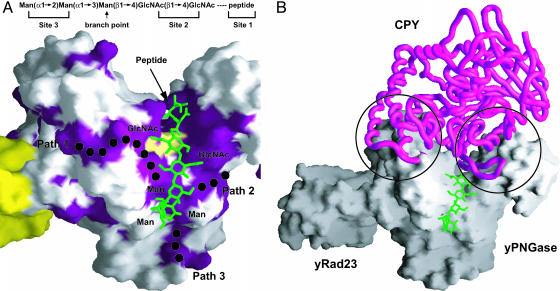
Fig. 6.
Molecular model of the yPNGase-substrate complex. (A) A glycan moiety containing two GlcNAc residues and three mannose molecules was modeled on the active site of the cleft. The dotted lines indicate the three conserved regions where the additional carbohydrate molecules could bind. The common structure of a glycan motif is shown at the top. The close-up view on these conserved regions is shown in Fig. 11. (B) Three Cα atoms from Val-366, Arg-367, and Asn-368 and the GlcNAc residue of yeast carboxypeptidase Y (CPY; Protein Data Bank ID code IYSC) were superimposed onto those from three residues of the inhibitor, Z-VAD-fmk, and a fructose molecule in the active site of yPNGase, respectively. The glycosylated Asn residue of CPY in native form is prevented from accessing the active site by both sides of the deep cleft, which are formed by the strands S2 and S3 in the Zn-binding domain and two loops between H10 and H11 helices and between H9 and S10 of the core domain.