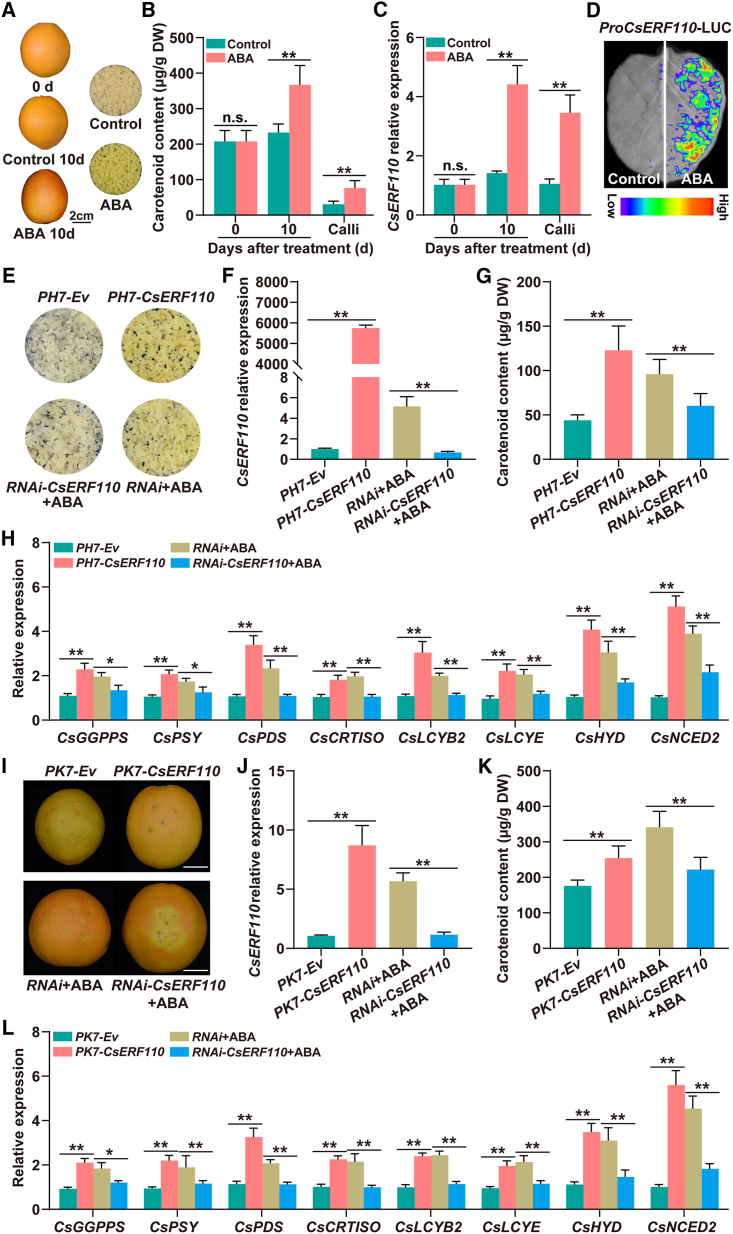
Figure 1.
CsERF110 is essential for ABA-mediated carotenoid biosynthesis in citrus.
(A–C) Phenotypes of citrus fruit and calli (A), (B) carotenoid content, and (C)CsERF110 expression under ABA treatment. Scale bars, 2 cm. (D)In vivo firefly luciferase (LUC) complementation imaging of N. benthamiana leaves treated with water or ABA. A suspension of Agrobacterium GV3101 carrying ProCsERF110-LUC was injected into N. benthamiana leaves. Two days after injection, N. benthamiana leaves were treated with water or 100 μM ABA, and 24 h later, bioluminescence imaging was used to measure the luciferase activity of ProCsERF110.
(E)–(H) Stable transformation of CsERF110 in citrus calli. (E) Phenotypes. PH7-CsERF110 and RNAi-ERF110 indicate CsERF110 overexpression and RNA interference, respectively. The PH7 (PH7-Ev) and RNAi empty vectors are controls. The expression levels of CsERF110(F) and CsGGPPS, CsPSY, CsPDS, CsCRTISO, CsLCYB2, CsLCYE, CsHYD, and CsNCED2(H) are shown. (G) Total carotenoid content (μg/g dry weight [DW]).
(I, J, and L) Transient expression of CsERF110 in citrus fruit. (I) Phenotypes. The PK7 (PK7-Ev) and RNAi empty vectors are controls. PK7-CsERF110 and RNAi-CsERF110 indicate CsERF110 overexpression and RNA interference, respectively. Scale bars, 2 cm. The transcript levels of CsERF110(J) and CsGGPPS, CsPSY, CsPDS, CsCRTISO, CsLCYB2, CsLCYE, CsHYD, and CsNCED2(L) are shown.
(K) Total carotenoid content (μg/g DW). Data are presented as means ± SDs of three biological replicates. Asterisks indicate statistically significant differences determined by Student’s t test (∗p < 0.05; ∗∗p < 0.01; n.s., no significant difference).