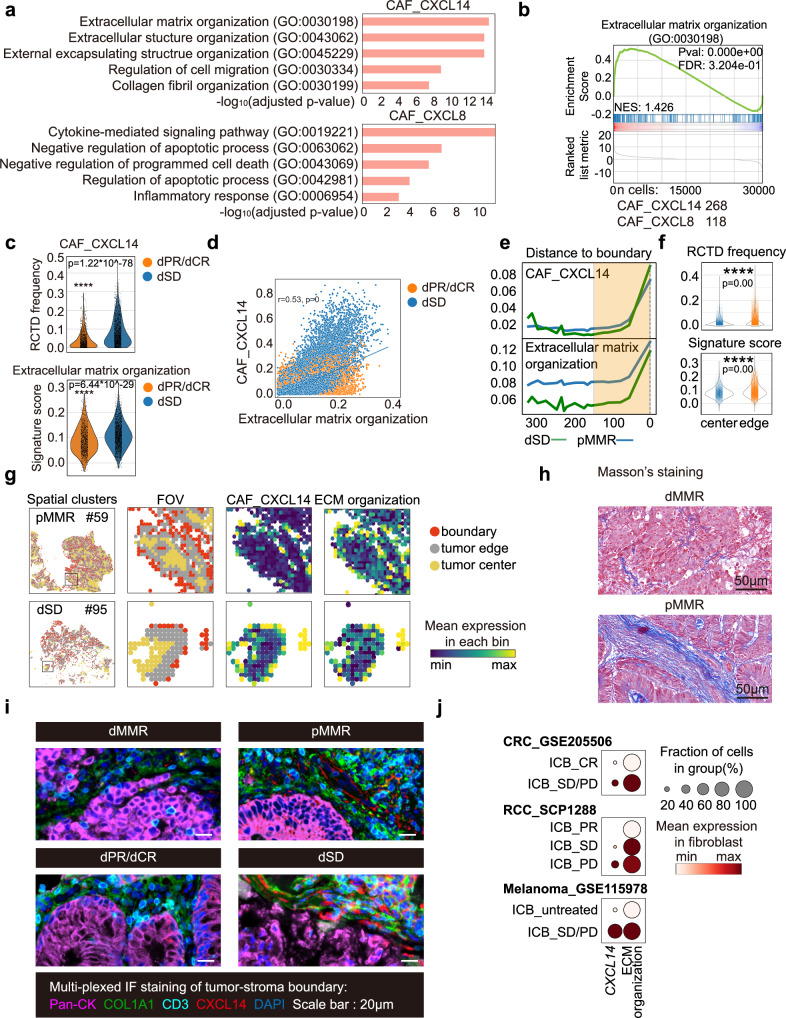
Fig. 5. CXCL14+CAFs may contribute to the well-organized matrix structure and T cell exclusion in TME of ICB non-responders.
a The top 5 GO terms of DEGs (p value < 0.001; fold change >2) identified from CAF_CXCL14 and CAF_CXCL8 clusters are shown. Comparison is made by two-tailed t test. b GSEA plot of the upregulated genes related to extracellular matrix organization in CAF_CXCL14 (n cells=268) compared to CAF_CXCL8 (n cells=118) is shown. Comparison is made by two-tailed t test. c RCTD frequencies of CAF_CXCL14 (up) and ECM organization signature scores (below) in the tumor-stroma boundary of dPR/dCR (n spots=1436) and dSD (n spots=2244) patients are shown as violim plots. Data are analyzed by unpaired 2-tailed Student t test. Ns, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001; ***, p < 0.0001. d Pearson correlation of RCTD frequencies of CAF_CXCL14 and ECM organization signature scores in dPR/dCR and dSD patients. e Loess smoothed curves and (f) violin plots of RCTD frequencies of CAF_CXCL14 (up) and ECM organization signature scores (below) in tumor ( + 300 μm, n spots = 9313) to the tumor-stroma boundary (0 μm, n spots = 56,357) from treatment naïve pMMR and dSD patients are shown. The tumor edge region ( + 150 μm to 0 μm) is highlighted in the yellow frame. g Representative images of FOV, CAF_CXCL14 and ECM organization scores in the tumor-stroma boundary of treatment naïve pMMR patient #59 and dSD patient #95 are shown. h Representative images of Masson’s trichrome staining from treatment naïve dMMR and pMMR patients (3 samples were analyzed in each group). Scale bar = 50 μm. i Representative mIF images of panCK, COL1A1, CD3 and CXCL14 in indicated patient groups (3 samples were analyzed in each group). DAPI was used as a positive control for cell nuclei staining. Scale bars, 50 μm. j CXCL14 expression level and ECM organization scores in fibroblasts from the three public scRNAseq datasets16,70,71.