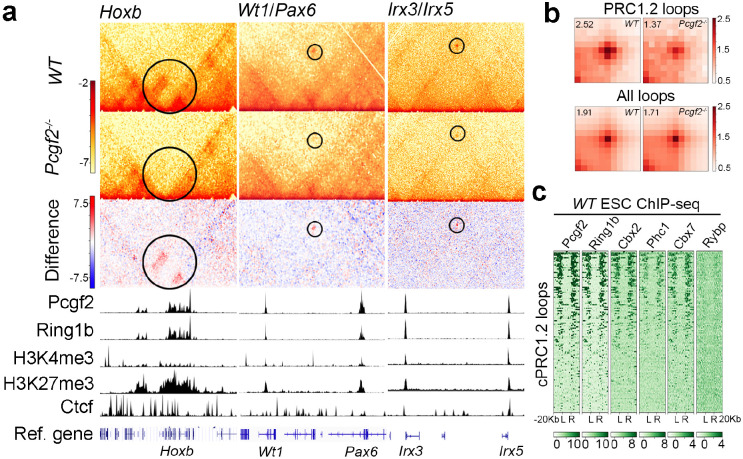
Figure 2. cPRC1.2 is required for chromatin loop formation.
a, Example Hi-C matrices for WT and Pcgf2−/− ESCs at 5-kb resolution showing reduced contact intensities at Hoxb, Wt1/Pax6, and Irx3/Irx5 loci in Pcgf2−/− ESCs. ChIP-seq tracks for Pcgf2, Ring1b, H3K4me3, H3K27me3, and Ctcf are displayed below the Hi-C matrices. b, Aggregate peak analysis (APA) of Hi-C for total chromatin loops and cPRC1.2 loops in WT and Pcgf2−/− ESCs at 10-kb resolution. The pileups are normalized to the average of the top-left and bottom-right corner pixels, and the value of the central pixels is displayed on the top-left side of the plot. c, ChIP-seq heatmaps show enrichment of cPRC1.2 components (Pcgf2, Cbx2, Phc1, and Cbx7) and lack of ncPRC1 component (Rybp) at cPRC1.2 loop anchors in WT ESCs. L and R denote left and right loop anchors, with 20 kb regions flanking each anchor.