] [Google Scholar]
- 64.Yang M, Li D, Chang Z, Yang Z, Tian Z, Dong Z. PDK1 orchestrates early NK cell development through induction of E4BP4 expression and maintenance of IL-15 responsiveness. J Exp Med. 2015. Feb 09;212(2):253–65. PubMed PMID: 25624444. PMCID: PMC4322053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Male V, Nisoli I, Kostrzewski T, Allan DS, Carlyle JR, Lord GM, Wack A, Brady HJ. The transcription factor E4bp4/Nfil3 controls commitment to the NK lineage and directly regulates Eomes and Id2 expression. J Exp Med. 2014. Apr 07;211(4):635–42. PubMed PMID: 24663216. PMCID: PMC3978281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Yu YL, Chiang YJ, Yen JJ. GATA factors are essential for transcription of the survival gene E4bp4 and the viability response of interleukin-3 in Ba/F3 hematopoietic cells. J Biol Chem. 2002. Jul 26;277(30):27144–53. PubMed PMID: 12023274. Epub 2002/05/23. [DOI] [PubMed] [Google Scholar]
- 67.Ikushima S, Inukai T, Inaba T, Nimer SD, Cleveland JL, Look AT. Pivotal role for the NFIL3/E4BP4 transcription factor in interleukin 3-mediated survival of pro-B lymphocytes. Proc Natl Acad Sci U S A. 1997. Mar 18;94(6):2609–14. PubMed PMID: 9122243. PMCID: PMC20136. Epub 1997/03/18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Yang C, Siebert JR, Burns R, Zheng Y, Mei A, Bonacci B, Wang D, Urrutia RA, Riese MJ, Rao S, Carlson KS, Thakar MS, Malarkannan S. Single-cell transcriptome reveals the novel role of T-bet in suppressing the immature NK gene signature. Elife. 2020. May 14;9. PubMed PMID: 32406817. PMCID: PMC7255804. Epub 2020/05/15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Yang C, Malarkannan S. Transcriptional regulation of NK cell development by mTOR complexes. Front Cell Dev Biol. 2020;8:566090. PubMed PMID: 33240877. PMCID: PMC7683515. Epub 2020/11/27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Townsend MJ, Weinmann AS, Matsuda JL, Salomon R, Farnham PJ, Biron CA, Gapin L, Glimcher LH. T-bet regulates the terminal maturation and homeostasis of NK and Valpha14i NKT cells. Immunity. 2004;20(4):477–94. [DOI] [PubMed] [Google Scholar]
- 71.Gordon SM, Chaix J, Rupp LJ, Wu J, Madera S, Sun JC, Lindsten T, Reiner SL. The transcription factors T-bet and Eomes control key checkpoints of natural killer cell maturation. Immunity. 2012. Jan 27;36(1):55–67. PubMed PMID: 22261438. PMCID: PMC3381976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Townsend MJ, Weinmann AS, Matsuda JL, Salomon R, Farnham PJ, Biron CA, Gapin L, Glimcher LH. T-bet regulates the terminal maturation and homeostasis of NK and Valpha14i NKT cells. Immunity. 2004. Apr;20(4):477–94. PubMed PMID: 15084276. Epub 2004/04/16. [DOI] [PubMed] [Google Scholar]
- 73.Grange M, Verdeil G, Arnoux F, Griffon A, Spicuglia S, Maurizio J, Buferne M, Schmitt-Verhulst AM, Auphan-Anezin N. Active STAT5 regulates T-bet and eomesodermin expression in CD8 T cells and imprints a T-bet-dependent Tc1 program with repressed IL-6/TGF-beta1 signaling. J Immunol. 2013. Oct 1;191(7):3712–24. PubMed PMID: 24006458. Epub 2013/09/06. [DOI] [PubMed] [Google Scholar]
- 74.Comijn J, Berx G, Vermassen P, Verschueren K, van Grunsven L, Bruyneel E, Mareel M, Huylebroeck D, van Roy F. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol Cell. 2001. Jun;7(6):1267–78. PubMed PMID: 11430829. Epub 2001/06/30. [DOI] [PubMed] [Google Scholar]
- 75.Fang X, Hu X, Zheng Z, Tao K, Wang H, Guan H, Shi J, Ji P, Cai W, Bai X, Zhu X, Han J, Liu J, Hu D. Smad interacting protein 1 influences transforming growth factor-beta1/Smad signaling in extracellular matrix protein production and hypertrophic scar formation. J Mol Histol. 2019. Dec;50(6):503–14. PubMed PMID: 31595443. Epub 2019/10/09. [DOI] [PubMed] [Google Scholar]
- 76.van Helden MJ, Goossens S, Daussy C, Mathieu AL, Faure F, Marcais A, Vandamme N, Farla N, Mayol K, Viel S, Degouve S, Debien E, Seuntjens E, Conidi A, Chaix J, Mangeot P, de Bernard S, Buffat L, Haigh JJ, Huylebroeck D, Lambrecht BN, Berx G, Walzer T. Terminal NK cell maturation is controlled by concerted actions of T-bet and Zeb2 and is essential for melanoma rejection. J Exp Med. 2015. Nov 16;212(12):2015–25. PubMed PMID: 26503444. PMCID: PMC4647267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Bagadia P, Huang X, Liu TT, Durai V, Grajales-Reyes GE, Nitschke M, Modrusan Z, Granja JM, Satpathy AT, Briseno CG, Gargaro M, Iwata A, Kim S, Chang HY, Shaw AS, Murphy TL, Murphy KM. An Nfil3-Zeb2-Id2 pathway imposes Irf8 enhancer switching during cDC1 development. Nat Immunol. 2019. Sep;20(9):1174–85. PubMed PMID: 31406377. PMCID: PMC6707889. Epub 2019/08/14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Ganapathi KA, Townsley DM, Hsu AP, Arthur DC, Zerbe CS, Cuellar-Rodriguez J, Hickstein DD, Rosenzweig SD, Braylan RC, Young NS, Holland SM, Calvo KR. GATA2 deficiency-associated bone marrow disorder differs from idiopathic aplastic anemia. Blood. 2015. Jan 1;125(1):56–70. PubMed PMID: 25359990. PMCID: PMC4281830. Epub 2014/11/02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Jung M, Cordes S, Zou J, Yu SJ, Guitart X, Hong SG, Dang V, Kang E, Donaires FS, Hassan SA, Albitar M, Hsu AP, Holland SM, Hickstein DD, Townsley D, Dunbar CE, Winkler T. GATA2 deficiency and human hematopoietic development modeled using induced pluripotent stem cells. Blood Adv. 2018. Dec 11;2(23):3553–65. PubMed PMID: 30538114. PMCID: PMC6290105. Epub 2018/12/13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Dickinson RE, Milne P, Jardine L, Zandi S, Swierczek SI, McGovern N, Cookson S, Ferozepurwalla Z, Langridge A, Pagan S, Gennery A, Heiskanen-Kosma T, Hamalainen S, Seppanen M, Helbert M, Tholouli E, Gambineri E, Reykdal S, Gottfreethsson M, Thaventhiran JE, Morris E, Hirschfield G, Richter AG, Jolles S, Bacon CM, Hambleton S, Haniffa M, Bryceson Y, Allen C, Prchal JT, Dick JE, Bigley V, Collin M. The evolution of cellular deficiency in GATA2 mutation. Blood. 2014. Feb 6;123(6):863–74. PubMed PMID: 24345756. PMCID: PMC3916878. Epub 2013/12/19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Schlums H, Jung M, Han H, Theorell J, Bigley V, Chiang SC, Allan DS, Davidson-Moncada JK, Dickinson RE, Holmes TD, Hsu AP, Townsley D, Winkler T, Wang W, Aukrust P, Nordoy I, Calvo KR, Holland SM, Collin M, Dunbar CE, Bryceson YT. Adaptive NK cells can persist in patients with GATA2 mutation depleted of stem and progenitor cells. Blood. 2017. Apr 6;129(14):1927–39. PubMed PMID: 28209719. PMCID: PMC5383869. Epub 2017/02/18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Maciejewski-Duval A, Meuris F, Bignon A, Aknin ML, Balabanian K, Faivre L, Pasquet M, Barlogis V, Fieschi C, Bellanne-Chantelot C, Donadieu J, Schlecht-Louf G, Marin-Esteban V, Bachelerie F. Altered chemotactic response to CXCL12 in patients carrying GATA2 mutations. J Leukoc Biol. 2016. Jun;99(6):1065–76. PubMed PMID: 26710799. Epub 2015/12/30. [DOI] [PubMed] [Google Scholar]
- 83.Almehmadi M, Flanagan BF, Khan N, Alomar S, Christmas SE. Increased numbers and functional activity of CD56+ T cells in healthy cytomegalovirus positive subjects. Immunology. 2014;142(2):258–68. PubMed PMID: 24433347.eng. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Waskow C, Liu K, Darrasse-Jeze G, Guermonprez P, Ginhoux F, Merad M, Shengelia T, Yao K, Nussenzweig M. The receptor tyrosine kinase Flt3 is required for dendritic cell development in peripheral lymphoid tissues. Nat Immunol. 2008. Jun;9(6):676–83. PubMed PMID: 18469816. PMCID: PMC2746085. Epub 2008/05/13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wlodarski MW, Collin M, Horwitz MS. GATA2 deficiency and related myeloid neoplasms. Semin Hematol. 2017. Apr;54(2):81–6. PubMed PMID: 28637621. PMCID: PMC5650112. Epub 2017/06/24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Bodor C, Renneville A, Smith M, Charazac A, Iqbal S, Etancelin P, Cavenagh J, Barnett MJ, Kramarzova K, Krishnan B, Matolcsy A, Preudhomme C, Fitzgibbon J, Owen C. Germ-line GATA2 p.THR354MET mutation in familial myelodysplastic syndrome with acquired monosomy 7 and ASXL1 mutation demonstrating rapid onset and poor survival. Haematologica. 2012. Jun;97(6):890–4. PubMed PMID: 22271902. PMCID: PMC3366655. Epub 2012/01/25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Micol JB, Abdel-Wahab O. Collaborating constitutive and somatic genetic events in myeloid malignancies: ASXL1 mutations in patients with germline GATA2 mutations. Haematologica. 2014. Feb;99(2):201–3. PubMed PMID: 24497555. PMCID: PMC3912947. Epub 2014/02/06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Grossmann V, Haferlach C, Nadarajah N, Fasan A, Weissmann S, Roller A, Eder C, Stopp E, Kern W, Haferlach T, Kohlmann A, Schnittger S. CEBPA double-mutated acute myeloid leukaemia harbours concomitant molecular mutations in 76.8% of cases with TET2 and GATA2 alterations impacting prognosis. Br J Haematol. 2013. Jun;161(5):649–58. PubMed PMID: 23521373. Epub 2013/03/26. [DOI] [PubMed] [Google Scholar]
- 89.Greif PA, Dufour A, Konstandin NP, Ksienzyk B, Zellmeier E, Tizazu B, Sturm J, Benthaus T, Herold T, Yaghmaie M, Dorge P, Hopfner KP, Hauser A, Graf A, Krebs S, Blum H, Kakadia PM, Schneider S, Hoster E, Schneider F, Stanulla M, Braess J, Sauerland MC, Berdel WE, Buchner T, Woermann BJ, Hiddemann W, Spiekermann K, Bohlander SK. GATA2 zinc finger 1 mutations associated with biallelic CEBPA mutations define a unique genetic entity of acute myeloid leukemia. Blood. 2012. Jul 12;120(2):395–403. PubMed PMID: 22649106. Epub 2012/06/01. [DOI] [PubMed] [Google Scholar]
- 90.Soukup AA, Zheng Y, Mehta C, Wu J, Liu P, Cao M, Hofmann I, Zhou Y, Zhang J, Johnson KD, Choi K, Keles S, Bresnick EH. Single-nucleotide human disease mutation inactivates a blood-regenerative GATA2 enhancer. J Clin Invest. 2019. Mar 1;129(3):1180–92. PubMed PMID: 30620726. PMCID: PMC6391105. Epub 2019/01/09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Fasan A, Eder C, Haferlach C, Grossmann V, Kohlmann A, Dicker F, Kern W, Haferlach T, Schnittger S. GATA2 mutations are frequent in intermediate-risk karyotype AML with biallelic CEBPA mutations and are associated with favorable prognosis. Leukemia. 2013. Feb;27(2):482–5. PubMed PMID: 22814295. Epub 2012/07/21. [DOI] [PubMed] [Google Scholar]
- 92.Cortes-Lavaud X, Landecho MF, Maicas M, Urquiza L, Merino J, Moreno-Miralles I, Odero MD. GATA2 germline mutations impair GATA2 transcription, causing haploinsufficiency: Functional analysis of the p.Arg-396Gln mutation. J Immunol. 2015. Mar 1;194(5):2190–8. PubMed PMID: 25624456. Epub 2015/01/28. [DOI] [PubMed] [Google Scholar]
- 93.Zhang SJ, Ma LY, Huang QH, Li G, Gu BW, Gao XD, Shi JY, Wang YY, Gao L, Cai X, Ren RB, Zhu J, Chen Z, Chen SJ. Gain-of-function mutation of GATA-2 in acute myeloid transformation of chronic myeloid leukemia. Proc Natl Acad Sci U S A. 2008. Feb 12;105(6):2076–81. PubMed PMID: 18250304. PMCID: PMC2538883. Epub 2008/02/06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Zhang SJ, Shi JY, Li JY. GATA-2 L359 V mutation is exclusively associated with CML progression but not other hematological malignancies and GATA-2 P250A is a novel single nucleotide polymorphism. Leuk Res. 2009. Aug;33(8):1141–3. PubMed PMID: 19304323. Epub 2009/03/24. [DOI] [PubMed] [Google Scholar]
- 95.Zhang P, Behre G, Pan J, Iwama A, Wara-Aswapati N, Radomska HS, Auron PE, Tenen DG, Sun Z. Negative cross-talk between hematopoietic regulators: GATA proteins repress PU.1. Proc Natl Acad Sci U S A. 1999. Jul 20;96(15):8705–10. PubMed PMID: 10411939. PMCID: PMC17580. Epub 1999/07/21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Carotta S, Dakic A, D’Amico A, Pang SH, Greig KT, Nutt SL, Wu L. The transcription factor PU.1 controls dendritic cell development and Flt3 cytokine receptor expression in a dose-dependent manner. Immunity. 2010. May 28;32(5):628–41. PubMed PMID: 20510871. Epub 2010/06/01. [DOI] [PubMed] [Google Scholar]
- 97.Chong CE, Venugopal P, Stokes PH, Lee YK, Brautigan PJ, Yeung DTO, Babic M, Engler GA, Lane SW, Klingler-Hoffmann M, Matthews JM, D’Andrea RJ, Brown AL, Hahn CN, Scott HS. Differential effects on gene transcription and hematopoietic differentiation correlate with GATA2 mutant disease phenotypes. Leukemia. 2018. Jan;32(1):194–202. PubMed PMID: 28642594. PMCID: PMC5770593. Epub 2017/06/24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Walsh JC, DeKoter RP, Lee HJ, Smith ED, Lancki DW, Gurish MF, Friend DS, Stevens RL, Anastasi J, Singh H. Cooperative and antagonistic interplay between PU.1 and GATA-2 in the specification of myeloid cell fates. Immunity. 2002. Nov;17(5):665–76. PubMed PMID: 12433372. Epub 2002/11/16. [DOI] [PubMed] [Google Scholar]
- 99.Ohmori S, Ishijima Y, Numata S, Takahashi M, Sekita M, Sato T, Chugun K, Yamamoto M, Ohneda K. GATA2 and PU.1 collaborate to activate the expression of the mouse Ms4a2 gene, encoding FcepsilonRIbeta, through distinct mechanisms. Mol Cell Biol. 2019. Nov 15;39(22):e00314–9. PubMed PMID: 31501274. PMCID: PMC6817753. Epub 2019/09/11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Kuhl C, Atzberger A, Iborra F, Nieswandt B, Porcher C, Vyas P. GATA1-mediated megakaryocyte differentiation and growth control can be uncoupled and mapped to different domains in GATA1. Mol Cell Biol. 2005. Oct;25(19):8592–606. PubMed PMID: 16166640. PMCID: PMC1265752. Epub 2005/09/17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Snow JW, Trowbridge JJ, Johnson KD, Fujiwara T, Emambokus NE, Grass JA, Orkin SH, Bresnick EH. Con text-dependent function of “GATA switch” sites in vivo. Blood. 2011. May 5;117(18):4769–72. PubMed PMID: 21398579. PMCID: PMC3100688. Epub 2011/03/15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Philipsen S A new twist to the GATA switch. Blood. 2013. Nov 14;122(20):3391–2. PubMed PMID: 24235125. Epub 2013/11/16. [DOI] [PubMed] [Google Scholar]
- 103.Bresnick EH, Lee HY, Fujiwara T, Johnson KD, Keles S. GATA switches as developmental drivers. J Biol Chem. 2010. Oct 8;285(41):31087–93. PubMed PMID: 20670937. PMCID: PMC2951181. Epub 2010/07/31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Gao J, Chen YH, Peterson LC. GATA family transcriptional factors: Emerging suspects in hematologic disorders. Exp Hematol Oncol. 2015;4:28. PubMed PMID: 26445707. PMCID: PMC4594744. Epub 2015/10/09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Takai J, Moriguchi T, Suzuki M, Yu L, Ohneda K, Yamamoto M. The Gata1 5’ region harbors distinct cis-regulatory modules that direct gene activation in erythroid cells and gene inactivation in HSCs. Blood. 2013. Nov 14;122(20):3450–60. PubMed PMID: 24021675. Epub 2013/09/12. [DOI] [PubMed] [Google Scholar]
- 106.Seillet C, Rankin LC, Groom JR, Mielke LA, Tellier J, Chopin M, Huntington ND, Belz GT, Carotta S. Nfil3 is required for the development of all innate lymphoid cell subsets. J Exp Med. 2014. Aug 25;211(9):1733–40. PubMed PMID: 25092873. PMCID: PMC4144736. Epub 2014/08/06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kashiwada M, Pham NL, Pewe LL, Harty JT, Rothman PB. NFIL3/E4BP4 is a key transcription factor for CD8alpha(+) dendritic cell development. Blood. 2011. Jun 9;117(23):6193–7. PubMed PMID: 21474667. PMCID: PMC3122942. Epub 2011/04/09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Onodera K, Fujiwara T, Onishi Y, Itoh-Nakadai A, Okitsu Y, Fukuhara N, Ishizawa K, Shimizu R, Yamamoto M, Harigae H. GATA2 regulates dendritic cell differentiation. Blood. 2016. Jul 28;128(4):508–18. PubMed PMID: 27259979. PMCID: PMC5026465. Epub 2016/06/05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Rodrigues NP, Janzen V, Forkert R, Dombkowski DM, Boyd AS, Orkin SH, Enver T, Vyas P, Scadden DT. Haploinsufficiency of GATA-2 perturbs adult hematopoietic stem-cell homeostasis. Blood. 2005. Jul 15;106(2):477–84. PubMed PMID: 15811962. Epub 2005/04/07. [DOI] [PubMed] [Google Scholar]
- 110.Tipping AJ, Pina C, Castor A, Hong D, Rodrigues NP, Lazzari L, May GE, Jacobsen SE, Enver T. High GATA-2 expression inhibits human hematopoietic stem and progenitor cell function by effects on cell cycle. Blood. 2009. Mar 19;113(12):2661–72. PubMed PMID: 19168794. Epub 2009/01/27. [DOI] [PubMed] [Google Scholar]
- 111.Luesink M, Hollink IH, van der Velden VH, Knops RH, Boezeman JB, de Haas V, Trka J, Baruchel A, Reinhardt D, van der Reijden BA, van den Heuvel-Eibrink MM, Zwaan CM, Jansen JH. High GATA2 expression is a poor prognostic marker in pediatric acute myeloid leukemia. Blood. 2012. Sep 6;120(10):2064–75. PubMed PMID: 22786876. Epub 2012/07/13. [DOI] [PubMed] [Google Scholar]
- 112.Vicente C, Vazquez I, Conchillo A, Garcia-Sanchez MA, Marcotegui N, Fuster O, Gonzalez M, Calasanz MJ, Lahortiga I, Odero MD. Overexpression of GATA2 predicts an adverse prognosis for patients with acute myeloid leukemia and it is associated with distinct molecular abnormalities. Leukemia. 2012. Mar;26(3):550–4. PubMed PMID: 21904383. Epub 2011/09/10. [DOI] [PubMed] [Google Scholar]
- 113.Constantinides MG, McDonald BD, Verhoef PA, Bendelac A. A committed precursor to innate lymphoid cells. Nature. 2014. Apr 17;508(7496):397–401. PubMed PMID: 24509713. PMCID: PMC4003507. Epub 2014/ 02/11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Beider K, Nagler A, Wald O, Franitza S, Dagan-Berger M, Wald H, Giladi H, Brocke S, Hanna J, Mandelboim O, Darash-Yahana M, Galun E, Peled A. Involvement of CXCR4 and IL-2 in the homing and retention of human NK and NK T cells to the bone marrow and spleen of NOD/SCID mice. Blood. 2003. Sep 15;102(6):1951–8. PubMed PMID: 12730102. Epub 2003/05/06. [DOI] [PubMed] [Google Scholar]
- 115.Tsuzuki S, Enver T. Interactions of GATA-2 with the promyelocytic leukemia zinc finger (PLZF) protein, its homologue FAZF, and the t(11;17)-generated PLZF-retinoic acid receptor alpha oncoprotein. Blood. 2002. May 1;99(9):3404–10. PubMed PMID: 11964310. Epub 2002/04/20. [DOI] [PubMed] [Google Scholar]