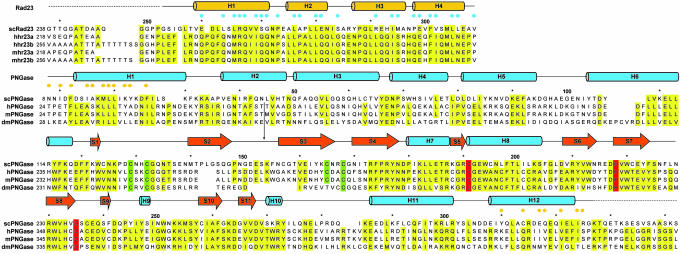
Fig. 2.
Structural alignment and conservation in yRad23XBD and yPNGase. Shown at the top is the sequence identity between Saccharomyces cerevisiae Rad23XBD (yRad23XBD) and its orthologues from human (Hs) and mouse (Mm). Blue dots indicate yRad23XBD residues that contact yPNGase. Every 10th residue is marked with an asterisk. The secondary structural elements are indicated above the alignment. At the bottom, sequence identity between S. cerevisiae PNGase (yPNGase) and its orthologues from human (Hs), mouse (Mm), and fly (Dm, bottom) is shown. Conserved residues are shaded in yellow, and the catalytic triad is highlighted in red. The four Cys residues that coordinate the Zn atom are shown in green. Orange dots indicate yPNGase residues that contact yRad23XBD. Omitted regions of HsPNGase (residues 64-174), MmPNGase (residues 64-171), and DmPNGase (residues 68-165) are indicated by arrows.